Abstract
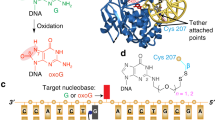
How DNA repair proteins distinguish between the rare sites of damage and the vast expanse of normal DNA is poorly understood. Recognizing the mutagenic lesion 8-oxoguanine (oxoG) represents an especially formidable challenge, because this oxidized nucleobase differs by only two atoms from its normal counterpart, guanine (G). Here we report the use of a covalent trapping strategy to capture a human oxoG repair protein, 8-oxoguanine DNA glycosylase I (hOGG1), in the act of interrogating normal DNA. The X-ray structure of the trapped complex features a target G nucleobase extruded from the DNA helix but denied insertion into the lesion recognition pocket of the enzyme. Free energy difference calculations show that both attractive and repulsive interactions have an important role in the preferential binding of oxoG compared with G to the active site. The structure reveals a remarkably effective gate-keeping strategy for lesion discrimination and suggests a mechanism for oxoG insertion into the hOGG1 active site.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Michaels, M. L. & Miller, J. H. The GO system protects organisms from the mutagenic effect of the spontaneous lesion 8-hydroxyguanine (7,8-dihydro-8-oxoguanine). J. Bacteriol. 174, 6321–6325 (1992)
Grollman, A. P. & Moriya, M. Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet. 9, 246–249 (1993)
Cappelli, E. et al. Rates of base excision repair are not solely dependent on levels of initiating enzymes. Carcinogenesis 22, 387–393 (2001)
Bruner, S. D., Norman, D. P. & Verdine, G. L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000)
Fromme, J. C., Bruner, S. D., Yang, W., Karplus, M. & Verdine, G. L. Product-assisted catalysis in base-excision DNA repair. Nature Struct. Biol. 10, 204–211 (2003)
Norman, D. P., Chung, S. J. & Verdine, G. L. Structural and biochemical exploration of a critical amino acid in human 8-oxoguanine glycosylase. Biochemistry 42, 1564–1572 (2003)
Norman, D. P. G., Bruner, S. D. & Verdine, G. L. Coupling of substrate recognition and catalysis by a human base-excision DNA repair protein. J. Am. Chem. Soc. 123, 359–360 (2001)
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997)
Viadiu, H. & Aggarwal, A. K. Structure of BamHI bound to nonspecific DNA: a model for DNA sliding. Mol. Cell 5, 889–895 (2000)
Robinson, H. et al. The hyperthermophile chromosomal protein Sac7d sharply kinks DNA. Nature 392, 202–205 (1998)
Gao, Y. G. et al. The crystal structure of the hyperthermophile chromosomal protein Sso7d bound to DNA. Nature Struct. Biol. 5, 782–786 (1998)
Huang, H., Harrison, S. C. & Verdine, G. L. Trapping of a catalytic HIV reverse transcriptase•template:primer complex through a disulfide bond. Chem. Biol. 7, 355–364 (2000)
Huang, H., Chopra, R., Verdine, G. L. & Harrison, S. C. Structure of a covalently trapped catalytic complex of HIV-1 reverse transcriptase: implications for drug resistance. Science 282, 1669–1675 (1998)
Verdine, G. L. & Norman, D. P. Covalent trapping of protein-DNA complexes. Annu. Rev. Biochem. 72, 337–366 (2003)
Fromme, J. C., Banerjee, A., Huang, S. J. & Verdine, G. L. Structural basis for removal of adenine mispaired with 8-oxoguanine by MutY adenine DNA glycosylase. Nature 427, 652–656 (2004)
Simonson, T., Archontis, G. & Karplus, M. Free energy simulations come of age: protein-ligand recognition. Acc. Chem. Res. 35, 430–437 (2002)
Yang, W., Bitetti-Putzer, R. & Karplus, M. Chaperoned alchemical free energy simulations: A general method for QM, MM, and QM/MM potentials. J. Chem. Phys. 120, 9450–9453 (2004)
Brooks, C. L. III, Karplus, M. & Pettitt, B. M. Advances in Chemical Physics Vol. 71 (Wiley & Sons, New York, 1988)
Verdine, G. L. & Bruner, S. D. How do DNA repair proteins locate damaged bases in the genome? Chem. Biol. 4, 329–334 (1997)
Bjoras, M., Seeberg, E., Luna, L., Pearl, L. H. & Barrett, T. E. Reciprocal “flipping” underlies substrate recognition and catalytic activation by the human 8-oxo-guanine DNA glycosylase. J. Mol. Biol. 317, 171–177 (2002)
Berdal, K. G., Johansen, R. F. & Seeberg, E. Release of normal bases from intact DNA by a native DNA repair enzyme. EMBO J. 17, 363–367 (1998)
O'Brien, P. J. & Ellenberger, T. The Escherichia coli 3-methyladenine DNA glycosylase AlkA has a remarkably versatile active site. J. Biol. Chem. 279, 26876–26884 (2004)
MacMillan, A. M. & Verdine, G. L. Engineering tethered DNA molecules by the convertible nucleoside approach. Tetrahedron 47, 2603–2616 (1991)
Carson, M. J. Ribbons 2.0. J. Appl. Crystallogr. 24, 379–384 (1991)
Schaefer, M., Sommer, M. & Karplus, M. pH-dependence of protein stability: absolute electrostatic free energy differences between conformations. J. Phys. Chem. B 101, 1663–1683 (1997)
Cui, Q., Elstner, M., Kaxiras, E., Frauenheim, T. & Karplus, M. A QM/MM implementation of the self-consistent charge density functional tight binding (SCC-DFTB) method. J. Phys. Chem. B 105, 569–585 (2001)
Dinner, A. R., Blackburn, G. M. & Karplus, M. Uracil-DNA glycosylase acts by substrate autocatalysis. Nature 413, 752–755 (2001)
MacKerell, A. D. Jr et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998)
Simonson, T., Archontis, G. & Karplus, M. Continuum treatment of long-range interactions in free energy calculations. Application to protein-ligand binding. J. Phys. Chem. B 101, 8347–8360 (1997)
Bash, P. A. et al. Computer simulation and analysis of the reaction pathway of triosephosphate isomerase. Biochemistry 30, 5826–5832 (1991)
Cervi, A. R., Guy, A., Leonard, G. A., Teoule, R. & Hunter, W. N. The crystal structure of N4-methylcytosine.guanosine base-pairs in the synthetic hexanucleotide d(CGCGm4CG). Nucleic Acids Res. 21, 5623–5629 (1993)
Acknowledgements
We are grateful to Y. Korkhin for help in data collection and processing. We thank Enanta Pharmaceuticals for use of their X-ray instrumentation. We acknowledge the entire staff at MACCHESS, especially C. Heaton and B. Miller, and NSLS X4A for assistance in data collection and processing. We thank D. Jeruzalmi and C. Fromme for valuable discussions. This work was supported by grants from the NIH to G.L.V. and M.K.Author contributions A.B. was responsible for performing the structural and biochemical experiments described herein, whereas W.Y. performed the computational simulations.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Table S1
Data collection and model statistics. (DOC 54 kb)
Supplementary Figure Legends
Figure captions for Supplementary Figures S1-S4. (DOC 35 kb)
Supplementary Methods
Additional details on computational methods and data collection and structure solution. (DOC 56 kb)
Supplementary Figure S1
Structural validation of the trapping strategy and crosslinking biochemistry. (PDF 658 kb)
Supplementary Figure S2
Electron density map around the G in the exo- site and the crosslinked C in the G-complex. (JPG 550 kb)
Supplementary Figure S3
Structural representations of the carbonyl of Gly42 and different nucleobases used in this study. (JPG 149 kb)
Supplementary Figure S4
Electron density maps around the analogs in the crosslinked complexes. (JPG 767 kb)
Rights and permissions
About this article
Cite this article
Banerjee, A., Yang, W., Karplus, M. et al. Structure of a repair enzyme interrogating undamaged DNA elucidates recognition of damaged DNA. Nature 434, 612–618 (2005). https://doi.org/10.1038/nature03458
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03458
This article is cited by
-
Small-molecule-mediated OGG1 inhibition attenuates pulmonary inflammation and lung fibrosis in a murine lung fibrosis model
Nature Communications (2023)
-
Tandem regulation of MutS activity by ATP and DNA during MMR initiation
Nature Structural & Molecular Biology (2022)
-
Crystal structure and DNA cleavage mechanism of the restriction DNA glycosylase R.CcoLI from Campylobacter coli
Scientific Reports (2021)
-
DNA repair glycosylase hNEIL1 triages damaged bases via competing interaction modes
Nature Communications (2021)
-
Non-flipping DNA glycosylase AlkD scans DNA without formation of a stable interrogation complex
Communications Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.