Abstract
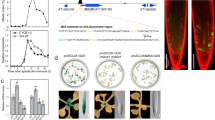
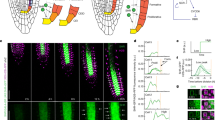
Factors with a graded distribution can program fields of cells in a dose-dependent manner1,2, but no evidence has hitherto surfaced for such mechanisms in plants. In the Arabidopsis thaliana root, two PLETHORA (PLT) genes encoding AP2-domain transcription factors have been shown to maintain the activity of stem cells3. Here we show that a clade of four PLT homologues is necessary for root formation. Promoter activity and protein fusions of PLT homologues display gradient distributions with maxima in the stem cell area. PLT activities are largely additive and dosage dependent. High levels of PLT activity promote stem cell identity and maintenance; lower levels promote mitotic activity of stem cell daughters; and further reduction in levels is required for cell differentiation. Our findings indicate that PLT protein dosage is translated into distinct cellular responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tabata, T. & Takei, Y. Morphogens, their identification and regulation. Development 131, 703–712 (2004)
Gurdon, J. B. & Bourillot, P. Y. Morphogen gradient interpretation. Nature 413, 797–803 (2001)
Aida, M. et al. The PLETHORA genes mediate patterning of the Arabidopsis root stem cell niche. Cell 119, 109–120 (2004)
Skoog, F. & Miller, C. O. Chemical regulation of growth and organ formation in plant tissues cultured in vitro . Symp. Soc. Exp. Biol. 54, 118–130 (1957)
Weigel, D. & Jurgens, G. Stem cells that make stems. Nature 415, 751–754 (2002)
Spradling, A., Drummond-Barbosa, D. & Kai, T. Stem cells find their niche. Nature 414, 98–104 (2001)
Xu, J. et al. A molecular framework for plant regeneration. Science 311, 385–388 (2006)
Sabatini, S. et al. An auxin-dependent distal organizer of pattern and polarity in the Arabidopsis root. Cell 99, 463–472 (1999)
Nole-Wilson, S., Tranby, T. L. & Krizek, B. A. AINTEGUMENTA-like (AIL) genes are expressed in young tissues and may specify meristematic or division-competent states. Plant Mol. Biol. 57, 613–628 (2005)
Birnbaum, K. et al. A gene expression map of the Arabidopsis root. Science 302, 1956–1960 (2003)
Blilou, I. et al. The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433, 39–44 (2005)
Hardtke, C. S. & Berleth, T. The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J. 17, 1405–1411 (1998)
Hellmann, H. et al. Arabidopsis AXR6 encodes CUL1 implicating SCF E3 ligases in auxin regulation of embryogenesis. EMBO J. 22, 3314–3325 (2003)
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119 (2005)
Srinivasan, C. et al. Heterologous expression of the BABY BOOM AP2/ERF transcription factor enhances the regeneration capacity of tobacco (Nicotiana tabacum L.). Planta 225, 341–351 (2007)
Wildwater, M. et al. The RETINOBLASTOMA-RELATED gene regulates stem cell maintenance in Arabidopsis roots. Cell 123, 1337–1349 (2005)
Grieneisen, V. A., Xu, J., Marée, A. F. M., Hogeweg, P. & Scheres, B. Auxin transport is sufficient to generate a maximum and gradient guiding root growth. Nature doi:10.1038/nature06215 (this issue).
O'Connor, M. B., Umulis, D., Othmer, H. G. & Blair, S. S. Shaping BMP morphogen gradients in the Drosophila embryo and pupal wing. Development 133, 183–193 (2006)
Jaeger, J. & Reinitz, J. On the dynamic nature of positional information. Bioessays 28, 1102–1111 (2006)
Hellens, R. P., Edwards, E. A., Leyland, N. R., Bean, S. & Mullineaux, P. M. pGreen: a versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol. Biol. 42, 819–832 (2000)
Aoyama, T. & Chua, N. H. A glucocorticoid-mediated transcriptional induction system in transgenic plants. Plant J. 11, 605–612 (1997)
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998)
Willemsen, V., Wolkenfelt, H., deVries, G., Weisbeek, P. & Scheres, B. The HOBBIT gene is required for formation of the root meristem in the Arabidopsis embryo. Development 125, 521–531 (1998)
Bougourd, S., Marrison, J. & Haseloff, J. Technical advance: an aniline blue staining procedure for confocal microscopy and 3D imaging of normal and perturbed cellular phenotypes in mature Arabidopsis embryos. Plant J. 24, 543–550 (2000)
Friml, J. et al. AtPIN4 mediates sink-driven auxin gradients and root patterning in Arabidopsis . Cell 108, 661–673 (2002)
Acknowledgements
We thank the Netherlands Genomics Initiative (M.L.) and the Portuguese Foundation for Science and Technology (C.G.) for funding, A. Shimotohno and J. M. Perez-Perez for sharing data and Frits Kindt for photography.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-9 with Legends and Supplementary Tables 1-3. (PDF 6235 kb)
Rights and permissions
About this article
Cite this article
Galinha, C., Hofhuis, H., Luijten, M. et al. PLETHORA proteins as dose-dependent master regulators of Arabidopsis root development. Nature 449, 1053–1057 (2007). https://doi.org/10.1038/nature06206
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature06206
This article is cited by
-
BRIP1 and BRIP2 maintain root meristem by affecting auxin-mediated regulation
Planta (2024)
-
Identification and functional analysis of a CbSHR homolog in controlling adventitious root development in Catalpa bungei
Plant Cell, Tissue and Organ Culture (PCTOC) (2024)
-
A New In Vitro Growth System for Phenotypic Characterization and Seed Propagation of Arabidopsis thaliana
Journal of Plant Growth Regulation (2024)
-
RWP-RK Domain 3 (OsRKD3) induces somatic embryogenesis in black rice
BMC Plant Biology (2023)
-
Dosage differences in 12-OXOPHYTODIENOATE REDUCTASE genes modulate wheat root growth
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.