Abstract
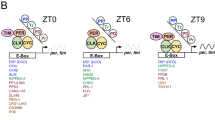
Circadian clocks drive rhythmic behaviour in animals and are regulated by transcriptional feedback loops1,2. For example, the Drosophila proteins Clock (Clk) and Cycle (Cyc) activate transcription of period (per) and timeless (tim). Per and Tim then associate, translocate to the nucleus, and repress the activity of Clk and Cyc. However, post-translational modifications are also critical to proper timing. Per and Tim undergo rhythmic changes in phosphorylation1, and evidence supports roles for two kinases in this process: Doubletime (Dbt) phosphorylates Per3,4, whereas Shaggy (Sgg) phosphorylates Tim5. Yet Sgg and Dbt often require a phosphoserine in their target site6,7, and analysis of Per phosphorylation in dbt mutants3,8 suggests a role for other kinases. Here we show that the catalytic subunit of Drosophila casein kinase 2 (CK2α) is expressed predominantly in the cytoplasm of key circadian pacemaker neurons. CK2α mutant flies show lengthened circadian period, decreased CK2 activity, and delayed nuclear entry of Per. These effects are probably direct, as CK2α specifically phosphorylates Per in vitro. We propose that CK2 is an evolutionary link between the divergent circadian systems of animals, plants and fungi.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Allada, R., Emery, P., Takahashi, J. S. & Rosbash, M. Stopping time: the genetics of fly and mouse circadian clocks. Annu. Rev. Neurosci. 24, 1091–1119 (2001)
Toh, K. L. et al. An hPer2 phosphorylation site mutation in familial advanced sleep phase syndrome. Science 291, 1040–1043 (2001)
Price, J. L. et al. double-time is a novel Drosophila clock gene that regulates PERIOD protein accumulation. Cell 94, 83–95 (1998)
Kloss, B. et al. The Drosophila clock gene double-time encodes a protein closely related to human casein kinase Iɛ. Cell 94, 97–107 (1998)
Martinek, S., Inonog, S., Manoukian, A. S. & Young, M. W. A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell 105, 769–779 (2001)
Fiol, C. J., Wang, A., Roeske, R. W. & Roach, P. J. Ordered multisite protein phosphorylation. Analysis of glycogen synthase kinase 3 action using model peptide substrates. J. Biol. Chem. 265, 6061–6065 (1990)
Flotow, H. et al. Phosphate groups as substrate determinants for casein kinase I action. J. Biol. Chem. 265, 14264–14269 (1990)
Suri, V., Hall, J. C. & Rosbash, M. Two novel doubletime mutants alter circadian properties and eliminate the delay between RNA and protein in Drosophila. J. Neurosci. 20, 7547–7555 (2000)
Chen, B., Chu, T., Harms, E., Gergen, J. P. & Strickland, S. Mapping of Drosophila mutations using site-specific male recombination. Genetics 149, 157–163 (1998)
Saxena, A., Padmanabha, R. & Glover, C. V. Isolation and sequencing of cDNA clones encoding alpha and beta subunits of Drosophila melanogaster casein kinase II. Mol. Cell Biol. 7, 3409–3417 (1987)
Niefind, K., Guerra, B., Pinna, L. A., Issinger, O. G. & Schomburg, D. Crystal structure of the catalytic subunit of protein kinase CK2 from Zea mays at 2.1 ? resolution. EMBO J. 17, 2451–2462 (1998)
Goueli, B. S., Hsiao, K., Tereba, A. & Goueli, S. A. A novel and simple method to assay the activity of individual protein kinases in a crude tissue extract. Anal. Biochem. 225, 10–17 (1995)
Dahmus, G. K., Glover, C. V., Brutlag, D. L. & Dahmus, M. E. Similarities in structure and function of calf thymus and Drosophila casein kinase II. J. Biol. Chem. 259, 9001–9006 (1984)
Helfrich-Forster, C. The period clock gene is expressed in central nervous system neurons which also produce a neuropeptide that reveals the projections of circadian pacemaker cells within the brain of Drosophila melanogaster. Proc. Natl Acad. Sci. USA 92, 612–616 (1995)
Park, J. H. & Hall, J. C. Isolation and chronobiological analysis of a neuropeptide pigment-dispersing factor gene in Drosophila melanogaster. J. Biol. Rhythms 13, 219–228 (1998)
Renn, S. C., Park, J. H., Rosbash, M., Hall, J. C. & Taghert, P. H. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila. Cell 99, 791–802 (1999); erratum 101, 113 (2000)
Zeng, H., Qian, Z., Myers, M. P. & Rosbash, M. A light-entrainment mechanism for the Drosophila circadian clock. Nature 380, 129–135 (1996)
Harmer, S. L., Panda, S. & Kay, S. A. Molecular bases of circadian rhythms. Annu. Rev. Cell Dev. Biol. 17, 215–253 (2001)
Sugano, S., Andronis, C., Green, R. M., Wang, Z. Y. & Tobin, E. M. Protein kinase CK2 interacts with and phosphorylates the Arabidopsis circadian clock-associated 1 protein. Proc. Natl Acad. Sci. USA 95, 11020–11025 (1998)
Sugano, S., Andronis, C., Ong, M. S., Green, R. M. & Tobin, E. M. The protein kinase CK2 is involved in regulation of circadian rhythms in Arabidopsis. Proc. Natl Acad. Sci. USA 96, 12362–12366 (1999)
Yang, Y., Cheng, P. & Liu, Y. Regulation of the Neurospora circadian clock by casein kinase II. Genes Dev. 16, 994–1006 (2002)
Ghavidel, A. & Schultz, M. C. TATA binding protein-associated CK2 transduces DNA damage signals to the RNA polymerase III transcriptional machinery. Cell 106, 575–584 (2001)
Keller, D. M. et al. A DNA damage-induced p53 serine 392 kinase complex contains CK2, hSpt16, and SSRP1. Mol. Cell 7, 283–292 (2001)
Cashmore, A. R., Jarillo, J. A., Wu, Y. J. & Liu, D. Cryptochromes: blue light receptors for plants and animals. Science 284, 760–765 (1999)
Allada, R., White, N. E., So, W. V., Hall, J. C. & Rosbash, M. A mutant Drosophila homolog of mammalian Clock disrupts circadian rhythms and transcription of period and timeless. Cell 93, 791–804 (1998)
Hamblen, M. et al. Germ-line transformation involving DNA from the period locus in Drosophila melanogaster: overlapping genomic fragments that restore circadian and ultradian rhythmicity to per0 and per- mutants. J. Neurogenet. 3, 249–291 (1986)
Kaneko, M., Helfrich-Forster, C. & Hall, J. C. Spatial and temporal expression of the period and timeless genes in the developing nervous system of Drosophila: newly identified pacemaker candidates and novel features of clock gene product cycling. J. Neurosci. 17, 6745–6760 (1997)
Park, J. H. et al. Differential regulation of circadian pacemaker output by separate clock genes in Drosophila. Proc. Natl Acad. Sci. USA 97, 3608–3613 (2000)
Acknowledgements
We thank J. Rutila for assistance and guidance in conducting the genetic screen; J. Biswas for assistance with sequencing; A. McElvaine for cloning; L. McCarty for western blots; R. Scharnweber for helping to set up the CK2 activity assays; Bloomington Stock Center for fly stocks; C. Glover for CK2 antibodies; and the Northwestern University Biological Imaging Facility for assistance with confocal microscopy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Lin, JM., Kilman, V., Keegan, K. et al. A role for casein kinase 2α in the Drosophila circadian clock. Nature 420, 816–820 (2002). https://doi.org/10.1038/nature01235
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01235
This article is cited by
-
Time-course RNASeq of Camponotus floridanus forager and nurse ant brains indicate links between plasticity in the biological clock and behavioral division of labor
BMC Genomics (2022)
-
Proteomic analysis of Drosophila CLOCK complexes identifies rhythmic interactions with SAGA and Tip60 complex component NIPPED-A
Scientific Reports (2020)
-
Effects of MUL1 and PARKIN on the circadian clock, brain and behaviour in Drosophila Parkinson’s disease models
BMC Neuroscience (2019)
-
Emerging roles for microRNA in the regulation of Drosophila circadian clock
BMC Neuroscience (2018)
-
Cardinal Epigenetic Role of non-coding Regulatory RNAs in Circadian Rhythm
Molecular Neurobiology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.