Abstract
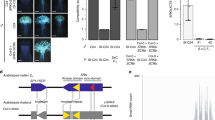
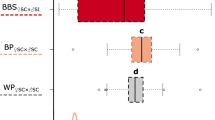
Ever since Darwin’s pioneering research, the evolution of self-fertilisation (selfing) has been regarded as one of the most prevalent evolutionary transitions in flowering plants1,2. A major mechanism to prevent selfing is the self-incompatibility (SI) recognition system, which consists of male and female specificity genes at the S-locus and SI modifier genes2,3,4. Under conditions that favour selfing, mutations disabling the male recognition component are predicted to enjoy a relative advantage over those disabling the female component, because male mutations would increase through both pollen and seeds whereas female mutations would increase only through seeds5,6. Despite many studies on the genetic basis of loss of SI in the predominantly selfing plant Arabidopsis thaliana7,8,9,10,11,12,13,14,15, it remains unknown whether selfing arose through mutations in the female specificity gene (S-receptor kinase, SRK), male specificity gene (S-locus cysteine-rich protein, SCR; also known as S-locus protein 11, SP11) or modifier genes, and whether any of them rose to high frequency across large geographic regions. Here we report that a disruptive 213-base-pair (bp) inversion in the SCR gene (or its derivative haplotypes with deletions encompassing the entire SCR-A and a large portion of SRK-A) is found in 95% of European accessions, which contrasts with the genome-wide pattern of polymorphism in European A. thaliana16,17. Importantly, interspecific crossings using Arabidopsis halleri as a pollen donor reveal that some A. thaliana accessions, including Wei-1, retain the female SI reaction, suggesting that all female components including SRK are still functional. Moreover, when the 213-bp inversion in SCR was inverted and expressed in transgenic Wei-1 plants, the functional SCR restored the SI reaction. The inversion within SCR is the first mutation disrupting SI shown to be nearly fixed in geographically wide samples, and its prevalence is consistent with theoretical predictions regarding the evolutionary advantage of mutations in male components.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Darwin, C. The Effects of Cross and Self Fertilisation in the Vegetable Kingdom (J. Murray, London, 1876)
Stebbins, G. L. Flowering Plants: Evolution Above the Species Level (Harvard Univ. Press, 1974)
de Nettancourt, D. Incompatibility and Incongruity in Wild and Cultivated Plants 2nd edn (Springer, 2001)
Takayama, S. & Isogai, A. Self-incompatibility in plants. Annu. Rev. Plant Biol. 56, 467–489 (2005)
Busch, J. W. & Schoen, D. J. The evolution of self-incompatibility when mates are limiting. Trends Plant Sci. 13, 128–136 (2008)
Uyenoyama, M. K., Zhang, Y. & Newbigin, E. On the origin of self-incompatibility haplotypes: transition through self-compatible intermediates. Genetics 157, 1805–1817 (2001)
Kusaba, M. et al. Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana . Plant Cell 13, 627–643 (2001)
Nasrallah, M. E., Liu, P. & Nasrallah, J. B. Generation of self-incompatible Arabidopsis thaliana by transfer of two S locus genes from A. lyrata . Science 297, 247–249 (2002)
Sherman-Broyles, S. et al. S locus genes and the evolution of self-fertility in Arabidopsis thaliana . Plant Cell 19, 94–106 (2007)
Tang, C. et al. The evolution of selfing in Arabidopsis thaliana . Science 317, 1070–1072 (2007)
Shimizu, K. K., Shimizu-Inatsugi, R., Tsuchimatsu, T. & Purugganan, M. D. Independent origins of self-compatibility in Arabidopsis thaliana . Mol. Ecol. 17, 704–714 (2008)
Boggs, N. A., Nasrallah, J. B. & Nasrallah, M. E. Independent S-locus mutations caused self-fertility in Arabidopsis thaliana . PLoS Genet. 5, e1000426 (2009)
Liu, P., Sherman-Broyles, S., Nasrallah, M. E. & Nasrallah, J. B. A cryptic modifier causing transient self-incompatibility in Arabidopsis thaliana . Curr. Biol. 17, 734–740 (2007)
Igic, B., Lande, R. & Kohn, J. R. Loss of self-incompatibility and its evolutionary consequences. Int. J. Plant Sci. 169, 93–104 (2008)
Mable, B. K. Genetic causes and consequences of the breakdown of self-incompatibility: case studies in the Brassicaceae. Genet. Res. 90, 47–60 (2008)
Sharbel, T. F., Haubold, B. & Mitchell-Olds, T. Genetic isolation by distance in Arabidopsis thaliana: biogeography and postglacial colonization of Europe. Mol. Ecol. 9, 2109–2118 (2000)
François, O., Blum, M. G. B., Jakobsson, M. & Rosenberg, N. A. Demographic history of European populations of Arabidopsis thaliana . PLoS Genet. 4, e1000075 (2008)
Goodwillie, C., Kalisz, S. & Eckert, C. G. The evolutionary enigma of mixed mating systems in plants: occurrence, theoretical explanations, and empirical evidence. Annu. Rev. Ecol. Evol. Syst. 36, 47–79 (2005)
Fisher, R. A. Average excess and average effect of a gene substitution. Ann. Eugen. 11, 53–63 (1941)
Schopfer, C. R., Nasrallah, M. E. & Nasrallah, J. B. The male determinant of self-incompatibility in Brassica . Science 286, 1697–1700 (1999)
Takasaki, T. et al. The S receptor kinase determines self-incompatibility in Brassica stigma. Nature 403, 913–916 (2000)
Takayama, S. et al. Direct ligand–receptor complex interaction controls Brassica self-incompatibility. Nature 413, 534–538 (2001)
Bechsgaard, J. S., Castric, V., Charlesworth, D., Vekemans, X. & Schierup, M. H. The transition to self-compatibility in Arabidopsis thaliana and evolution within S-haplotypes over 10 Myr. Mol. Biol. Evol. 23, 1741–1750 (2006)
Abbott, R. J. & Gomes, M. F. Population genetic structure and outcrossing rate of Arabidopsis thaliana (L.) Heynh. Heredity 62, 411–418 (1989)
Kärkkäinen, K. et al. Genetic basis of inbreeding depression in Arabis petraea . Evolution 53, 1354–1365 (1999)
Okamoto, S. et al. Self-compatibility in Brassica napus is caused by independent mutations in S-locus genes. Plant J. 50, 391–400 (2007)
Shiba, H., Hinata, K., Suzuki, A. & Isogai, A. Breakdown of self-incompatibility in Brassica by the antisense RNA of SLG gene. Proc. Japan Acad. Ser. B 71, 81–83 (1995)
Guo, Y.-L. et al. Recent speciation of Capsella rubella from Capsella grandiflora, associated with loss of self-incompatibility and an extreme bottleneck. Proc. Natl Acad. Sci. USA 106, 5246–5251 (2009)
Baker, H. G. Self-compatibility and establishment after ‘long-distance’ dispersal. Evolution 9, 347–349 (1955)
Goring, D. R., Glavin, T. L., Schafer, U. & Rothstein, S. J. An S receptor kinase gene in self-compatible Brassica napus has a 1-bp deletion. Plant Cell 5, 531–539 (1993)
Nou, I. S., Watanabe, M., Isogai, A. & Hinata, K. Comparison of S-alleles and S-glycoproteins between two wild populations of Brassica campestris in Turkey and Japan. Sex. Plant Reprod. 6, 79–86 (1993)
Shimizu, K. K. Ecology meets molecular genetics in Arabidopsis . Popul. Ecol. 44, 221–233 (2002)
Smyth, D. R., Bowman, J. L. & Meyerowitz, E. M. Early flower development in Arabidopsis . Plant Cell 2, 755–767 (1990)
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999)
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana . Plant J. 16, 735–743 (1998)
Park, J. I. et al. Molecular characterization of two anther-specific genes encoding putative RNA-binding proteins, AtRBP45s, in Arabidopsis thaliana . Genes Genet. Syst. 81, 355–359 (2006)
Acknowledgements
We thank P. Awadalla, C. Bustamante, A. Caicedo, G. Coop, U. Grossniklaus, T.-h. Kao, C. Mays, R. Moore, K. Olsen, M. Purugganan, J. Reininga, L. Rose, S. Ruzsa and W. Stephan for discussions or technical advice. This work was supported by grants from the University Research Priority Program in Systems Biology/Functional Genomics of the University of Zurich and from the Swiss National Science Foundation (SNF) to K.K.S., and by Grants-in-Aid for Special Research on Priority Areas to S.T., M.W. and K.K.S.; by a grant from the Ministry of Education, Culture, Sports, Science, and Technology of Japan (MEXT) and by a Grant-in-Aid for Creative Scientific Research to S.T.; by a Young Scientific Research (S) grant to M.W.; and by a grant from the Japan Society for the Promotion of Science (JSPS). P.P. is the recipient of a grant from the Volkswagen Foundation and a STIBET scholarship of the Deutscher Akademischer Austauschdienst (DAAD). T.T. and K.S. are recipients of a Research Fellowship for Young Scientists from JSPS.
Author information
Authors and Affiliations
Contributions
T.T., K.S., M.W. and K.K.S. conceived and designed the study; T.T., R.S.-I. and K.K.S. performed sequencing, genotyping and crossing experiments; T.T., P.P., T.S. and K.K.S. conducted molecular population genetic analysis; K.S., S.I., S.T. and M.W. conducted transgenic analysis; T.T., K.S., G.S., T.S., M.W. and K.K.S. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Notes 1-4, Supplementary Tables 1-5 and Supplementary Figures 1-11 with legends. (PDF 1704 kb)
Rights and permissions
About this article
Cite this article
Tsuchimatsu, T., Suwabe, K., Shimizu-Inatsugi, R. et al. Evolution of self-compatibility in Arabidopsis by a mutation in the male specificity gene. Nature 464, 1342–1346 (2010). https://doi.org/10.1038/nature08927
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08927
This article is cited by
-
Dominance in self-compatibility between subgenomes of allopolyploid Arabidopsis kamchatica shown by transgenic restoration of self-incompatibility
Nature Communications (2023)
-
MLPK function is not required for self-incompatibility in the S29 haplotype of Brassica rapa L.
Plant Reproduction (2023)
-
Ancestral self-compatibility facilitates the establishment of allopolyploids in Brassicaceae
Plant Reproduction (2023)
-
Genetics Behind Sexual Incompatibility in Plants: How Much We Know and What More to Uncover?
Journal of Plant Growth Regulation (2023)
-
Stigma receptors control intraspecies and interspecies barriers in Brassicaceae
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.