Abstract
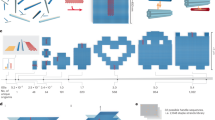
‘Bottom-up fabrication’, which exploits the intrinsic properties of atoms and molecules to direct their self-organization, is widely used to make relatively simple nanostructures. A key goal for this approach is to create nanostructures of high complexity, matching that routinely achieved by ‘top-down’ methods. The self-assembly of DNA molecules provides an attractive route towards this goal. Here I describe a simple method for folding long, single-stranded DNA molecules into arbitrary two-dimensional shapes. The design for a desired shape is made by raster-filling the shape with a 7-kilobase single-stranded scaffold and by choosing over 200 short oligonucleotide ‘staple strands’ to hold the scaffold in place. Once synthesized and mixed, the staple and scaffold strands self-assemble in a single step. The resulting DNA structures are roughly 100 nm in diameter and approximate desired shapes such as squares, disks and five-pointed stars with a spatial resolution of 6 nm. Because each oligonucleotide can serve as a 6-nm pixel, the structures can be programmed to bear complex patterns such as words and images on their surfaces. Finally, individual DNA structures can be programmed to form larger assemblies, including extended periodic lattices and a hexamer of triangles (which constitutes a 30-megadalton molecular complex).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Feynman, R. P. There's plenty of room at the bottom. Engineering and Science 23(5), 22–36 (Caltech, February, 1960).
Junno, T., Deppert, K., Montelius, L. & Samuelson, L. Controlled manipulation of nanoparticles with an atomic force microscope. Appl. Phys. Lett. 66, 3627–3629 (1995)
Eigler, D. M. & Schweizer, E. K. Positioning single atoms with a scanning tunnelling microscope. Nature 344, 524–526 (1990)
Heinrich, A. J., Lutz, C. P., Gupta, J. A. & Eigler, D. M. Molecular cascades. Science 298, 1381–1387 (2002)
Whitesides, G. M., Mathias, J. P. & Seto, C. T. Molecular self-assembly and nanochemistry: a chemical strategy for the synthesis of nanostructures. Science 254, 1312–1319 (1991)
Yokoyama, T., Yokoyama, S., Kamikado, T., Okuno, Y. & Mashiko, S. Self-assembly on a surface of supramolecular aggregates with controlled size and shape. Nature 413, 619–621 (2001)
Mao, C. B. et al. Virus-based toolkit for the directed synthesis of magnetic and semiconducting nanowires. Science 303, 213–217 (2004)
Seeman, N. C. Nucleic-acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982)
Seeman, N. C. & Lukeman, P. S. Nucleic acid nanostructures: bottom-up control of geometry on the nanoscale. Rep. Prog. Phys. 68, 237–270 (2005)
Chworos, A. et al. Building programmable jigsaw puzzles with RNA. Science 306, 2068–2072 (2004)
Park, S. H. et al. Finite-size, fully-addressable DNA tile lattices formed by hierarchical assembly procedures. Angew. Chem. 118, 749–753 (2006)
Chen, J. & Seeman, N. C. The synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991)
Zhang, Y. & Seeman, N. C. The construction of a DNA truncated octahedron. J. Am. Chem. Soc. 116, 1661–1669 (1994)
Shih, W. M., Quispe, J. D. & Joyce, G. F. A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004)
Rothemund, P. W. K. et al. Design and characterization of programmable DNA nanotubes. J. Am. Chem. Soc. 26, 16344–16353 (2004)
Fu, T.-J. & Seeman, N. C. DNA double-crossover molecules. Biochemistry 32, 3211–3220 (1993)
LaBean, T. H., Winfree, E. & Reif, J. H. in DNA Based Computers V (eds Winfree, E. & Gifford, D. K.) 123–140 ( Vol. 54 of DIMACS, AMS Press, Providence, Rhode Island, 1999)
Yan, H., LaBean, T. H., Feng, L. & Reif, J. H. Directed nucleation assembly of DNA tile complexes for barcode-patterned lattices. Proc. Natl Acad. Sci. USA 100, 8103–8108 (2003)
Reif, J. H. in Proc. 29th Int. Colloquium on Automata, Languages, and Programming (ICALP) (eds Widmayer, P., Ruiz, F. T., Bueno, R. M., Hennessy, M., Eidenbenz, S. & Conejo, R.) 1–21 (Vol. 2380 of Lecture Notes in Computer Science, Springer, New York, 2002)
Seeman, N. C. De novo design of sequences for nucleic acid structural engineering. J. Biomol. Struct. Dyn. 8, 573–581 (1990)
Winfree, E. in DNA Based Computers (eds Lipton, R. J. & Baum, E. B.) 199–221 (Vol. 27 of DIMACS, AMS Press, Providence, Rhode Island, 1996)
Rothemund, P. W. K., Papadakis, N. & Winfree, E. Algorithmic self-assembly of DNA Sierpinski triangles. PLoS Biol. 2, e424 (2004)
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003)
Le, J. D. et al. DNA-templated self-assembly of metallic nanocomponent arrays on a surface. Nano Lett. 4, 2343–2347 (2004)
Acknowledgements
I thank E. Winfree for discussions and providing a stimulating laboratory environment; B. Yurke for the term ‘nanobreadboard’; N. Papadakis, L. Adleman, J. Goto, R. Barish, R. Schulman, R. Hariadi, M. Cook and M. Diehl for discussions; B. Shaw for a gift of AFM tips; A. Schmidt for coordinating DNA synthesis; and K. Yong, J. Crouch and L. Hein for administrative support. This work was supported by National Science Foundation Career and Nano grants to E. Winfree as well as fellowships from the Beckman Foundation and Caltech Center for the Physics of Information.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The California Institute of Technology has filed a provisional patent on the method for design and creation of the folded DNA structures (scaffolded DNA origami) described in this paper. Paul W. K. Rothemund is named as the inventor on this patent.
Supplementary information
Supplementary Notes 1–11
Notes on the design process; helix bending and the inter-helix gap; models and sequences; experimental methods; control experiments; patterning with dumbbell hairpins; the combination of shapes into larger structures; secondary structure of the scaffold and staples; the robustness of the scaffolded approach; the cost of the scaffold versus staples; and additiona references. (PDF 6156 kb)
Supplementary Note 12
Full designs for all structures. Staple sequences are drawn out explicitly where they occur in the design. Because the designs are very large and the fonts are very small, this file will not print legibly. Instead of printing this file, open it in a PDF viewer and use the zoom feature to inspect the designs. (PDF 188 kb)
Rights and permissions
About this article
Cite this article
Rothemund, P. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006). https://doi.org/10.1038/nature04586
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04586
This article is cited by
-
Imaging DNA origami by fluorescence in situ hybridization
Nature Nanotechnology (2024)
-
Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly
Nature (2024)
-
Universal, label-free, single-molecule visualization of DNA origami nanodevices across biological samples using origamiFISH
Nature Nanotechnology (2024)
-
Cooperative control of a DNA origami force sensor
Scientific Reports (2024)
-
An intelligent DNA nanodevice for precision thrombolysis
Nature Materials (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.