Abstract
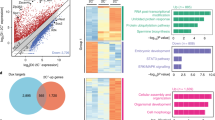
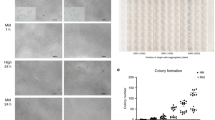
We present an integrated approach to identify genetic mechanisms that control self-renewal in mouse embryonic stem cells. We use short hairpin RNA (shRNA) loss-of-function techniques to downregulate a set of gene products whose expression patterns suggest self-renewal regulatory functions. We focus on transcriptional regulators and identify seven genes for which shRNA-mediated depletion negatively affects self-renewal, including four genes with previously unrecognized roles in self-renewal. Perturbations of these gene products are combined with dynamic, global analyses of gene expression. Our studies suggest specific biological roles for these molecules and reveal the complexity of cell fate regulation in embryonic stem cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Smith, A. G. Embryo-derived stem cells: of mice and men. Annu. Rev. Cell Dev. Biol. 17, 435–462 (2001)
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998)
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nature Genet. 24, 372–376 (2000)
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003)
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003)
Niwa, H., Burdon, T., Chambers, I. & Smith, A. Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev. 12, 2048–2060 (1998)
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003)
Burdon, T., Stracey, C., Chambers, I., Nichols, J. & Smith, A. Suppression of SHP-2 and ERK signalling promotes self-renewal of mouse embryonic stem cells. Dev. Biol. 210, 30–43 (1999)
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A. H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nature Med. 10, 55–63 (2004)
Loebel, D. A., Watson, C. M., De Young, R. A. & Tam, P. P. Lineage choice and differentiation in mouse embryos and embryonic stem cells. Dev. Biol. 264, 1–14 (2003)
Bortvin, A. et al. Incomplete reactivation of Oct4-related genes in mouse embryos cloned from somatic nuclei. Development 130, 1673–1680 (2003)
Tighe, A. P. & Gudas, L. J. Retinoic acid inhibits leukemia inhibitory factor signaling pathways in mouse embryonic stem cells. J. Cell. Physiol. 198, 223–229 (2004)
Ivanova, N. B. et al. A stem cell molecular signature. Science 298, 601–604 (2002)
Burdon, T., Smith, A. & Savatier, P. Signalling, cell cycle and pluripotency in embryonic stem cells. Trends Cell Biol. 12, 432–438 (2002)
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003)
Bamshad, M. et al. Mutations in human TBX3 alter limb, apocrine and genital development in ulnar-mammary syndrome. Nature Genet. 16, 311–315 (1997)
Mitsunaga, K. et al. Loss of PGC-specific expression of the orphan nuclear receptor ERR-β results in reduction of germ cell number in mouse embryos. Mech. Dev. 121, 237–246 (2004)
Pekarsky, Y. et al. Tcl1 enhances Akt kinase activity and mediates its nuclear translocation. Proc. Natl Acad. Sci. USA 97, 3028–3033 (2000)
Ting, D. T., Kyba, M. & Daley, G. Q. Inducible transgene expression in mouse stem cells. Methods Mol. Med. 105, 23–46 (2005)
Yasunaga, M. et al. Induction and monitoring of definitive and visceral endoderm differentiation of mouse ES cells. Nature Biotechnol. 23, 1542–1550 (2005)
Corson, L. B., Yamanaka, Y., Lai, K. M. & Rossant, J. Spatial and temporal patterns of ERK signaling during mouse embryogenesis. Development 130, 4527–4537 (2003)
Saba-El-Leil, M. K. et al. An essential function of the mitogen-activated protein kinase Erk2 in mouse trophoblast development. EMBO Rep. 4, 964–968 (2003)
Keller, G. Embryonic stem cell differentiation: emergence of a new era in biology and medicine. Genes Dev. 19, 1129–1155 (2005)
Ferri, A. L. et al. Sox2 deficiency causes neurodegeneration and impaired neurogenesis in the adult mouse brain. Development 131, 3805–3819 (2004)
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005)
Kurokawa, D. et al. Regulation of Otx2 expression and its functions in mouse epiblast and anterior neuroectoderm. Development 131, 3307–3317 (2004)
Essner, J. J., Branford, W. W., Zhang, J. & Yost, H. J. Mesendoderm and left-right brain, heart and gut development are differentially regulated by pitx2 isoforms. Development 127, 1081–1093 (2000)
Evans, A. L. & Gage, P. J. Expression of the homeobox gene Pitx2 in neural crest is required for optic stalk and ocular anterior segment development. Hum. Mol. Genet. 14, 3347–3359 (2005)
Zhang, C., Basta, T. & Klymkowsky, M. W. SOX7 and SOX18 are essential for cardiogenesis in Xenopus. Dev. Dyn. 234, 878–891 (2005)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Luo, J. et al. Placental abnormalities in mouse embryos lacking the orphan nuclear receptor ERR-β. Nature 388, 778–782 (1997)
Davenport, T. G., Jerome-Majewska, L. A. & Papaioannou, V. E. Mammary gland, limb and yolk sac defects in mice lacking Tbx3, the gene mutated in human ulnar mammary syndrome. Development 130, 2263–2273 (2003)
Kang, S. M. et al. Impaired T- and B-cell development in Tcl1-deficient mice. Blood 105, 1288–1294 (2005)
Narducci, M. G. et al. TCL1 participates in early embryonic development and is overexpressed in human seminomas. Proc. Natl Acad. Sci. USA 99, 11712–11717 (2002)
Berns, K. et al. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004)
Paddison, P. J. et al. A resource for large-scale RNA-interference-based screens in mammals. Nature 428, 427–431 (2004)
Rao, M. Conserved and divergent paths that regulate self-renewal in mouse and human embryonic stem cells. Dev. Biol. 275, 269–286 (2004)
Acknowledgements
We thank A. Smith for E14/T cells and the episomal expression system, G. Daley for Aniv15 cells, and D. Baltimore for the FUGW lentiviral expression system. We are grateful to K. A. Moore and members of the Lemischka laboratory for their assistance at various stages of these studies. We also thank A. Smith for constructive criticisms. This work was supported by funding from the NIDDK. Author Contributions N.I. and I.R.L. designed the experiments. N.I., R.L., I.K., J.L., C.D., X.S. and Y.L. performed the experiments. N.I., R.D. and I.R.L. analysed the data. N.I. and I.R.L. wrote the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
This file contains Supplementary Materials, Supplementary Tables 1–4, Supplementary Figures 1–18 and additional references. (PDF 2301 kb)
Supplementary Data 1
MAS5.0 normalized microarray data (ZIP 16494 kb)
Supplementary Data 2
MIAME description of microarray data (DOC 55 kb)
Supplementary Data 3
Description of microarray files (XLS 27 kb)
Supplementary Data 4
Non-redundant probes used in microarray analysis (XLS 3771 kb)
Supplementary Data 5
Probes affected at least in one shRNA time-course (XLS 503 kb)
Supplementary Data 6
901 probes down-regulated in RA differentiation time-course (XLS 732 kb)
Supplementary Data 7
Information for genes used in the over-expression studies (XLS 43 kb)
Supplementary Data 8
List of genes affected by Nanog shRNA (XLS 252 kb)
Supplementary Data 9
List of genes affected by Oct4 shRNA (XLS 189 kb)
Supplementary Data 10
List of genes affected by Sox2 shRNA (XLS 228 kb)
Supplementary Data 11
List of genes affected by Esrrb shRNA (XLS 149 kb)
Supplementary Data 12
List of genes affected by Tbx3 shRNA (XLS 136 kb)
Supplementary Data 13
List of genes affected by Tcl1 shRNA (XLS 100 kb)
Supplementary Data 14
List of genes affected by Mm.343880 shRNA (XLS 111 kb)
Supplementary Data 15
List of genes affected by Dppa4 shRNA (XLS 74 kb)
Supplementary Data 16
List of genes down-regulated during the Dppa4 shRNA time-course only. (XLS 20 kb)
Supplementary Data 17
List of genes down-regulated during the Mm.343880 shRNA time-course only. (XLS 17 kb)
Supplementary Data 18
List of genes up-regulated during the Mm.343880 shRNA time-course only. (XLS 11 kb)
Supplementary Data 19
List of genes down-regulated during the Nanog and Oct4 shRNA time-courses only. (XLS 13 kb)
Supplementary Data 20
List of genes down-regulated during the Nanog shRNA time-course only. (XLS 36 kb)
Supplementary Data 21
List of genes up-regulated during the Nanog shRNA time-course only. (XLS 25 kb)
Supplementary Data 22
List of genes down-regulated during the Oct4 shRNA time-course only. (XLS 21 kb)
Supplementary Data 23
List of genes up-regulated during the Oct4 shRNA time-course only. (XLS 20 kb)
Supplementary Data 24
List of genes up-regulated during the Oct4 and Sox2 shRNA time-courses only. (XLS 14 kb)
Supplementary Data 25
List of genes down-regulated in Pattern1 (XLS 27 kb)
Supplementary Data 26
List of genes up-regulated in Pattern1 (XLS 116 kb)
Supplementary Data 27
List of genes down-regulated in Pattern2 (XLS 48 kb)
Supplementary Data 28
List of genes up-regulated in Pattern2 (XLS 48 kb)
Supplementary Data 29
List of genes down-regulated in Pattern3 (XLS 15 kb)
Supplementary Data 30
List of genes up-regulated in Pattern3 (XLS 49 kb)
Supplementary Data 31
List of genes down-regulated during the Sox2 shRNA time-course only (XLS 16 kb)
Supplementary Data 32
List of genes up-regulated during the Sox2 shRNA time-course only (XLS 19 kb)
Supplementary Data 33
List of genes down-regulated during the Tbx3 shRNA time-course only (XLS 18 kb)
Rights and permissions
About this article
Cite this article
Ivanova, N., Dobrin, R., Lu, R. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006). https://doi.org/10.1038/nature04915
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04915
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.