Abstract
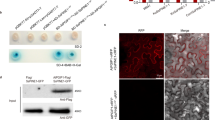
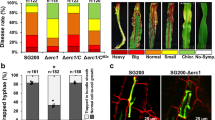
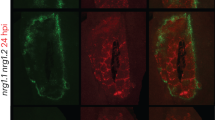
Plant disease resistance can be conferred by constitutive features such as structural barriers or preformed antimicrobial secondary metabolites. Additional defence mechanisms are activated in response to pathogen attack and include localized cell death (the hypersensitive response)1,2. Pathogens use different strategies to counter constitutive and induced plant defences, including degradation of preformed antimicrobial compounds3 and the production of molecules that suppress induced plant defences4,5,6. Here we present evidence for a two-component process in which a fungal pathogen subverts the preformed antimicrobial compounds of its host and uses them to interfere with induced defence responses. Antimicrobial saponins are first hydrolysed by a fungal saponin-detoxifying enzyme. The degradation product of this hydrolysis then suppresses induced defence responses by interfering with fundamental signal transduction processes leading to disease resistance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hammond-Kosack, K. & Jones, J. D. G. in Biochemistry and Molecular Biology of Plants (eds Buchanan, B. B., Gruissem, W. & Jones, R. L.) 1102–1156 (American Society of Plant Physiologists, Rockville, Maryland, 2001)
Flor, H. H. Current status of the gene-for-gene concept. Annu. Rev. Phytopathol. 9, 275–296 (1971)
Morrissey, J. P. & Osbourn, A. E. Fungal resistance to plant antibiotics as a mechanism of pathogenesis. Microbiol. Mol. Biol. Rev. 63, 708–724 (1999)
Doke, N. et al. in Molecular Genetics of Host-Specific Toxins in Plant Disease (eds Kohmoto, K. & Yoder, O. C.) 331–341 (Kluwer, Dordrecht, 1998)
Shiraishi, T. et al. in Molecular Genetics of Host-Specific Toxins in Plant Disease (eds Kohmoto, K. & Yoder, O. C.) 343–349 (Kluwer, Dordrecht, 1998)
Heath, M. C. Hypersensitive response-related death. Plant Mol. Biol. 44, 321–334 (2002)
Roddick, J. The steroidal glycoalkaloid tomatine. Phytochemistry 13, 9–25 (1974)
Arneson, P. A. & Durbin, R. D. Studies on the mode of action of tomatine as a fungitoxic agent. Plant Physiol. 43, 683–686 (1968)
Arneson, P. A. & Durbin, R. D. The sensitivity of fungi to α-tomatine. Phytopathology 58, 536–537 (1968)
Sandrock, R. W., DellaPenna, D. & VanEtten, H. D. Purification and characterization of β2-tomatinase, an enzyme involved in the degradation of α-tomatine and isolation of the gene encoding β2-tomatinase from Septoria lycopersici. Mol. Plant Microbe Interact. 8, 960–970 (1995)
Sandrock, R. W. & VanEtten, H. D. Fungal sensitivity to and enzymatic degradation of the phytoanticipin α-tomatine. Phytopathology 88, 137–143 (1998)
Martin-Hernandez, A. M., Dufresne, M., Hugouvieux, V., Melton, R. & Osbourn, A. Effects of targeted replacement of the tomatinase gene on the interaction of Septoria lycopersici with tomato plants. Mol. Plant Microbe Interact. 13, 1301–1311 (2000)
Arneson, P. A. & Durbin, R. D. Hydrolysis of tomatine by Septoria lycopersici: a detoxification mechanism. Phytopathology 57, 1358–1360 (1967)
Durbin, R. D. & Uchytil, T. F. Purification and properties of a fungal β-glucosidase acting on α-tomatine. Biochim. Biophys. Acta. 191, 176–178 (1969)
Osbourn, A. E., Bowyer, P., Lunness, P., Clarke, B. & Daniels, M. Fungal pathogens of oat roots and tomato leaves employ closely related enzymes to detoxify different host plant saponins. Mol. Plant Microbe Interact. 8, 971–978 (1995)
Sandrock, R. W. & VanEtten, H. D. The relevance of tomatinase activity in pathogens of tomato: disruption of the β2-tomatinase gene in Colletotrichum coccodes and Septoria lycopersici and heterologous expression of the Septoria lycopersici β2-tomatinase in Nectria haematococca, a pathogen of tomato fruit. Physiol. Mol. Plant Pathol. 58, 159–171 (2001)
Baulcombe, D. C. Fast forward genetics based on virus-induced gene silencing. Curr. Opin. Plant Biol. 2, 109–113 (1999)
Tronchet, M., Ranty, B., Marco, Y. & Roby, D. HSR203 antisense suppression in tobacco accelerates development of hypersensitive cell death. Plant J. 27, 115–127 (2001)
Yoshioka, K. et al. Environmentally sensitive, SA-dependent defense responses in the cpr22 mutant of Arabidopsis. Plant J. 26, 447–459 (2001)
Froehlich, J. E., Itoh, A. & Howe, G. A. Tomato allene oxide synthase and fatty acid hydroperoxide lyase, two cytochrome P450s involved in oxylipin metabolism, are targeted to different membranes of chloroplast envelope. Plant Physiol. 125, 306–317 (2001)
Ziegler, J., Keinanen, M. & Baldwin, I. T. Herbivore-induced allene oxide synthase transcripts and jasmonic acid in Nicotiana attenuata. Phytochemistry 58, 729–738 (2001)
Austin, M. J. et al. Regulatory role of SGT1 in early R gene-mediated plant defenses. Science 292, 2077–2080 (2002)
Azevedo, C. et al. The RAR1 interactor SGT1, an essential component of R gene-triggered disease resistance. Science 292, 2073–2076 (2002)
Peart, J. et al. SGT1 is required for host and nonhost disease resistance in plants. Proc. Natl Acad. Sci. USA advance online publication, 15 July 2002 (DOI 10.1073/pnas.152330599)
Martin, G. B. et al. Map-based cloning of a protein kinase gene conferring disease resistance in tomato. Science 262, 1432–1436 (1993)
Scofield, S. R. et al. Molecular basis of gene-for-gene specificity in bacterial speck disease of tomato. Science 274, 2063–2065 (1996)
Rommens, C. M. T., Salmeron, J. M., Oldroyd, G. E. D. & Staskawicz, B. J. Intergenic transfer and functional expression of the tomato disease resistance gene Pto. Plant Cell 7, 1537–1544 (1995)
Voinnet, O. & Baulcombe, D. Systemic signalling in gene silencing. Nature 389, 553–553 (1997)
Singh, S., Khanna, N. M. & Dhar, M. M. Solaplumbin, a new anticancer glycoside from Nicotiana plumbaginifolia. Phytochemistry 13, 2020–2022 (1974)
Grünweller, S., Schröder, E. & Kesselmeier, J. Biological activities of furostanol saponins from Nicotiana tabacum. Phytochemistry 29, 2485–2490 (1990)
Acknowledgements
We thank R. Sandrock and H. Van Etten for strains of S. lycopersici; P. Moffett for supplying SGT1-silenced N. benthamiana plants; and D. Holden and J. Rathjen for criticism of the manuscript. K.B. is supported by a Marie Curie European Community fellowship and the Sainsbury Laboratory is supported by the Gatsby Charitable Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Bouarab, K., Melton, R., Peart, J. et al. A saponin-detoxifying enzyme mediates suppression of plant defences. Nature 418, 889–892 (2002). https://doi.org/10.1038/nature00950
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature00950
This article is cited by
-
Yeast lacking the sterol C-5 desaturase Erg3 are tolerant to the anti-inflammatory triterpenoid saponin escin
Scientific Reports (2023)
-
Pathogenic Factors of Plant Pathogenic Streptomyces
Potato Research (2023)
-
Multiomics analysis of the giant triton snail salivary gland, a crown-of-thorns starfish predator
Scientific Reports (2017)
-
Response of peanut Arachis hypogaea roots to the presence of beneficial and pathogenic fungi by transcriptome analysis
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.