Abstract
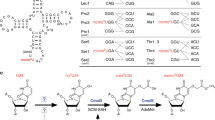
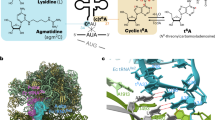
Maturation of precursor transfer RNA (pre-tRNA) includes excision of the 5′ leader and 3′ trailer sequences, removal of introns and addition of the CCA terminus1,2,3. Nucleotide modifications are incorporated at different stages of tRNA processing, after the RNA molecule adopts the proper conformation. In bacteria, tRNAIle2 lysidine synthetase (TilS) modifies cytidine into lysidine (L; 2-lysyl-cytidine) at the first anticodon of tRNAIle2 (refs 4–9). This modification switches tRNAIle2 from a methionine-specific to an isoleucine-specific tRNA9. However, the aminoacylation of tRNAIle2 by methionyl-tRNA synthetase (MetRS), before the modification by TilS, might lead to the misincorporation of methionine in response to isoleucine codons. The mechanism used by bacteria to avoid this pitfall is unknown. Here we show that the TilS enzyme specifically recognizes and modifies tRNAIle2 in its precursor form, thereby avoiding translation errors. We identified the lysidine modification in pre-tRNAIle2 isolated from RNase-E-deficient Escherichia coli and did not detect mature tRNAIle2 lacking this modification. Our kinetic analyses revealed that TilS can modify both types of RNA molecule with comparable efficiencies. X-ray crystallography and mutational analyses revealed that TilS specifically recognizes the entire L-shape structure in pre-tRNAIle2 through extensive interactions coupled with sequential domain movements. Our results demonstrate how TilS prevents the recognition of tRNAIle2 by MetRS and achieves high specificity for its substrate. These two key points form the basis for maintaining the fidelity of isoleucine codon translation in bacteria. Our findings also provide a rationale for the necessity of incorporating specific modifications at the precursor level during tRNA biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bjork, G. R. in tRNA: Structure, Biosynthesis and Function (eds Söll, D. & RajBhandary, U. L.) 165–206 (ASM Press, 1995)
Hendrickson, T. & Schimmel, P. in Translation Mechanisms (eds Lapointe, J. & Brakier-Gingras, L.) 33–64 (Landes Bioscience, 2003)
Nakanishi, K. & Nureki, O. Recent progress of structural biology of tRNA processing and modification. Mol. Cells 19, 157–166 (2005)
Harada, F. & Nishimura, S. Purification and characterization of AUA specific isoleucine transfer ribonucleic-acid from Escherichia coli B. Biochemistry 13, 300–307 (1974)
Matsugi, J., Murao, K. & Ishikura, H. Characterization of a B. subtilis minor isoleucine tRNA deduced from tDNA having a methionine anticodon CAT. J. Biochem. 119, 811–816 (1996)
Muramatsu, T. et al. A novel lysine-substituted nucleoside in the 1st position of the anticodon of minor isoleucine transfer-RNA from Escherichia coli . J. Biol. Chem. 263, 9261–9267 (1988)
Weber, F., Dietrich, A., Weil, J. H. & Marechal-Drouard, L. A potato mitochondrial isoleucine tRNA is coded for by a mitochondrial gene possessing a methionine anticodon. Nucleic Acids Res. 18, 5027–5030 (1990)
Soma, A. et al. An RNA-modifying enzyme that governs both the codon and amino acid specificities of isoleucine tRNA. Mol. Cell 12, 689–698 (2003)
Muramatsu, T. et al. Codon and amino-acid specificities of a transfer-RNA are both converted by a single post-transcriptional modification. Nature 336, 179–181 (1988)
Sakano, H. & Shimura, Y. Characterization and in vitro processing of transfer RNA precursors accumulated in a temperature sensitive mutant of Escherichia coli . J. Mol. Biol. 123, 287–326 (1978)
Etcheverry, T., Colby, D. & Guthrie, C. Precursor to a minor species of yeast tRNASer contains an intervening sequence. Cell 18, 11–26 (1979)
Nishikura, K. & De Robertis, E. M. RNA processing in micro injected Xenopus laevis oocytes. Sequential addition of base modifications in a spliced transfer RNA. J. Mol. Biol. 145, 405–420 (1981)
Nakanishi, K., Ogiso, Y., Nakama, T., Fukai, S. & Nureki, O. Structural basis for anticodon recognition by methionyl-tRNA synthetase. Nature Struct. Mol. Biol. 12, 931–932 (2005)
Kimata, K., Tanaka, Y., Inada, T. & Aiba, H. Expression of the glucose transporter gene, ptsG, is regulated at the mRNA degradation step in response to glycolytic flux in Escherichia coli . EMBO J. 20, 3587–3595 (2001)
Kaneko, T. et al. Wobble modification differences and subcellular localization of tRNAs in Leishmania tarentolae: implication for tRNA sorting mechanism. EMBO J. 22, 657–667 (2003)
Suzuki, T. & Suzuki, T. Chaplet column chromatography: isolation of a large set of individual RNAs in a single step. Methods Enzymol. 425, 231–239 (2007)
Suzuki, T., Ikeuchi, Y., Noma, A., Suzuki, T. & Sakaguchi, Y. Mass spectrometric identification and characterization of RNA-modifying enzymes. Methods Enzymol. 425, 212–229 (2007)
Sugiura, I. et al. The 2.0 Angstrom crystal structure of Thermus thermophilus methionyl-tRNA synthetase reveals two RNA-binding modules. Structure 8, 197–208 (2000)
Rould, M. A., Perona, J. J. & Steitz, T. A. Structural basis of anticodon loop recognition by glutaminyl-tRNA synthetase. Nature 352, 213–218 (1991)
Numata, T., Ikeuchi, Y., Fukai, S., Suzuki, T. & Nureki, O. Snapshots of tRNA sulphuration via an adenylated intermediate. Nature 442, 419–424 (2006)
Losey, H. C., Ruthenburg, A. J. & Verdine, G. L. Crystal structure of Staphylococcus aureus tRNA adenosine deaminase TadA in complex with RNA. Nature Struct. Mol. Biol. 13, 153–159 (2006)
Xie, W., Liu, X. J. & Huang, R. H. Chemical trapping and crystal structure of a catalytic tRNA guanine transglycosylase covalent intermediate. Nature Struct. Biol. 10, 781–788 (2003)
Nagaswamy, U., Voss, N., Zhang, Z. D. & Fox, G. E. Database of non-canonical base pairs found in known RNA structures. Nucleic Acids Res. 28, 375–376 (2000)
Shi, H. J. & Moore, P. B. The crystal structure of yeast phenylalanine tRNA at 1.93 Angstrom resolution: a classic structure revisited. RNA 6, 1091–1105 (2000)
Ikeuchi, Y. et al. Molecular mechanism of lysidine synthesis that determines tRNA identity and codon recognition. Mol. Cell 19, 235–246 (2005)
Hoang, C. et al. Crystal structure of pseudouridine synthase RluA: Indirect sequence readout through protein-induced RNA structure. Mol. Cell 24, 535–545 (2006)
Nakanishi, K. et al. Structural basis for lysidine formation by ATP pyrophosphatase accompanied by a lysine-specific loop and a tRNA-recognition domain. Proc. Natl Acad. Sci. USA 102, 7487–7492 (2005)
Jordan, S. R. & Pabo, C. O. Structure of the lambda complex at 2.5 Å resolution: details of the repressor-operator interactions. Science 242, 893–899 (1988)
Chomczynski, P. & Sacchi, N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162, 156–159 (1987)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Weeks, C. M. & Miller, R. The design and implementation of SnB version 2.0. J. Appl. Cryst. 32, 120–124 (1999)
de La Fortelle, E., Irwin, J. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for the multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Abrahams, J. P. & Leslie, A. G. W. Methods used in the structure determination of bovine mitochondrial F-1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
Collaborative Computational Project, 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron-density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr D 58, 1948–1954 (2002)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Vagin, A. & Teplyakov, A. An approach to multi-copy search in molecular replacement. Acta Crystallogr. D 56, 1622–1624 (2000)
Emsley, P. C. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Acknowledgements
We thank the beamline staff at BL41XU of SPring-8 (Harima, Japan) and NW12 of PF-AR (Tsukuba, Japan) for help during data collection. We thank H. Aiba for providing the temperature-sensitive E. coli strain of RNase E, A. Soma and Y. Sekine for discussions, and T. Suzuki and Y. Sakaguchi for support with mass spectrometry. This work was supported by a SORST program grant from Japan Science and Technology to O.N., by a grant for the National Project on Protein Structural and Functional Analyses from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) to O.N. and T.S., by grants from MEXT to R.I. and O.N., and by Mitsubishi Foundation and Kurata Memorial Hitachi Science and Technology Foundation grants to O.N. L.B. and K.N. were supported by a postdoctoral fellowship for foreign researchers and a young scientist fellowship, respectively, from the Japan Society for the Promotion of Science.
Author Contributions K.N. performed purification, crystallization and structure determination of the complex, and biochemical analyses. L.B. performed purification, crystallization and structure determination of the ASB domain, and biochemical analyses. S.K. purified the native tRNAs and performed the mass spectrometry analysis under the supervision of T.S. R.I. performed the molecular dynamics analysis and assisted with the structural determination. O.N. assisted with the structure determination. All authors discussed the results and commented on the manuscript. O.N. supervised all the work.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary Methods, Supplementary References, Supplementary Tables 1- 4 and Supplementary Figures 1-12 with Legends. (PDF 10766 kb)
Rights and permissions
About this article
Cite this article
Nakanishi, K., Bonnefond, L., Kimura, S. et al. Structural basis for translational fidelity ensured by transfer RNA lysidine synthetase. Nature 461, 1144–1148 (2009). https://doi.org/10.1038/nature08474
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08474
This article is cited by
-
The occurrence order and cross-talk of different tRNA modifications
Science China Life Sciences (2021)
-
Analysis of the complete plastid genome of the unicellular red alga Porphyridium purpureum
Journal of Plant Research (2014)
-
Identification of highly-disrupted tRNA genes in nuclear genome of the red alga, Cyanidioschyzon merolae 10D
Scientific Reports (2013)
-
Evolution in an oncogenic bacterial species with extreme genome plasticity: Helicobacter pyloriEast Asian genomes
BMC Microbiology (2011)
-
Biogenesis of 2-agmatinylcytidine catalyzed by the dual protein and RNA kinase TiaS
Nature Structural & Molecular Biology (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.