Abstract
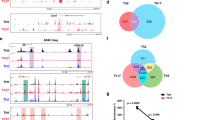
Activator protein 1 (AP-1, also known as JUN) transcription factors are dimers of JUN, FOS, MAF and activating transcription factor (ATF) family proteins characterized by basic region and leucine zipper domains1. Many AP-1 proteins contain defined transcriptional activation domains, but BATF and the closely related BATF3 (refs 2, 3) contain only a basic region and leucine zipper, and are considered to be inhibitors of AP-1 activity3,4,5,6,7,8. Here we show that Batf is required for the differentiation of IL17-producing T helper (TH17) cells9. TH17 cells comprise a CD4+ T-cell subset that coordinates inflammatory responses in host defence but is pathogenic in autoimmunity10,11,12,13. Batf-/- mice have normal TH1 and TH2 differentiation, but show a defect in TH17 differentiation, and are resistant to experimental autoimmune encephalomyelitis. Batf-/- T cells fail to induce known factors required for TH17 differentiation, such as RORγt11 (encoded by Rorc) and the cytokine IL21 (refs 14–17). Neither the addition of IL21 nor the overexpression of RORγt fully restores IL17 production in Batf-/- T cells. The Il17 promoter is BATF-responsive, and after TH17 differentiation, BATF binds conserved intergenic elements in the Il17a–Il17f locus and to the Il17, Il21 and Il22 (ref. 18) promoters. These results demonstrate that the AP-1 protein BATF has a critical role in TH17 differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Data deposits
Microarray data are available at Array Express (http://www.ebi.ac.uk/arrayexpress/) under the accession numbers E-MEXP-1518, E-MEXP-2152 and E-MEXP-2153.
References
Wagner, E. F. & Eferl, R. Fos/AP-1 proteins in bone and the immune system. Immunol. Rev. 208, 126–140 (2005)
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8α+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008)
Iacobelli, M., Wachsman, W. & McGuire, K. L. Repression of IL-2 promoter activity by the novel basic leucine zipper p21SNFT protein. J. Immunol. 165, 860–868 (2000)
Blank, V. Small Maf proteins in mammalian gene control: mere dimerization partners or dynamic transcriptional regulators? J. Mol. Biol. 376, 913–925 (2008)
Williams, K. L. et al. Characterization of murine BATF: a negative regulator of activator protein-1 activity in the thymus. Eur. J. Immunol. 31, 1620–1627 (2001)
Echlin, D. R., Tae, H. J., Mitin, N. & Taparowsky, E. J. B-ATF functions as a negative regulator of AP-1 mediated transcription and blocks cellular transformation by Ras and Fos. Oncogene 19, 1752–1763 (2000)
Dorsey, M. J. et al. B-ATF: a novel human bZIP protein that associates with members of the AP-1 transcription factor family. Oncogene 11, 2255–2265 (1995)
Thornton, T. M., Zullo, A. J., Williams, K. L. & Taparowsky, E. J. Direct manipulation of activator protein-1 controls thymocyte proliferation in vitro. Eur. J. Immunol. 36, 160–169 (2006)
Harrington, L. E. et al. Interleukin 17-producing CD4+ effector T cells develop via a lineage distinct from the T helper type 1 and 2 lineages. Nature Immunol. 6, 1123–1132 (2005)
Langrish, C. L. et al. IL-23 drives a pathogenic T cell population that induces autoimmune inflammation. J. Exp. Med. 201, 233–240 (2005)
Ivanov, I. I. et al. The orphan nuclear receptor RORγt directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126, 1121–1133 (2006)
Bettelli, E. et al. Reciprocal developmental pathways for the generation of pathogenic effector TH17 and regulatory T cells. Nature 441, 235–238 (2006)
Brustle, A. et al. The development of inflammatory TH17 cells requires interferon-regulatory factor 4. Nature Immunol. 8, 958–966 (2007)
Korn, T. et al. IL-21 initiates an alternative pathway to induce proinflammatory TH17 cells. Nature 448, 484–487 (2007)
Nurieva, R. et al. Essential autocrine regulation by IL-21 in the generation of inflammatory T cells. Nature 448, 480–483 (2007)
Wei, L., Laurence, A., Elias, K. M. & O’Shea, J. J. IL-21 is produced by Th17 cells and drives IL-17 production in a STAT3-dependent manner. J. Biol. Chem. 282, 34605–34610 (2007)
Zhou, L. et al. IL-6 programs TH17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nature Immunol. 8, 967–974 (2007)
Liang, S. C. et al. Interleukin (IL)-22 and IL-17 are coexpressed by Th17 cells and cooperatively enhance expression of antimicrobial peptides. J. Exp. Med. 203, 2271–2279 (2006)
Bower, K. E., Fritz, J. M. & McGuire, K. L. Transcriptional repression of MMP-1 by p21SNFT and reduced in vitro invasiveness of hepatocarcinoma cells. Oncogene 23, 8805–8814 (2004)
Hess, J., Angel, P. & Schorpp-Kistner, M. AP-1 subunits: quarrel and harmony among siblings. J. Cell Sci. 117, 5965–5973 (2004)
Williams, K. L. et al. BATF transgenic mice reveal a role for activator protein-1 in NKT cell development. J. Immunol. 170, 2417–2426 (2003)
Zhumabekov, T., Corbella, P., Tolaini, M. & Kioussis, D. Improved version of a human CD2 minigene based vector for T cell-specific expression in transgenic mice. J. Immunol. Methods 185, 133–140 (1995)
Haak, S. et al. IL-17A and IL-17F do not contribute vitally to autoimmune neuro-inflammation in mice. J. Clin. Invest. 119, 61–69 (2009)
Yang, X. O. et al. T helper 17 lineage differentiation is programmed by orphan nuclear receptors RORα and RORγ. Immunity 28, 29–39 (2008)
Veldhoen, M. et al. The aryl hydrocarbon receptor links TH17-cell-mediated autoimmunity to environmental toxins. Nature 453, 106–109 (2008)
Quintana, F. J. et al. Control of Treg and TH17 cell differentiation by the aryl hydrocarbon receptor. Nature 453, 65–71 (2008)
Ichiyama, K. et al. Foxp3 inhibits RORγt-mediated IL-17A mRNA transcription through direct interaction with RORγt. J. Biol. Chem. 283, 17003–17008 (2008)
Zhang, F., Meng, G. & Strober, W. Interactions among the transcription factors Runx1, RORγt and Foxp3 regulate the differentiation of interleukin 17-producing T cells. Nature Immunol. 9, 1297–1306 (2008)
Zhu, H. et al. Unexpected characteristics of the IFN-γ reporters in nontransformed T cells. J. Immunol. 167, 855–865 (2001)
Hertz, G. Z. & Stormo, G. D. Identifying DNA and protein patterns with statistically significant alignments of multiple sequences. Bioinformatics 15, 563–577 (1999)
Gorman, J. R. et al. The Igκ enhancer influences the ratio of Igκ versus Igλ B lymphocytes. Immunity 5, 241–252 (1996)
Ranganath, S. et al. GATA-3-dependent enhancer activity in IL-4 gene regulation. J. Immunol. 161, 3822–3826 (1998)
Zhumabekov, T., Corbella, P., Tolaini, M. & Kioussis, D. Improved version of a human CD2 minigene based vector for T cell-specific expression in transgenic mice. J. Immunol. Methods 185, 133–140 (1995)
Sun, Z. et al. Requirement for RORγ in thymocyte survival and lymphoid organ development. Science 288, 2369–2373 (2000)
Acknowledgements
We thank R. Lallone for anti-BATF antibody preparation, and B. Sleckman for Cre-expressing adenovirus. This work was supported by the Howard Hughes Medical Institute (K.M.M.), and grants from the National Institutes of Health HG00249 and training grant GM07200 (G.D.S.), AI035783 (C.T.W.), AR049293 (R.D.H.), and from Daiichi-Sankyo Co. Ltd (C.T.W.).
Author Contributions B.U.S. generated Batf-/- mice, designed and analysed the experiments, interpreted results and wrote the manuscript. K.H. constructed the targeting vector and probes, transgenic vector, and recombinant BATF. W.I. helped with retroviral expression experiments. W.-L.L. helped with reverse-strand reporter analysis. W.A.-E.S. helped with mouse generation. B.S. helped with EMSA analysis. G.S. and G.D.S. performed bioinformatics analysis for the BATF binding elements. J.S. and J.H.R. helped with EAE experiments. R.M., R.D.H. and C.T.W. performed ChIP experiments. T.L.M. and S.C. performed confocal microscopy for BATF. K.M.M. directed the study and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-14 with Legends, Supplementary Tables 1, 5-6, Supplementary Methods and Supplementary References. (PDF 1930 kb)
Supplementary Table 2
This file contains microarray data in Excel format accompanying Figure 3c. (XLS 27 kb)
Supplementary Table 3
This file contains microarray data in Excel format accompanying Supplementary Figure 9a. (XLS 1 kb)
Supplementary Table 4
This file contains Gene ChIP data in Excel format accompanying Supplementary Figure 9b. (XLS 1 kb)
Rights and permissions
About this article
Cite this article
Schraml, B., Hildner, K., Ise, W. et al. The AP-1 transcription factor Batf controls TH17 differentiation. Nature 460, 405–409 (2009). https://doi.org/10.1038/nature08114
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08114
This article is cited by
-
Mitochondrial and metabolic dysfunction of peripheral immune cells in multiple sclerosis
Journal of Neuroinflammation (2024)
-
Single cell transcriptomics shows that malaria promotes unique regulatory responses across multiple immune cell subsets
Nature Communications (2023)
-
Cardinal features of immune memory in innate lymphocytes
Nature Immunology (2023)
-
Markers and makers of NKT17 cells
Experimental & Molecular Medicine (2023)
-
The role of transcription factors in shaping regulatory T cell identity
Nature Reviews Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.