Abstract
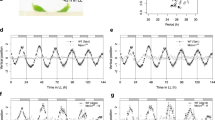
Many plants use day length as an environmental cue to ensure proper timing of the switch from vegetative to reproductive growth. Day-length sensing involves an interaction between the relative length of day and night, and endogenous rhythms that are controlled by the plant circadian clock1. Thus, plants with defects in circadian regulation cannot properly regulate the timing of the floral transition2. Here we describe the gene EARLY FLOWERING 4 (ELF4), which is involved in photoperiod perception and circadian regulation. ELF4 promotes clock accuracy and is required for sustained rhythms in the absence of daily light/dark cycles. elf4 mutants show attenuated expression of CIRCADIAN CLOCK ASSOCIATED 1 (CCA1), a gene that is thought to function as a central oscillator component3,4. In addition, elf4 plants transiently show output rhythms with highly variable period lengths before becoming arrhythmic. Mutations in elf4 result in early flowering in non-inductive photoperiods, which is probably caused by elevated amounts of CONSTANS (CO), a gene that promotes floral induction5.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Thomas, B. & Vince-Prue, D. Photoperiodism in Plants (Academic, San Diego, 1997)
Carré, I. A. Day-length perception and the photoperiodic regulation of flowering in Arabidopsis. J. Biol. Rhythms 16, 416–424 (2001)
Alabadi, D. et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880–883 (2001)
Wang, Z. & Tobin, E. M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 93, 1207–1217 (1998)
Putterill, J., Robson, F., Lee, K., Simon, R. & Coupland, G. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell 80, 847–857 (1995)
Koornneef, M., Hanhart, C. J. & van der Veen, J. H. A genetic and physiological analysis of late flowering mutants in Arabidopsis thaliana. Mol. Gen. Genet. 229, 57–66 (1991)
Samach, A. & Coupland, G. Time measurement and the control of flowering in plants. BioEssays 22, 38–47 (2000)
Dowson-Day, M. J. & Millar, A. J. Circadian dysfunction causes aberrant hypocotyl elongation patterns in Arabidopsis. Plant J. 17, 63–71 (1999)
Hicks, K. A. et al. Conditional circadian dysfunction of the Arabidopsis early-flowering 3 mutant. Science 274, 790–792 (1996)
Zagotta, M. T. et al. The Arabidopsis ELF3 gene regulates vegetative photomorphogenesis and the photoperiodic induction of flowering. Plant J. 10, 691–702 (1996)
McWatters, H. G., Bastow, R. M., Hall, A. & Millar, A. J. The ELF3 zeitnehmer regulates light signalling to the circadian clock. Nature 408, 716–720 (2000)
Liu, X. L., Covington, M. F., Fankhauser, C., Chory, J. & Wagner, D. R. ELF3 encodes a circadian clock-regulated nuclear protein that functions in an Arabidopsis PHYB signal transduction pathway. Plant Cell 13, 1293–1304 (2001)
Covington, M. F. et al. ELF3 modulates resetting of the circadian clock in Arabidopsis. Plant Cell 13, 1305–1315 (2001)
Hicks, K. A., Albertson, T. M. & Wagner, D. R. EARLY FLOWERING 3 encodes a novel protein that regulates circadian clock function and flowering in Arabidopsis. Plant Cell 13, 1281–1292 (2001)
Reed, J. W. et al. Independent action of ELF3 and phyB to control hypocotyl elongation and flowering time. Plant Physiol. 122, 1149–1160 (2000)
Bell-Pedersen, D., Crosthwaite, S. K., Lakin-Thomas, P. L., Merrow, M. & Okland, M. The Neurospora circadian clock: simple or complex? Phil. Trans. R. Soc. Lond. B 356, 1697–1709 (2001)
Aronson, B. D., Johnson, K. A. & Dunlap, J. C. Circadian clock locus frequency: protein encoded by a single open reading frame defines period length and temperature compensation. Proc. Natl Acad. Sci. USA 91, 7683–7687 (1994)
Strayer, C. et al. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science 289, 768–771 (2000)
Feldmann, K. A. T-DNA insertion mutagenesis in Arabidopsis: mutational spectrum. Plant J. 1, 71–82 (1991)
Onouchi, H., Igeno, M. I., Perilleux, C., Graves, K. & Coupland, G. Mutagenesis of plants overexpressing CONSTANS demonstrates novel interactions among Arabidopsis flowering-time genes. Plant Cell 12, 885–900 (2000)
Suarez-Lopez, P. et al. CONSTANS mediates between the circadian clock and the control of flowering in Arabidopsis. Nature 410, 1116–1120 (2001)
Davis, S. J., Bhoo, S. H., Durski, A. M., Walker, J. M. & Vierstra, R. D. The heme-oxygenase family required for phytochrome chromophore biosynthesis is necessary for proper photomorphogenesis in higher plants. Plant Physiol. 126, 656–669 (2001)
Kardailsky, I. et al. Activation tagging of the floral inducer FT. Science 286, 1962–1965 (1999)
Bognar, L. K. et al. The circadian clock controls the expression pattern of the circadian input photoreceptor, phytochrome B. Proc. Natl Acad. Sci. USA 96, 14652–14657 (1999)
Carré, I. A. & Kay, S. A. Multiple DNA–protein complexes at a circadian-regulated promoter element. Plant Cell 7, 2039–2051 (1995)
Acknowledgements
This work was supported by the College of Agricultural and Life Sciences of the University of Wisconsin and by a grant to R.A. from the National Science Foundation. M.R.D. was supported by a Molecular Biosciences Training Grant (NIH); R.M.B. was supported by a Gatsby graduate studentship; S.J.D. is a Department of Energy Bioscience fellow of the Life Sciences Research Foundation. Work in Warwick was supported by grants from the Biotechnology and Biological Sciences Research Council and the Human Frontier Science Program (HFSP) to A.J.M. The work in Hungary was supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Doyle, M., Davis, S., Bastow, R. et al. The ELF4 gene controls circadian rhythms and flowering time in Arabidopsis thaliana. Nature 419, 74–77 (2002). https://doi.org/10.1038/nature00954
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature00954
This article is cited by
-
Assembly and operation of an imaging system for long-term monitoring of bioluminescent and fluorescent reporters in plants
Plant Methods (2023)
-
Limited water stress modulates expression of circadian clock genes in Brachypodium distachyon roots
Scientific Reports (2023)
-
Transcriptomal dissection of soybean circadian rhythmicity in two geographically, phenotypically and genetically distinct cultivars
BMC Genomics (2021)
-
A central circadian oscillator confers defense heterosis in hybrids without growth vigor costs
Nature Communications (2021)
-
Comparative transcriptomic analyses of normal and peloric mutant flowers in Cymbidium goeringii Rchb.f identifies differentially expressed genes associated with floral development
Molecular Biology Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.