Abstract
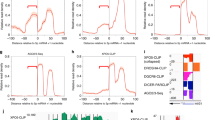
MicroRNAs (miRNAs) are a growing family of small non-protein-coding regulatory genes that regulate the expression of homologous target-gene transcripts. They have been implicated in the control of cell death and proliferation in flies1,2, haematopoietic lineage differentiation in mammals3, neuronal patterning in nematodes4 and leaf and flower development in plants5,6,7,8. miRNAs are processed by the RNA-mediated interference machinery. Drosha is an RNase III enzyme that was recently implicated in miRNA processing. Here we show that human Drosha is a component of two multi-protein complexes. The larger complex contains multiple classes of RNA-associated proteins including RNA helicases, proteins that bind double-stranded RNA, novel heterogeneous nuclear ribonucleoproteins and the Ewing's sarcoma family of proteins. The smaller complex is composed of Drosha and the double-stranded-RNA-binding protein, DGCR8, the product of a gene deleted in DiGeorge syndrome. In vivo knock-down and in vitro reconstitution studies revealed that both components of this smaller complex, termed Microprocessor, are necessary and sufficient in mediating the genesis of miRNAs from the primary miRNA transcript.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Brennecke, J. et al. bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113, 25–36 (2003)
Xu, P. et al. The Drosophila microRNA mir-14 suppresses cell death and is required for normal fat metabolism. Curr. Biol. 13, 790–795 (2003)
Chen, C. Z. et al. MicroRNAs modulate hematopoietic lineage differentiation. Science 303, 83–86 (2004)
Johnston, R. J. & Hobert, O. A microRNA controlling left/right neuronal asymmetry in Caenorhabditis elegans. Nature 426, 845–849 (2003)
Aukerman, M. J. & Sakai, H. Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-like target genes. Plant Cell 15, 2730–2741 (2003)
Chen, X. et al. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022–2025 (2004)
Emery, J. F. et al. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr. Biol. 13, 1768–1774 (2003)
Palatnik, J. F. et al. Control of leaf morphogenesis by microRNAs. Nature 425, 257–263 (2003)
Lagos-Quintana, M. et al. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001)
Lau, N. C. et al. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862 (2001)
Lee, R. C. & Ambros, V. An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864 (2001)
Aravin, A. A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003)
Lagos-Quintana, M. et al. New microRNAs from mouse and human. RNA 9, 175–179 (2003)
Lim, L. P. et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 17, 991–1008 (2003)
Lee, Y., Jeon, K., Lee, J. T., Kim, S. & Kim, V. N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002)
Zeng, Y. & Cullen, B. R. MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc. Natl Acad. Sci. USA 100, 9779–9784 (2003)
Lee, Y. et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003)
Denali, A. M., Tops, B. B. J., Plasterk, R. H. A., Ketting, R. F. & Hannon, G. J. Processing of primary microRNAs by the Microprocessor complex. Nature doi:10.1038/nature03049 (this issue)
Yamagishi, H. & Srivastava, D. Unraveling the genetic and developmental mysteries of 22q11 deletion syndrome. Trends Mol. Med. 9, 383–389 (2003)
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Molecular cloning and expression analysis of a novel gene DGCR8 located in DiGeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 304, 184–190 (2003)
Arvand, A. & Denny, C. T. Biology of EWS/ETS fusions in Ewing's family tumors. Oncogene 20, 5747–5754 (2001)
Wu, H., Xu, H., Miraglia, L. J. & Crooke, S. T. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J. Biol. Chem. 275, 36957–36965 (2000)
Bochar, D. A. et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell 102, 257–265 (2000)
Dong, Y. et al. Regulation of BRCC, a holoenzyme complex containing BRCA1 and BRCA2, by a signalosome-like subunit and its role in DNA repair. Mol. Cell 12, 1087–1099 (2003)
Acknowledgements
We thank V. N. Kim for the Drosha cDNA; O. Delattre and A. I. Lamond for the gift of EWS and DDX17 antibodies, respectively; K. Nishikura for providing recombinant Drosha; and T. Beer (Wistar Proteomocs Facility) for expertise in the microcapillary HPLC/mass spectrometry. R.S. was supported by grants from the NIH and the American Cancer Institute. R.G. is a fellow of the Jane Coffin Child Memorial Fund for Medical Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure 1
Diagrammatic depiction of Drosha-associated proteins and their domain structures. (PDF 47 kb)
Supplementary Figure 2
The bulk of miRNA processing activity resides in Drosha/DGCR8 complex. (PDF 281 kb)
Supplementary Figure 3
Analysis of miRNA precursors. (PDF 425 kb)
Rights and permissions
About this article
Cite this article
Gregory, R., Yan, Kp., Amuthan, G. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004). https://doi.org/10.1038/nature03120
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature03120
This article is cited by
-
N4-acetylcytidine modifies primary microRNAs for processing in cancer cells
Cellular and Molecular Life Sciences (2024)
-
An Overview of Targeted Genome Editing Strategies for Reducing the Biosynthesis of Phytic Acid: an Anti-nutrient in Crop Plants
Molecular Biotechnology (2024)
-
Nuclear receptor RXRα binds the precursor of miR-103 to inhibit its maturation
BMC Biology (2023)
-
The various role of microRNAs in breast cancer angiogenesis, with a special focus on novel miRNA-based delivery strategies
Cancer Cell International (2023)
-
Non-coding RNAs identification and regulatory networks in pathogen-host interaction in the microsporidia congenital infection
BMC Genomics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.