Abstract
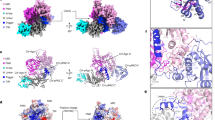
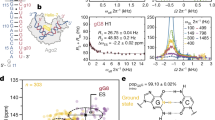
MicroRNAs (miRNAs) mediate post-transcriptional gene regulation through association with Argonaute proteins (AGOs)1. Crystal structures of archaeal and bacterial homologues of AGOs have shown that the MID (middle) domain mediates the interaction with the phosphorylated 5′ end of the miRNA guide strand and this interaction is thought to be independent of the identity of the 5′ nucleotide in these systems2,3. However, analysis of the known sequences of eukaryotic miRNAs and co-immunoprecipitation experiments indicate that there is a clear bias for U or A at the 5′ position4,5,6,7. Here we report the crystal structure of a MID domain from a eukaryotic AGO protein, human AGO2. The structure, in complex with nucleoside monophosphates (AMP, CMP, GMP, and UMP) mimicking the 5′ end of miRNAs, shows that there are specific contacts made between the base of UMP or AMP and a rigid loop in the MID domain. Notably, the structure of the loop discriminates against CMP and GMP and dissociation constants calculated from NMR titration experiments confirm these results, showing that AMP (0.26 mM) and UMP (0.12 mM) bind with up to 30-fold higher affinity than either CMP (3.6 mM) or GMP (3.3 mM). This study provides structural evidence for nucleotide-specific interactions in the MID domain of eukaryotic AGO proteins and explains the observed preference for U or A at the 5′ end of miRNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nature Rev. Mol. Cell Biol. 9, 22–32 (2008)
Ma, J.-B. et al. Structural basis for 5′-end-specific recognition of guide RNA by the A. fulgidus Piwi protein. Nature 434, 666–670 (2005)
Parker, J. S., Roe, S. M. & Barford, D. Structural insights into mRNA recognition from a PIWI domain-siRNA guide complex. Nature 434, 663–666 (2005)
Ghildiyal, M. et al. Endogenous siRNAs derived from transposons and mRNAs in Drosophila somatic cells. Science 320, 1077–1081 (2008)
Ghildiyal, M., Xu, J., Seitz, H., Weng, Z. & Zamore, P. Sorting of Drosophila small silencing RNAs partitions microRNA* strands into the RNA interference pathway. RNA 16, 43–56 (2009)
Hu, H. Y. et al. Sequence features associated with microRNA strand selection in humans and flies. BMC Genomics 10 10.1186/1471-2164-10-413 (2009)
Lau, N. C., Lim, L. P., Weinstein, E. G. & Bartel, D. P. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans . Science 294, 858–862 (2001)
Ghildiyal, M. & Zamore, P. D. Small silencing RNAs: an expanding universe. Nature Rev. Genet. 10, 94–108 (2009)
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nature Rev. Genet. 9, 102–114 (2008)
Seitz, H., Ghildiyal, M. & Zamore, P. D. Argonaute loading improves the 5′ precision of both microRNAs and their miRNA strands in flies. Curr. Biol. 18, 147–151 (2008)
Mi, S. et al. Sorting of small RNAs into Arabidopsis Argonaute complexes is directed by the 5′ terminal nucleotide. Cell 133, 116–127 (2008)
Montgomery, T. A. et al. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133, 128–141 (2008)
Takeda, A., Iwasaki, S., Watanabe, T., Utsumi, M. & Watanabe, Y. The mechanism selecting the guide strand from small RNA duplexes is different among Argonaute proteins. Plant Cell Physiol. 49, 493–500 (2008)
Ma, J.-B., Ye, K. & Patel, D. J. Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429, 318–322 (2004)
Wang, Y. et al. Structure of an Argonaute silencing complex with a seed-containing guide DNA and target RNA duplex. Nature 456, 921–926 (2008)
Wang, Y. et al. Nucleation, propagation and cleavage of target RNAs in Ago silencing complexes. Nature 461, 754–761 (2009)
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D. J. Structure of the guide-strand-containing Argonaute silencing complex. Nature 456, 209–213 (2008)
Parker, J. S., Parizotto, E. A., Wang, M., Roe, S. M. & Barford, D. Enhancement of the seed-target recognition step in RNA silencing by a PIWI/MID domain protein. Mol. Cell 33, 204–214 (2009)
Song, J.-J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004)
Wahl, M. C. & Sundaralingam, M. C–H…O hydrogen bonding in biology. Trends Biochem. Sci. 22, 97–102 (1997)
Felice, K. M., Salzman, D. W., Shubert-Coleman, J., Jensen, K. P. & Furneaux, H. M. The 5′ terminal uracil of let-7a is critical for the recruitment of mRNA to Argonaute2. Biochem. J. 422, 329–341 (2009)
Wu, L. et al. Rice microRNA effector complexes and targets. Plant Cell 21, 3421–3435 (2009)
Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila . Nature 453, 798–802 (2008)
Kiriakidou, M. et al. An mRNA m7G cap binding-like motif within human Ago2 represses translation. Cell 129, 1141–1151 (2007)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Mossessova, E. & Lima, C. D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000)
Otwinowski, Z. Processing of X-ray diffraction data collected in oscillation mode. Meth. Enzymol. 276, 307–326 (1997)
Terwilliger, T. Automated main-chain model building by template matching and iterative fragment extension. Acta Crystallogr. D 59, 38–44 (2002)
Langer, G., Cohen, S. X., Lamzin, V. S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nature Protoc. 3, 1171–1179 (2008)
Adams, P., Grosse-Kunstleve, R. & Hung, L. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Acknowledgements
We thank O. Larsson for help with bioinformatic analysis; J.-F. Trempe, G. Kozlov, S. Azeroual and D. Rodionov for help with X-ray and NMR data collection at the QANUC NMR facility; and K. Gehring and A. Berghuis for sharing resources and critical reading of the manuscript. We also thank M. Fabian, T. Sundermeier and T. Duchaine for comments on the manuscript; W. Filipowicz for discussions; R. Szittner, K. Illes and G. Virgili for technical assistance. B.N. is supported by a Canada Research Chair, a Career Development Award from the Human Frontiers Science Program (CDA 0018/2006-C/1) and an operating grant from the Canadian Institutes of Health Research (CIHR grant MOP-82929). N.S. is funded by a CIHR grant. F.F. is supported by a Boehringer Ingelheim Fonds PhD Fellowship.
Author information
Authors and Affiliations
Contributions
B.N., N.S. and F.F. designed the project. B.N. and F.F. wrote the manuscript. F.F. performed all of the NMR and crystallographic work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures S1-S15 with legends and References. (PDF 10361 kb)
Rights and permissions
About this article
Cite this article
Frank, F., Sonenberg, N. & Nagar, B. Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 465, 818–822 (2010). https://doi.org/10.1038/nature09039
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09039
This article is cited by
-
Structural basis of antiphage immunity generated by a prokaryotic Argonaute-associated SPARSA system
Nature Communications (2024)
-
Unlocking the potential of RNAi as a therapeutic strategy against infectious viruses: an in-silico study
Chemical Papers (2024)
-
Exosomal miRNA-mediated intercellular communications and immunomodulatory effects in tumor microenvironments
Journal of Biomedical Science (2023)
-
Structural basis for sequence-specific recognition of guide and target strands by the Archaeoglobus fulgidus Argonaute protein
Scientific Reports (2023)
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.