Abstract
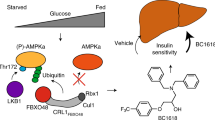
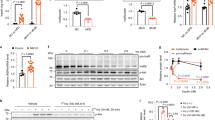
G proteins are an important class of regulatory switches in all living systems. They are activated by guanine nucleotide exchange factors (GEFs), which facilitate the exchange of GDP for GTP1,2. This activity makes GEFs attractive targets for modulating disease-relevant G-protein-controlled signalling networks3,4,5. GEF inhibitors are therefore of interest as tools for elucidating the function of these proteins and for therapeutic intervention; however, only one small molecule GEF inhibitor, brefeldin A (BFA), is currently available6,7,8,9. Here we used an aptamer displacement screen to identify SecinH3, a small molecule antagonist of cytohesins. The cytohesins are a class of BFA-resistant small GEFs for ADP-ribosylation factors (ARFs), which regulate cytoskeletal organization10, integrin activation11 or integrin signalling12. The application of SecinH3 in human liver cells showed that insulin-receptor-complex-associated cytohesins are required for insulin signalling. SecinH3-treated mice show increased expression of gluconeogenic genes, reduced expression of glycolytic, fatty acid and ketone body metabolism genes in the liver, reduced liver glycogen stores, and a compensatory increase in plasma insulin. Thus, cytohesin inhibition results in hepatic insulin resistance. Because insulin resistance is among the earliest pathological changes in type 2 diabetes, our results show the potential of chemical biology for dissecting the molecular pathogenesis of this disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cherfils, J. & Chardin, P. GEFs: structural basis for their activation of small GTP-binding proteins. Trends Biochem. Sci. 24, 306–311 (1999)
Jackson, C. L. & Casanova, J. E. Turning on ARF: the Sec7 family of guanine-nucleotide-exchange factors. Trends Cell Biol. 10, 60–67 (2000)
Hashimoto, S. et al. Requirement for Arf6 in breast cancer invasive activities. Proc. Natl Acad. Sci. USA 101, 6647–6652 (2004)
Gonzalez-Garcia, A. et al. RalGDS is required for tumor formation in a model of skin carcinogenesis. Cancer Cell 7, 219–226 (2005)
Malliri, A. et al. Mice deficient in the Rac activator Tiam1 are resistant to Ras-induced skin tumours. Nature 417, 867–871 (2002)
Donaldson, J. G., Finazzi, D. & Klausner, R. D. Brefeldin A inhibits Golgi membrane-catalysed exchange of guanine nucleotide onto ARF protein. Nature 360, 350–352 (1992)
Helms, J. B. & Rothman, J. E. Inhibition by brefeldin A of a Golgi membrane enzyme that catalyses exchange of guanine nucleotide bound to ARF. Nature 360, 352–354 (1992)
Morinaga, N., Tsai, S.-C., Moss, J. & Vaughan, M. Isolation of a brefeldin A-inhibited guanine nucleotide-exchange protein for ADP ribosylation factor (ARF) 1 and ARF3 that contains a Sec7-like domain. Proc. Natl Acad. Sci. USA 93, 12856–12860 (1996)
Peyroche, A. et al. Brefeldin A acts to stabilize an abortive ARF–GDP–Sec7 domain protein complex: involvement of specific residues of the Sec7 domain. Mol. Cell. Biol. 3, 275–285 (1999)
Mayer, G. et al. Controlling small guanine-nucleotide-exchange factor function through cytoplasmic RNA intramers. Proc. Natl Acad. Sci. USA 98, 4961–4965 (2001)
Kolanus, W. et al. αLβ2 integrin/LFA-1 binding to ICAM-1 induced by cytohesin-1, a cytoplasmic regulatory molecule. Cell 86, 233–242 (1996)
Perez, O. D. et al. Leukocyte functional antigen 1 lowers T cell activation thresholds and signaling through cytohesin-1 and Jun-activating binding protein 1. Nature Immunol. 4, 1083–1092 (2003)
Renault, L., Guibert, B. & Cherfils, J. Structural snapshots of the mechanism and inhibition of a guanine nucleotide exchange factor. Nature 426, 525–530 (2003)
Mossessova, E., Corpina, R. A. & Goldberg, J. Crystal structure of ARF1*Sec7 complexed with Brefeldin A and its implications for the guanine nucleotide exchange mechanism. Mol. Cell 12, 1403–1411 (2003)
Kruljac-Letunic, A., Moelleken, J., Kallin, A., Wieland, F. & Blaukat, A. The tyrosine kinase Pyk2 regulates Arf1 activity by phosphorylation and inhibition of the Arf-GTPase-activating protein ASAP1. J. Biol. Chem. 278, 29560–29570 (2003)
Ahmadian, M. R., Wittinghofer, A. & Herrmann, C. Fluorescence methods in the study of small GTP-binding proteins. Methods Mol. Biol. 189, 45–63 (2002)
Moss, J. & Vaughan, M. Molecules in the ARF Orbit. J. Biol. Chem. 273, 21431–21434 (1998)
Fuss, B., Becker, T., Zinke, I. & Hoch, M. The cytohesin Steppke is essential for insulin signalling in Drosophila. Nature doi:10.1038/nature05412 (this issue).
Takatsu, H., Yoshino, K., Toda, K. & Nakayama, K. GGA proteins associate with Golgi membranes through interaction between their GGAH domains and ADP-ribosylation factors. Biochem. J. 365, 369–378 (2002)
Brunet, A. et al. Akt promotes cell survival by phosphorylating and inhibiting a Forkhead transcription factor. Cell 96, 857–868 (1999)
Kops, G. J. et al. Direct control of the Forkhead transcription factor AFX by protein kinase B. Nature 398, 630–634 (1999)
Burgering, B. M. & Kops, G. J. Cell cycle and death control: long live Forkheads. Trends Biochem. Sci. 27, 352–360 (2002)
Accili, D. & Arden, K. C. FoxOs at the crossroads of cellular metabolism, differentiation, and transformation. Cell 117, 421–426 (2004)
Knight, Z. A. et al. A pharmacological map of the PI3-K family defines a role for p110α in insulin signaling. Cell 125, 733–747 (2006)
Michael, M. D. et al. Loss of insulin signaling in hepatocytes leads to severe insulin resistance and progressive hepatic dysfunction. Mol. Cell 6, 87–97 (2000)
Biddinger, S. B. & Kahn, C. R. From mice to men: insights into the insulin resistance syndromes. Annu. Rev. Physiol. 68, 123–158 (2006)
Kubota, N. et al. Disruption of insulin receptor substrate 2 causes type 2 diabetes because of liver insulin resistance and lack of compensatory beta-cell hyperplasia. Diabetes 49, 1880–1889 (2000)
Withers, D. J. et al. Disruption of IRS-2 causes type 2 diabetes in mice. Nature 391, 900–904 (1998)
Nakae, J. et al. Regulation of insulin action and pancreatic beta-cell function by mutated alleles of the gene encoding forkhead transcription factor Foxo1. Nature Genet. 32, 245–253 (2002)
Malecki, M. T. Genetics of type 2 diabetes mellitus. Diabetes Res. Clin. Pract. 68, S10–S21 (2005)
Mora, A., Lipina, C., Tronche, F., Sutherland, C. & Alessi, D. R. Deficiency of PDK1 in liver results in glucose intolerance, impairment of insulin-regulated gene expression and liver failure. Biochem. J. 385, 639–648 (2005)
Acknowledgements
We thank M. Franco for ARF6 and EFA6 plasmids, R. Quirion for the EGFP–FoxO1A plasmid, J. Kuriyan for the Ras and Sos plasmids, J.-L. Parent for the GST–GGA3 plasmid, M. Hoch for the steppke plasmid, V. Fieberg, K. Rotscheidt, N. Kuhn, R. Tolba, and A. Carney for technical assistance and the members of the Famulok laboratory for helpful discussions. This work was supported by grants from the Deutsche Forschungsgemeinschaft, the Sonderforschungsbereiche 645 and 704, the Fonds der Chemischen Industrie (to M.F.), and the Alexander von Humboldt foundation (to S.G.S.). Author Contributions M. H. and A.S. contributed equally to this work. M.H. and A.S. performed and designed, with M.F., most of the included studies. I.G. performed the aptamer displacement screen and binding and in vitro inhibition analyses of Secins. S.G.S. synthesized all Secin derivatives. W.K. provided cytohesin and ARF expression plasmids, E.K. produced the cyh3 monoclonal antibody and B.P. characterized it. T.Q. performed the analysis of Golgi integrity and ARF6 membrane recruitment. I.B. and A.S. did the immunoprecipitation experiments. M.F. supervised the research project, and assisted in the experimental design. All authors discussed the experimental results. A.S. and M.F. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains complete Supplementary Information including Supplementary Methods, Supplementary Figures 1-14 and Supplementary Tables (PDF 1079 kb)
Rights and permissions
About this article
Cite this article
Hafner, M., Schmitz, A., Grüne, I. et al. Inhibition of cytohesins by SecinH3 leads to hepatic insulin resistance. Nature 444, 941–944 (2006). https://doi.org/10.1038/nature05415
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature05415
This article is cited by
-
Targeting cytohesin-1 suppresses acute myeloid leukemia progression and overcomes resistance to ABT-199
Acta Pharmacologica Sinica (2024)
-
Activation of the GTPase ARF6 regulates invasion of human vascular smooth muscle cells by stimulating MMP14 activity
Scientific Reports (2022)
-
An ARF GTPase module promoting invasion and metastasis through regulating phosphoinositide metabolism
Nature Communications (2021)
-
The GEF Cytohesin-2/ARNO Mediates Resistin induced Phenotypic Switching in Vascular Smooth Muscle Cells
Scientific Reports (2020)
-
Detection of the in vitro modulation of Plasmodium falciparum Arf1 by Sec7 and ArfGAP domains using a colorimetric plate-based assay
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.