Abstract
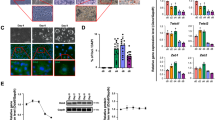
The worldwide epidemic of obesity has increased the urgency to develop a deeper understanding of physiological systems related to energy balance and energy storage, including the mechanisms controlling the development of fat cells (adipocytes). The differentiation of committed preadipocytes to adipocytes is controlled by PPARγ and several other transcription factors1, but the molecular basis for preadipocyte determination is not understood. Using a new method for the quantitative analysis of transcriptional components, we identified the zinc-finger protein Zfp423 as a factor enriched in preadipose versus non-preadipose fibroblasts. Ectopic expression of Zfp423 in non-adipogenic NIH 3T3 fibroblasts robustly activates expression of Pparg in undifferentiated cells and permits cells to undergo adipocyte differentiation under permissive conditions. Short hairpin RNA (shRNA)-mediated reduction of Zfp423 expression in 3T3-L1 cells blunts preadipocyte Pparg expression and diminishes the ability of these cells to differentiate. Furthermore, both brown and white adipocyte differentiation is markedly impaired in Zfp423-deficient mouse embryos. Zfp423 regulates Pparg expression, in part, through amplification of the BMP signalling pathway, an effect dependent on the SMAD-binding capacity of Zfp423. This study identifies Zfp423 as a transcriptional regulator of preadipocyte determination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Complete microarray data is available at Gene Expression Omnibus under accession GSE19732 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE19732).
References
Farmer, S. R. Transcriptional control of adipocyte formation. Cell Metab. 4, 263–273 (2006)
Green, H. & Kehinde, O. An established preadipose cell line and its differentiation in culture. II. Factors affecting the adipose conversion. Cell 5, 19–27 (1975)
Rodeheffer, M. S., Birsoy, K. & Friedman, J. M. Identification of white adipocyte progenitor cells in vivo . Cell 135, 240–249 (2008)
Tang, W. et al. White fat progenitor cells reside in the adipose vasculature. Science 322, 583–586 (2008)
Green, H. & Kehinde, O. Spontaneous heritable changes leading to increased adipose conversion in 3T3 cells. Cell 7, 105–113 (1976)
Todaro, G. J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963)
Gray, P. A. et al. Mouse brain organization revealed through direct genome-scale TF expression analysis. Science 306, 2255–2257 (2004)
Barak, Y. et al. PPARγ is required for placental, cardiac, and adipose tissue development. Mol. Cell 4, 585–595 (1999)
Kubota, N. et al. PPARγ mediates high-fat diet-induced adipocyte hypertrophy and insulin resistance. Mol. Cell 4, 597–609 (1999)
Rosen, E. D. et al. PPARγ is required for the differentiation of adipose tissue in vivo and in vitro . Mol. Cell 4, 611–617 (1999)
Tsai, R. Y. & Reed, R. R. Cloning and functional characterization of Roaz, a zinc finger protein that interacts with O/E-1 to regulate gene expression: implications for olfactory neuronal development. J. Neurosci. 17, 4159–4169 (1997)
Kajimura, S. et al. Initiation of myoblast to brown fat switch by a PRDM16–C/EBP-β transcriptional complex. Nature 460, 1154–1158 (2009)
Seale, P. et al. PRDM16 controls a brown fat/skeletal muscle switch. Nature 454, 961–967 (2008)
Cheng, L. E., Zhang, J. & Reed, R. R. The transcription factor Zfp423/OAZ is required for cerebellar development and CNS midline patterning. Dev. Biol. 307, 43–52 (2007)
Bowers, R. R., Kim, J. W., Otto, T. C. & Lane, M. D. Stable stem cell commitment to the adipocyte lineage by inhibition of DNA methylation: role of the BMP-4 gene. Proc. Natl Acad. Sci. USA 103, 13022–13027 (2006)
Hata, K. et al. Differential roles of Smad1 and p38 kinase in regulation of peroxisome proliferator-activating receptor γ during bone morphogenetic protein 2-induced adipogenesis. Mol. Biol. Cell 14, 545–555 (2003)
Jin, W. et al. Schnurri-2 controls BMP-dependent adipogenesis via interaction with Smad proteins. Dev. Cell 10, 461–471 (2006)
Tang, Q. Q., Otto, T. C. & Lane, M. D. Commitment of C3H10T1/2 pluripotent stem cells to the adipocyte lineage. Proc. Natl Acad. Sci. USA 101, 9607–9611 (2004)
Tseng, Y. H. et al. New role of bone morphogenetic protein 7 in brown adipogenesis and energy expenditure. Nature 454, 1000–1004 (2008)
Hata, A. et al. OAZ uses distinct DNA- and protein-binding zinc fingers in separate BMP-Smad and Olf signaling pathways. Cell 100, 229–240 (2000)
Warming, S., Rachel, R. A., Jenkins, N. A. & Copeland, N. G. Zfp423 is required for normal cerebellar development. Mol. Cell. Biol. 26, 6913–6922 (2006)
Alcaraz, W. A. et al. Zfp423 controls proliferation and differentiation of neural precursors in cerebellar vermis formation. Proc. Natl Acad. Sci. USA 103, 19424–19429 (2006)
Cheng, L. E. & Reed, R. R. Zfp423/OAZ participates in a developmental switch during olfactory neurogenesis. Neuron 54, 547–557 (2007)
Jimenez, M. A., Akerblad, P., Sigvardsson, M. & Rosen, E. D. Critical role for Ebf1 and Ebf2 in the adipogenic transcriptional cascade. Mol. Cell. Biol. 27, 743–757 (2007)
Lockhart, D. J. et al. Expression monitoring by hybridization to high-density oligonucleotide arrays. Nature Biotechnol. 14, 1675–1680 (1996)
Li, C. & Wong, W. H. Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc. Natl Acad. Sci. USA 98, 31–36 (2001)
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002)
Acknowledgements
We are grateful to S. Kleiner and S. Kajimura for critical reading of the manuscript and to all members of the Spiegelman laboratory for useful discussions. We thank B. Wagner for technical assistance in using robotic liquid handlers, B. Seed’s laboratory for help with high-through query of Primer bank, and J. Brestelli for performing high-throughput queries of the Primer 3 program. We are also grateful to D. Bernlohr for the FABP4 antiserum. R.K.G. is supported by the Ruth Kirstein NRSA (F32 DK079507-01), Z.A. is supported by K08 HL79172-01 (NHLBI) and the Smith Family Foundation Grant, P.S. is supported by NIH DK081605, and the research described in this study was supported by NIH DK31405 to B.M.S. and by NIDCD R01DC008295 to R.R.R.
Author Contributions R.K.G. and B.M.S. conceived and designed the experiments. R.K.G., Z.A., P.S., R.J.M., L.Y. and H.M.C. performed experiments. All authors analysed the data. Y.A.R., H.K. and R.R.R. provided reagents and samples, and R.K.G. and B.M.S. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-14 with legends, and Supplementary Tables 1-2. (PDF 1177 kb)
Supplementary Data
Supplementary Data Set 1: Raw Cycle Threshold data from Quanttrx assays (XLS 479 kb)
Supplementary Data
Supplementary Data Set 2: Primer sequences used in the Quanttrx Assays (XLS 522 kb)
Rights and permissions
About this article
Cite this article
Gupta, R., Arany, Z., Seale, P. et al. Transcriptional control of preadipocyte determination by Zfp423. Nature 464, 619–623 (2010). https://doi.org/10.1038/nature08816
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08816
This article is cited by
-
Anti-obesity effect of the bacterial product nisin in an NIH Swiss mouse model
Lipids in Health and Disease (2023)
-
Characteristic and fate determination of adipose precursors during adipose tissue remodeling
Cell Regeneration (2023)
-
Stem cell-based strategies and challenges for production of cultivated meat
Nature Food (2023)
-
Deciphering the interaction between Twist1 and PPARγ during adipocyte differentiation
Cell Death & Disease (2023)
-
Promotion effect of TGF-β-Zfp423-ApoD pathway on lip sensory recovery after nerve sacrifice caused by nerve collateral compensation
International Journal of Oral Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.