Abstract
Specialized DNA polymerases (DNA pols) are required for lesion bypass in human cells1. Auxiliary factors have an important, but so far poorly understood, role. Here we analyse the effects of human proliferating cell nuclear antigen (PCNA) and replication protein A (RP-A) on six different human DNA pols—belonging to the B, Y and X classes—during in vitro bypass of different lesions. The mutagenic lesion 8-oxo-guanine (8-oxo-G) has high miscoding potential2,3,4. A major and specific effect was found for 8-oxo-G bypass with DNA pols λ and η. PCNA and RP-A allowed correct incorporation of dCTP opposite a 8-oxo-G template 1,200-fold more efficiently than the incorrect dATP by DNA pol λ, and 68-fold by DNA pol η, respectively. Experiments with DNA-pol-λ-null cell extracts suggested an important role for DNA pol λ. On the other hand, DNA pol ι, together with DNA pols α, δ and β, showed a much lower correct bypass efficiency. Our findings show the existence of an accurate mechanism to reduce the deleterious consequences of oxidative damage and, in addition, point to an important role for PCNA and RP-A in determining a functional hierarchy among different DNA pols in lesion bypass.
Similar content being viewed by others
Main
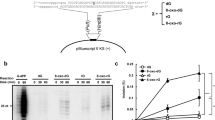
Our experiments show that DNA pol λ inserted either dATP or dCTP in vitro opposite an 8-oxo-G lesion on a template DNA oligonucleotide, and, unlike pol μ (ref. 5), did so without -1 frameshifts (Supplementary Fig. 2a), and showing a 12.5-fold higher incorporation efficiency (kcat/Km) for faithful dCTP versus error-prone dATP incorporation (Supplementary Table 1). DNA pol λ physically and functionally interacts with the proteins PCNA and RP-A (refs 6–8). PCNA increased the incorporation rates (kobs) of dCTP opposite a normal G and 8-oxo-G, but not of dATP incorporation opposite the lesion (Supplementary Fig. 2b–d). dATP (Fig. 1a, lanes 1–5) but not dCTP (lanes 6–9) incorporation opposite 8-oxo-G was inhibited by RP-A alone and, even more efficiently, by RP-A and PCNA together (Fig. 1a, lanes 13 and 17). In the presence of PCNA, addition of RP-A reduced dATP incorporation opposite 8-oxo-G to background levels (Fig. 1b compare lanes 1–5 with lanes 6–10), whereas dCTP incorporation was affected less than twofold (Fig. 1b lanes 11–15 and 16–20). In the presence of PCNA and RP-A, DNA pol λ catalysed only 1 error-prone dATP incorporation every 1.2 × 103 faithful dCTP incorporation events opposite 8-oxo-G (Table 1 and Supplementary Fig. 2e), which is sufficient to efficiently cope with the ∼103 lesions of 8-oxo-G estimated to be present in normal cells2. From the difference in the kcat/Km values for dATP versus dCTP incorporation in the presence of PCNA (Supplementary Table 1), a relative free energy change (ΔΔG) value of +1.9 kcal mol-1 was estimated to be required for generation of an A:8-oxo-G mismatch by DNA pol λ. Such a decreased thermodynamic stability may facilitate displacement of DNA pol λ by RP-A when it is incorporating dATP opposite 8-oxo-G. Identical results were obtained with two unrelated sequence contexts (Supplementary Fig. 2f, g). BSA or Escherichia coli single strand DNA binding protein could not substitute for RP-A (data not shown and refs 6, 8).
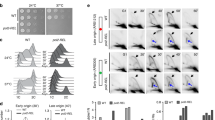
a, dATP (lanes 1–5 and 10–14) or dCTP (lanes 6–9 and 15–18) incorporation opposite 8-oxo-G with RP-A or PCNA. b, Time-dependent dATP or dCTP incorporation opposite 8-oxo-G with or without PCNA and RP-A. c, 8-oxo-G bypass by POLL+/+ or POLL-/- MEF extracts with PCNA, RP-A and dATP (lanes 1–3 and 7–9) or dCTP (lanes 4–6 and 10–12). d, Relative dATP or dCTP incorporation opposite 8-oxo-G with or without PCNA and RP-A by POLL+/+ (black) or POLL-/- (grey) MEF extracts. Values are means of three independent replicates ± s.d.
Next, extracts of mouse embryonic fibroblasts (MEFs) with or without DNA pol λ (POLL+/+ and POLL-/-, respectively) were used for in vitro 8-oxo-G bypass reactions. Aphidicolin was added to reduce background error-prone synthesis by DNA pols α, δ, ε and ζ. With the POLL+/+ extract, the ratio of dCTP versus dATP incorporation increased from 1.8 in the absence of PCNA and RP-A to 6.6 in their presence, contrary to the POLL-/- extract (1.5 and 1.8 ratio values, respectively, Fig. 1c, d). The endogenous concentrations of PCNA and RP-A were too low to contribute to the observed effects (1 µg of extract corresponded to a 1 nM final concentration of PCNA in the assay, Supplementary Fig. 3a). Addition of recombinant DNA pol λ to the POLL-/- extract restored the stimulation of the dCTP versus dATP incorporation by PCNA and RP-A (Supplementary Fig. 3b), suggesting a specific role of DNA pol λ in faithful 8-oxo-G bypass together with PCNA and RP-A.
Next we tested other DNA pols in the same experimental system. Human DNA pol η incorporated dCTP opposite an 8-oxo-G with the same efficiency (kcat/Km) as opposite a normal G (Supplementary Table 1)9, showing a 2.5-fold preference for dCTP versus dATP incorporation opposite an 8-oxo-G (Table 1). RP-A and PCNA, selectively suppressed dATP versus dCTP incorporation opposite an 8-oxo-G (Fig. 2 a–c), raising the preference for faithful versus error-prone 8-oxo-G bypass by DNA pol η to 68-fold (Table 1). This makes DNA pol η the second most accurate enzyme after DNA pol λ at 8-oxo-G.
a, dATP or dCTP incorporation by DNA pol η with PCNA. b, dATP (lanes 1–6 and 13–18) or dCTP (lanes 7–12 and 19–24) incorporation by DNA pol η with RP-A (lanes 4–6 and 10–12) or RP-A and PCNA (lanes 13–24). c, dATP (black symbols) or dCTP (open symbols) incorporation by human DNA pol η with RP-A, or RP-A and PCNA. Error bars, ±s.d. of three independent replicates. d, dCTP (lanes 1–10) or dGTP (lanes 11–21) incorporation by DNA pol ι with PCNA (lanes 2–4 and 12–14), RP-A (lanes 5–8 and 15–18) or both (lanes 9, 10, 19 and 20).
DNA pol ι incorporates dGTP with similar efficiency to dCTP opposite an 8-oxo-G (Supplementary Table 1)10. PCNA stimulated dCTP incorporation opposite the lesion (Fig. 2d, compare lanes 2 and 3 with lanes 12 and 13), but no further elongation was observed10, whereas RP-A completely inhibited 8-oxo-G bypass (lanes 5–8, 15–18). PCNA restored the ability of DNA pol ι to bypass the 8-oxo-G lesion in the presence of RP-A, resulting in a 5-fold preference for dCTP (lanes 9, 10) versus dGTP (lanes 19, 20) incorporation (Table 1). Thus, PCNA is important for 8-oxo-G lesion bypass by DNA pol ι in the presence of RP-A. DNA pol α does not interact with PCNA, but its fidelity is increased by RP-A11. As shown in Supplementary Fig. 4a, b, RP-A selectively inhibited dATP more than dCTP incorporation by DNA pol α opposite an 8-oxo-G (Supplementary Table 1). However, lesion bypass was still highly error-prone even in the presence of RP-A (Table 1). Finally, PCNA and RP-A under similar conditions did not affect the bypass of 8-oxo-G by DNA pols δ and β (Table 1 and Supplementary Fig. 4c, d).
DNA pol λ bypasses an abasic site exclusively by template-slippage, generating -1 frameshifts, whereas pol β can either skip the lesion, or incorporate dATP (ref. 12). PCNA and RP-A have been shown to promote bypass of an abasic site by Y-family DNA pols13,14, and PCNA alone stimulated abasic site bypass by DNA pol λ (ref. 7; Supplementary Fig. 5a). In the presence of PCNA, DNA pol λ bypassed the abasic site by inserting dCTP opposite the G on the template position downstream of the lesion (Supplementary Fig. 5b, lanes 1–4) but could not incorporate dATP (Supplementary Fig. 5b, lanes 5–10). RP-A did not influence the abasic-site bypass by DNA pol λ, either in the absence or in the presence of PCNA (Supplementary Fig. 5b, lanes 12–20). Also, PCNA and RP-A did not show any effect on abasic-site bypass by DNA pol β (Supplementary Fig. 5c).
The GG cis-platinum adduct has been shown to block synthesis by DNA pols α, δ and λ, even in the presence of PCNA7,15, whereas DNA pols β and η can bypass this lesion15,16. PCNA and RP-A did not significantly influence the bypass of a single cis-platinum adduct by DNA pol β (Supplementary Fig. 5d) or DNA pol η (Supplementary Fig. 5e), suggesting that bypass of bulky adducts is primarily dependent on the intrinsic properties of the DNA pol involved.
In summary our results suggest a specific role of RP-A and PCNA in the 8-oxo-G lesion bypass by DNA pol λ and—to a lower extent—by DNA pol η, and indicate that the effects of these two auxiliary proteins on the bypass activity of different DNA pols can be modulated by the nature of the lesions.
Methods summary
Nucleic acid substrates
All the oligonucleotide templates, either undamaged or containing the lesions, have been chemically synthesized and purified from polyacrylamide gels. Concentrations were determined spectrophotometrically. For sequences and details see Methods.
Proteins production and purification
All recombinant human proteins were expressed and purified according to published procedures (see Methods).
Enzymatic assays
In vitro assays were performed as described in Methods. Products were resolved on a 7 M urea/10% polyacrylamide gel and visualized by PhosphoImager (Typhoon Trio GE Healthcare).
Cells and extracts
Extracts were prepared from 107 immortalized POLL+/+ and POLL-/- MEFs in the presence of 0.4 M NaCl. For full details see Methods.
Online Methods
Chemicals
[γ-32P]ATP (3,000 Ci mmol-1) was from GE Healthcare Biosciences; unlabelled dNTPs were from Roche Molecular Biochemicals. All other reagents were of analytical grade and were purchased from Merck or Fluka.
Nucleic acids substrates
The sequences are:
72-mer 8-oxo-G template, 3′ATGTTGGTTCTCGTATGCTGCCGGTCACGGCTTAAGTGT GGCGGCCGCGGGTTGGAGGGCTTATAGATTATG;
73-mer abasic template, 3′ATAGGTGGTATGATGGGAGGTGATAGAGGTGAGTTGAGTTGGGAAGTA GGAGTTATAAGGATGGGAGGGCTAG;
60-mer cis-platinum template, 3′ATGTTGGTTCTCGTATGCTGCCGGTCACGGCTTTTCTTGGTTCCTATCGGTGGTTAGTCG.
The 72-mer and 73-mer, either undamaged or containing the 8-oxo G or abasic lesions (8-oxo-G-CE Phosphoramidite, or Tetrahydrofuran from Glen Research), and the corresponding primer were chemically synthesized. The 60-mer template containing a single cis-platinum adduct was prepared as described15. All substrate oligonucleotides and the corresponding primers were purified on a 15% (w/v) polyacrylamide, 7 M urea, 30% formamide gel. After elution and ethanol precipitation, their concentrations were determined by spectrophotometry. The bold letters correspond to the positions of the 8-oxo-G, abasic or cis-platinum lesions, respectively. The underlined sequences correspond to the primer annealing sites. The primer was 5′-labelled with T4 polynucleotide kinase (New England Biolabs) in the presence of [γ-32P]ATP. Each labelled primer was mixed with the complementary template oligonucleotide at 1:1 (M/M) ratio in the presence of 25 mM Tris-HCl (pH 8.0) and 50 mM KCl, heated at 80 °C for 3 min and then slowly cooled down at to 20 °C.
Proteins production and purification
Recombinant His-tagged human wild-type DNA pol λ, human PCNA and human RP-A were expressed and purified as described6. Human DNA pol α and human DNA pol δ were purified from HeLa cells according to the same procedure previously established for calf thymus17. Human DNA pol β was from TREVIGEN. After purification, the proteins were >90% homogenous, as judged by SDS–PAGE and Coomassie staining.
Cells and extracts.
Primary POLL+/+ and POLL-/- MEFs were generated as previously described18. Immortalized MEFs were obtained after 15 passages with a 3-day transfer regime according to a standard 3T3 culture protocol19. The cells were grown at 37 °C in a 5% CO2 incubator in DMEM containing GlutaMAX-I (Gibco BRL), 10% fetal calf serum (Brunschwig), and 100 U ml-1 penicillin–streptomycin (Gibco BRL). Extracts were prepared by adding 0.5 ml of lysis buffer (10 mM TrisHCl (pH 7.5) 1 mM EDTA (pH 8.0), 10% sucrose, 0.4 M NaCl, 1 mM DTT, 0.1 mM PMSF, protease inhibitors) to 107 POLL+/+ or POLL-/- MEFs. Cells were incubated in lysis buffer for 30 min on ice, homogenized by Dounce and centrifuged for 20 min at 4 °C at 15,000 × g. The supernatant (cell extract) was aliquoted and kept at -80 °C until use. Protein concentrations were 7 mg ml-1 for POLL+/+ and 8.5 mg ml-1 for POLL-/- MEFs.
Enzymatic assays
For denaturing gel analysis of DNA synthesis products, the reaction mixtures contained 50 mM Tris-HCl (pH 7.0), 0.25 mg ml-1 BSA, 1 mM DTT and 20 nM (0.2 pmol of 3′ OH ends) of the 5′ 32P-labelled primer/template (unless otherwise stated). DNA pols α, δ, η and ι were assayed in the presence of 5 mM Mg2+, whereas for DNA pols λ and β a final Mg2+ concentration of 1 mM was used. Concentrations of DNA pols, PCNA, RP-A, dNTPs were as indicated. Reactions were incubated for 5 min at 37 °C, unless otherwise stated, and then stopped by addition of standard denaturing gel loading buffer (95% formamide, 10 mM EDTA, xylene cyanol and bromophenol blue), heated at 95 °C for 3 min and loaded on a 7 M urea/10% polyacrylamide gel.
Steady state kinetic analysis
Reactions were performed as described above. Quantification was done by scanning densitometry. The initial velocities of the reaction were calculated from the values of integrated gel band intensities:

where T is the target site, the template position of interest; I*T is the sum of the integrated intensities at positions T, T + 1, ..., T + n.
All the intensity values were normalized to the total intensity of the corresponding lane to correct for differences in gel loading. The apparent Km and kcat values were calculated by plotting the initial velocities in dependence of the nucleotide concentrations [dNTP] and fitting the data according to the Michaelis–Menten equation:

where [E]0, was the input enzyme concentration. Nucleotide concentrations used (unless otherwise stated in the Figs) were 0.002 µM, 0.005 µM, 0.02 µM, 0.05 µM, 0.2 µM, 0.5 µM, 2 µM, 5 µM and 10 µM.
Nucleotide incorporation efficiencies were defined as the kcat/Km ratio. Under single nucleotide incorporation conditions kcat = kpolkoff/(kpol + koff) and Km = Kskoff/(kpol + koff), where kpol is the true polymerization rate, koff is the dissociation rate of the enzyme–primer complex and Ks is the true Michaelis constant for nucleotide binding. Thus, kcat/Km values are equal to kpol/Ks.
Time-dependent accumulation of products was fitted to the mixed exponential equation:

where [P] are the products formed, A is the burst amplitude, t is time, kobs is the apparent burst rate and kss is the steady-state rate, which corresponds to koff.
Free energy change (ΔΔG) values were estimated according to the relationship:

where R is the gas constant (1.9872 cal mol-1 K-1), T is the absolute temperature (K), and k is the ratio between the kinetic constants considered.
References
Hubscher, U., Maga, G. & Spadari, S. Eukaryotic DNA polymerases. Annu. Rev. Biochem. 71, 133–163 (2002)
Collins, A. R. Oxidative DNA damage, antioxidants, and cancer. Bioessays 21, 238–246 (1999)
Krahn, J. M., Beard, W. A., Miller, H., Grollman, A. P. & Wilson, S. H. Structure of DNA polymerase β with the mutagenic DNA lesion 8-oxodeoxyguanine reveals structural insights into its coding potential. Structure 11, 121–127 (2003)
Shibutani, S., Takeshita, M. & Grollman, A. P. Insertion of specific bases during DNA synthesis past the oxidation-damaged base 8-oxodG. Nature 349, 431–434 (1991)
Zhang, Y. et al. Lesion bypass activities of human DNA polymerase μ. J. Biol. Chem. 277, 44582–44587 (2002)
Maga, G., Shevelev, I., Villani, G., Spadari, S. & Hubscher, U. Human replication protein A can suppress the intrinsic in vitro mutator phenotype of human DNA polymerase λ. Nucleic Acids Res. 34, 1405–1415 (2006)
Maga, G. et al. Human DNA polymerase λ functionally and physically interacts with proliferating cell nuclear antigen in normal and translesion DNA synthesis. J. Biol. Chem. 277, 48434–48440 (2002)
Maga, G. et al. DNA elongation by the human DNA polymerase λ polymerase and terminal transferase activities are differentially coordinated by proliferating cell nuclear antigen and replication protein A. J. Biol. Chem. 280, 1971–1981 (2005)
Haracska, L., Yu, S. L., Johnson, R. E., Prakash, L. & Prakash, S. Efficient and accurate replication in the presence of 7,8-dihydro-8-oxoguanine by DNA polymerase η. Nature Genet. 25, 458–461 (2000)
Vaisman, A. & Woodgate, R. Unique misinsertion specificity of pol ι may decrease the mutagenic potential of deaminated cytosines. EMBO J. 20, 6520–6529 (2001)
Maga, G., Frouin, I., Spadari, S. & Hubscher, U. Replication protein A as a “fidelity clamp” for DNA polymerase α. J. Biol. Chem. 276, 18235–18242 (2001)
Blanca, G. et al. Human DNA polymerases λ and β show different efficiencies of translesion DNA synthesis past abasic sites and alternative mechanisms for frameshift generation. Biochemistry 43, 11605–11615 (2004)
Haracska, L., Kondratick, C. M., Unk, I., Prakash, S. & Prakash, L. Interaction with PCNA is essential for yeast DNA polymerase η function. Mol. Cell 8, 407–415 (2001)
Haracska, L. et al. Targeting of human DNA polymerase ι to the replication machinery via interaction with PCNA. Proc. Natl Acad. Sci. USA 98, 14256–14261 (2001)
Hoffmann, J. S. et al. DNA polymerase β bypasses in vitro a single d(GpG)-cisplatin adduct placed on codon 13 of the HRAS gene. Proc. Natl Acad. Sci. USA 92, 5356–5360 (1995)
Bassett, E. et al. Frameshifts and deletions during in vitro translesion synthesis past Pt-DNA adducts by DNA polymerases β and η. DNA Repair (Amst.) 1, 1003–1016 (2002)
Weiser, T. et al. Biochemical and functional comparison of DNA polymerases α, δ and ε from calf thymus. J. Biol. Chem. 266, 10420–10428 (1991)
Bertocci, B., De Smet, A., Weill, J. C. & Reynaud, C. A. Nonoverlapping functions of DNA polymerases μ, λ, and terminal deoxynucleotidyltransferase during immunoglobulin V(D)J recombination in vivo. Immunity 25, 31–41 (2006)
Todaro, G. J. & Green, H. Quantitative studies of the growth of mouse embryo cells in culture and their development into established lines. J. Cell Biol. 17, 299–313 (1963)
Acknowledgements
We thank R. Woodgate for his generous gift of recombinant human DNA pol η and ι. U.H. and U.W. are supported by the Swiss National Science Foundation, by the UBS “im Auftrag eines Kunden”, and U.H. and E.F. by the University of Zürich, which gave a grant in aid to G.M. G.M. is supported partially by the CARIPLO Foundation Project “Oncogenetica e Proteomica della Replicazione”. G.V. is supported by CNRS and ARC.
Author Contributions G.M. and U.H. had the original idea. G.M. supervised the overall experimental strategy and performed all the experiments with human DNA pol λ and cell extracts; E.C. performed all the experiments with human DNA polymerases α, δ, β, ι and η; U.W. and B.B. generated the DNA POLL+/+ and POLL-/- MEFs and characterized their phenotype; G.V. provided the damaged templates; E.F. purified DNA pols α, δ, PCNA and RP-A; G.M., G.V. and U.H. designed and interpreted all the experiments and equally contributed to manuscript writing and figures preparation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S5 with Legends and Supplementary Table S1. The Supplementary Figures provide additional evidences for the specificity of the observed effects of PCNA and RP-A on DNA polymerases lambda and eta during 8-oxo-G bypass, including results with other lesions such as abasic site and cis-platinum adduct.The Supplementary Table 1 summarizes all the kinetic constants for nucleotide incorporation by DNA polymerases lambda, alpha eta and iota opposite a normal G or an 8-oxo-G. (PDF 977 kb)
Rights and permissions
About this article
Cite this article
Maga, G., Villani, G., Crespan, E. et al. 8-oxo-guanine bypass by human DNA polymerases in the presence of auxiliary proteins. Nature 447, 606–608 (2007). https://doi.org/10.1038/nature05843
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature05843
This article is cited by
-
Distinctive roles of translesion polymerases DinB1 and DnaE2 in diversification of the mycobacterial genome through substitution and frameshift mutagenesis
Nature Communications (2022)
-
Analysis of DNA Polymerase λ Activity and Gene Expression in Response to Salt and Drought Stress in Oryza sativa Indica Rice Cultivars
Journal of Plant Growth Regulation (2022)
-
Structural basis of DNA synthesis opposite 8-oxoguanine by human PrimPol primase-polymerase
Nature Communications (2021)
-
Mutations induced by 8-hydroxyguanine (8-oxo-7,8-dihydroguanine), a representative oxidized base, in mammalian cells
Genes and Environment (2017)
-
Embryonic exposure to the widely-used herbicide atrazine disrupts meiosis and normal follicle formation in female mice
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.