Abstract
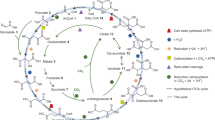
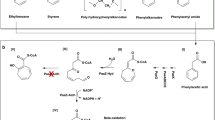
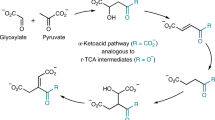
The human pathogenic bacterium Clostridium difficile thrives by the fermentation of l-leucine to ammonia, CO2, 3-methylbutanoate and 4-methylpentanoate under anaerobic conditions1. The reductive branch to 4-methylpentanoate proceeds by means of the dehydration of (R)-2-hydroxy-4-methylpentanoyl-CoA to 4-methylpent-2-enoyl-CoA, which is chemically the most demanding step. Ketyl radicals have been proposed2 to mediate this reaction catalysed by an iron–sulphur-cluster-containing dehydratase, which requires activation by ATP-dependent electron transfer from a second iron–sulphur protein functionally similar to the iron protein of nitrogenase. Here we identify a kinetically competent product-related allylic ketyl radical bound to the enzyme by electron paramagnetic resonance spectroscopy employing isotope-labelled (R)-2-hydroxy-4-methylpentanoyl-CoA species. We also found that the enzyme generated the stabilized pentadienoyl ketyl radical from the substrate analogue 2-hydroxypent-4-enoyl-CoA, supporting the proposed mechanism. Our results imply that also other 2-hydroxyacyl-CoA dehydratases3 and the related benzoyl-CoA reductases4—present in anaerobically living bacteria—employ ketyl radical intermediates. The absence of radical generators such as coenzyme B12, S-adenosylmethionine or oxygen makes these enzymes unprecedented in biochemistry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Buckel, W. in Biology of the Prokaryotes (eds Lengeler, J. W., Drews, G. & Schlegel, H. G.) 278–326 (Thieme, Stuttgart, 1999)
Buckel, W. & Keese, R. One electron redox reactions of CoASH esters in anaerobic bacteria. A mechanistic proposal. Angew. Chem. Int. Edn Engl. 34, 1502–1506 (1995)
Buckel, W., Hetzel, M. & Kim, J. ATP-driven electron transfer in enzymatic radical reactions. Curr. Opin. Chem. Biol. 8, 462–467 (2004)
Boll, M. & Fuchs, G. Unusual reactions involved in anaerobic metabolism of phenolic compounds. Biol. Chem. 386, 989–997 (2005)
Seebach, D. Methods of reactivity Umpolung. Angew. Chem. Int. Edn Engl. 18, 239–258 (1979)
Brückner, R. Reaktionsmechanismen (Spektrum Akademischer, Heidelberg, 2003)
Buckel, W. & Golding, B. T. Radical enzymes in anaerobes. Annu. Rev. Microbiol. 60, 27–49 (2006)
Smith, D. M., Buckel, W. & Zipse, H. Deprotonation of enoxy radicals: theoretical validation of a 50-year-old mechanistic proposal. Angew. Chem. Int. Edn Engl. 42, 1867–1870 (2003)
Sebaihia, M. & Thomson, N. R. Colonic irritation. Nature Rev. Microbiol. 4, 882–883 (2006)
Reineke, J. et al. Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446, 415–419 (2007)
Selmer, T. & Andrei, P. I. p-Hydroxyphenylacetate decarboxylase from Clostridium difficile. A novel glycyl radical enzyme catalysing the formation of p-cresol. Eur. J. Biochem. 268, 1363–1372 (2001)
Sebaihia, M. et al. The multidrug-resistant human pathogen Clostridium difficile has a highly mobile, mosaic genome. Nature Genet. 38, 779–786 (2006)
Barker, H. A. Amino acid degradation by anaerobic bacteria. Annu. Rev. Biochem. 50, 23–40 (1981)
Elsden, S. R. & Hilton, M. G. Volatile acid production from threonine, valine, leucine and isoleucine by clostridia. Arch. Microbiol. 117, 165–172 (1978)
Kim, J., Darley, D., Selmer, T. & Buckel, W. Characterization of (R)-2-hydroxyisocaproate dehydrogenase and a family III coenzyme A transferase involved in reduction of l-leucine to isocaproate by Clostridium difficile. Appl. Environ. Microbiol. 72, 6062–6069 (2006)
Kim, J., Darley, D. & Buckel, W. 2-Hydroxyisocaproyl-CoA dehydratase and its activator from Clostridium difficile. FEBS J. 272, 550–561 (2005)
Hans, M., Buckel, W. & Bill, E. The iron–sulfur clusters in 2-hydroxyglutaryl-CoA dehydratase from Acidaminococcus fermentans. Biochemical and spectroscopic investigations. Eur. J. Biochem. 267, 7082–7093 (2000)
Weil, J. A. & Bolton, J. R. Electron Paramagnetic Resonance: Elementary Theory and Practical Applications 2nd edn (Wiley, Bognor Regis, 2007)
Wu, W. et al. Lysine 2,3-aminomutase and trans-4,5-dehydrolysine: characterization of an allylic analogue of a substrate-based radical in the catalytic mechanism. Biochemistry 39, 9561–9570 (2000)
Magnusson, O. T., Reed, G. H. & Frey, P. A. Characterization of an allylic analogue of the 5′-deoxyadenosyl radical: an intermediate in the reaction of lysine 2,3-aminomutase. Biochemistry 40, 7773–7782 (2001)
Layer, G. et al. The substrate radical of Escherichia coli oxygen-independent coproporphyrinogen III oxidase HemN. J. Biol. Chem. 281, 15727–15734 (2006)
Stubbe, J. & Van der Donk, W. A. Protein radicals in enzyme catalysis. Chem. Rev. 98, 705–762 (1998)
Buckel, W. The reversible dehydration of (R)-2-hydroxyglutarate to (E)-glutaconate. Eur. J. Biochem. 106, 439–447 (1980)
Beinert, H. & Albracht, S. P. J. New insights, ideas and unanswered questions concerning iron–sulfur clusters in mitochondria. Biochim. Biophys. Acta 683, 245–277 (1982)
Tsai, A. L., Berka, V., Kulmacz, R. J., Wu, G. & Palmer, G. An improved sample packing device for rapid freeze-trap electron paramagnetic resonance spectroscopy kinetic measurements. Anal. Biochem. 264, 165–171 (1998)
Ballinger, M. D., Reed, G. H. & Frey, P. A. An organic radical in the lysine 2,3-aminomutase reaction. Biochemistry 31, 949–953 (1992)
Acknowledgements
We thank R. K. Thauer for the use of his EPR spectrometer, and V. Schünemann and M. Bennati for the use of their freeze-quench instruments.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
The file contains Supplementary Figures S1-S3 with Legends, Supplementary Methods describing Synthesis and characterization of (labeled) substrates, Supplementary Tables S1-S5, Supplementary Discussion, and additional references. (PDF 980 kb)
Rights and permissions
About this article
Cite this article
Kim, J., Darley, D., Buckel, W. et al. An allylic ketyl radical intermediate in clostridial amino-acid fermentation. Nature 452, 239–242 (2008). https://doi.org/10.1038/nature06637
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06637
This article is cited by
-
Rapid Freeze-Quench EPR Spectroscopy: Improved Collection of Frozen Particles
Applied Magnetic Resonance (2016)
-
Structural basis of enzymatic benzene ring reduction
Nature Chemical Biology (2015)
-
Dark-operative protochlorophyllide oxidoreductase generates substrate radicals by an iron-sulphur cluster in bacteriochlorophyll biosynthesis
Scientific Reports (2014)
-
Phenylalanine catabolism in Archaeoglobus fulgidus VC-16
Archives of Microbiology (2013)
-
Clostridium sticklandii, a specialist in amino acid degradation:revisiting its metabolism through its genome sequence
BMC Genomics (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.