Abstract
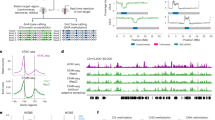
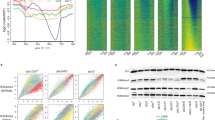
Cytosine DNA methylation is important in regulating gene expression and in silencing transposons and other repetitive sequences1,2. Recent genomic studies in Arabidopsis thaliana have revealed that many endogenous genes are methylated either within their promoters or within their transcribed regions, and that gene methylation is highly correlated with transcription levels3,4,5. However, plants have different types of methylation controlled by different genetic pathways, and detailed information on the methylation status of each cytosine in any given genome is lacking. To this end, we generated a map at single-base-pair resolution of methylated cytosines for Arabidopsis, by combining bisulphite treatment of genomic DNA with ultra-high-throughput sequencing using the Illumina 1G Genome Analyser and Solexa sequencing technology6. This approach, termed BS-Seq, unlike previous microarray-based methods, allows one to sensitively measure cytosine methylation on a genome-wide scale within specific sequence contexts. Here we describe methylation on previously inaccessible components of the genome and analyse the DNA methylation sequence composition and distribution. We also describe the effect of various DNA methylation mutants on genome-wide methylation patterns, and demonstrate that our newly developed library construction and computational methods can be applied to large genomes such as that of mouse.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Henderson, I. R. & Jacobsen, S. E. Epigenetic inheritance in plants. Nature 447, 418–424 (2007)
Goll, M. G. & Bestor, T. H. Eukaryotic cytosine methyltransferases. Annu. Rev. Biochem. 74, 481–514 (2005)
Zhang, X. et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell 126, 1189–1201 (2006)
Zilberman, D., Gehring, M., Tran, R. K., Ballinger, T. & Henikoff, S. Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nature Genet. 39, 61–69 (2007)
Vaughn, M. W. et al. Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol. 5, e174 (2007)
Bentley, D. R. Whole-genome re-sequencing. Curr. Opin. Genet. Dev. 16, 545–552 (2006)
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl Acad. Sci. USA 89, 1827–1831 (1992)
Ngernprasirtsiri, J., Kobayashi, H. & Akazawa, T. DNA methylation as a mechanism of transcriptional regulation in nonphotosynthetic plastids in plant cells. Proc. Natl Acad. Sci. USA 85, 4750–4754 (1988)
Tran, R. K. et al. DNA methylation profiling identifies CG methylation clusters in Arabidopsis genes. Curr. Biol. 15, 154–159 (2005)
Gruenbaum, Y., Naveh-Many, T., Cedar, H. & Razin, A. Sequence specificity of methylation in higher plant DNA. Nature 292, 860–862 (1981)
Meyer, P., Niedenhof, I. & ten Lohuis, M. Evidence for cytosine methylation of non-symmetrical sequences in transgenic Petunia hybrida. EMBO J. 13, 2084–2088 (1994)
Cao, X. & Jacobsen, S. E. Locus-specific control of asymmetric and CpNpG methylation by the DRM and CMT3 methyltransferase genes. Proc. Natl Acad. Sci. USA 99 (Suppl 4). 16491–16498 (2002)
Dieguez, M. J., Vaucheret, H., Paszkowski, J. & Mittelsten Scheid, O. Cytosine methylation at CG and CNG sites is not a prerequisite for the initiation of transcriptional gene silencing in plants, but it is required for its maintenance. Mol. Gen. Genet. 259, 207–215 (1998)
Ramsahoye, B. H. et al. Non-CpG methylation is prevalent in embryonic stem cells and may be mediated by DNA methyltransferase 3a. Proc. Natl Acad. Sci. USA 97, 5237–5242 (2000)
Jia, D., Jurkowska, R. Z., Zhang, X., Jeltsch, A. & Cheng, X. Structure of Dnmt3a bound to Dnmt3L suggests a model for de novo DNA methylation. Nature 449, 248–251 (2007)
Cao, X. et al. Conserved plant genes with similarity to mammalian de novo DNA methyltransferases. Proc. Natl Acad. Sci. USA 97, 4979–4984 (2000)
Bers, E. P., Singh, N. P., Pardonen, V. A., Lutova, L. A. & Zalensky, A. O. Nucleosomal structure and histone H1 subfractional composition of pea (Pisum sativum) root nodules, radicles and callus chromatin. Plant Mol. Biol. 20, 1089–1096 (1992)
Vershinin, A. V. & Heslop-Harrison, J. S. Comparative analysis of the nucleosomal structure of rye, wheat and their relatives. Plant Mol. Biol. 36, 149–161 (1998)
Fulnecek, J., Matyasek, R., Kovarik, A. & Bezdek, M. Mapping of 5-methylcytosine residues in Nicotiana tabacum 5S rRNA genes by genomic sequencing. Mol. Gen. Genet. 259, 133–141 (1998)
Fan, Y. et al. Histone H1 depletion in mammals alters global chromatin structure but causes specific changes in gene regulation. Cell 123, 1199–1212 (2005)
Zhang, X. & Jacobsen, S. E. Genetic analyses of DNA methyltransferases in Arabidopsis thaliana. Cold Spring Harb. Symp. Quant. Biol. 71, 439–447 (2006)
Henderson, I. R. et al. Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nature Genet. 38, 721–725 (2006)
Bostick, M. et al. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007)
Sharif, J. et al. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450, 908–912 (2007)
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005)
Rajagopalan, R., Vaucheret, H., Trejo, J. & Bartel, D. P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 20, 3407–3425 (2006)
Acknowledgements
We thank Y. Bernatavichute for nuclear DNA isolation protocols, A. Clarke for providing embryonic stem cell DNA, A. Girard and G. Hannon for providing mouse germ cell DNA, J. Hetzel for technical assistance, and C. F. Li for assistance with rDNA annotation. S.F. is a Howard Hughes Medical Institute Fellow of the Life Science Research Foundation. X.Z. was supported by a fellowship from the Jonsson Cancer Center Foundation. S.E.J. is an investigator of the Howard Hughes Medical Institute. This work was supported in part by grants from the NSF Plant Genome Research Program and the NIH, and some aspects of the work were performed in the UCLA DNA Microarray Facility.
Author Contributions S.J.C. developed computational methods for mapping and base-calling. S.F. designed and created DNA libraries and performed all molecular biology experiments. S.F., Z.C., B.M. and S.F.N. sequenced the libraries. M.P., S.J.C., S.F. and S.E.J. analysed data. S.E.J. and M.P. designed and directed the study. X.Z., C.D.H. and S.P. assisted in the design of experiments. S.F. and S.J.C. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
C. Haudenschild is an employee of Illumina Inc., the manufacturer of the high throughput sequencing system used in this work.
Supplementary information
TITLE
This file contains Supplementary Figures 1-18 with Legends and Supplementary Methods. (PDF 5784 kb)
TITLE
This file contains Supplementary Table 1. (XLS 17 kb)
TITLE
This file contains Supplementary Table 2. (XLS 18 kb)
TITLE
This file contains Supplementary Table 3. (XLS 978 kb)
TITLE
This file contains Supplementary Table 4. (XLS 807 kb)
TITLE
This file contains Supplementary Table 5. (XLS 3593 kb)
TITLE
This file contains Supplementary Table 6. (XLS 9 kb)
TITLE
This file contains Supplementary Table 7. (XLS 14 kb)
Rights and permissions
About this article
Cite this article
Cokus, S., Feng, S., Zhang, X. et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219 (2008). https://doi.org/10.1038/nature06745
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06745
This article is cited by
-
Genome-wide characterization of DNA methyltransferase family genes implies GhDMT6 improving tolerance of salt and drought on cotton
BMC Plant Biology (2024)
-
Epigenetics: Toward improving crop disease resistance and agronomic characteristics
Plant Biotechnology Reports (2024)
-
Global effects of identity and aging on the human sperm methylome
Clinical Epigenetics (2023)
-
Single-cell DNA methylation sequencing by combinatorial indexing and enzymatic DNA methylation conversion
Cell & Bioscience (2023)
-
MethylC-analyzer: a comprehensive downstream pipeline for the analysis of genome-wide DNA methylation
Botanical Studies (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.