Abstract
Multiple myeloma is cytogenetically heterogeneous and a hyperdiploid karyotype is considered currently to have standard risk. In this study, we investigated the clinical impact of additional–structural–chromosomal aberrations assessed by chromosome analysis in 284 patients with a hyperdiploid karyotype that were subdivided into four groups based on the complexity of additional–structural–chromosomal aberrations: group 1, no additional–structural–chromosomal aberrations (n=35); group 2, one additional–structural–chromosomal aberration (n=46); group 3, two additional–structural–chromosomal aberrations (n=39); group 4, ≥three additional–structural–chromosomal aberrations (n=164). Clinicopathological data among these groups showed no differences, except patients in group 1 had higher hemoglobin (P=0.031) and albumin (P=0.045) levels. The median follow-up was 55 months (range, 3–221). The median overall survival of patients in groups 1–4 was negatively correlated with the number of the additional–structural–chromosomal aberrations: 98, 76, 61, and 48 months, respectively (P<0.0001). In group 4, CKS1B gain, RB1, or TP53 deletions had no additional impact on overall survival; however, trisomy 3 or 15 conferred a much better overall survival, and monosomy 13 and 14 predicted a worse outcome. In addition, the overall survival of patients in groups 3 and 4 was similar to a subset of high-risk multiple myeloma cases (n=21) (P=0.387). About 192 (67.6%) patients who received stem cell transplantation did not show improved overall survival compared with non-stem cell transplantation patients (n=92; P=0.142) overall; however, they did show significantly improved overall survival in patients with refractory disease in group 4 (P=0.0084). Multivariate analysis showed that two or more additional–structural–chromosomal aberrations (P<0.0001), stages (P=0.02 and P=0.002) and relapsed disease (P=0.009) negatively impacted the overall survival. We conclude that hyperdiploid karyotypes in multiple myeloma are associated with additional–structural–chromosomal aberrations and a greater number of additional–structural–chromosomal aberrations predicts poorer clinical outcome. A hyperdiploid karyotype with ≥2 additional–structural–chromosomal aberrations at chromosomal level should be considered an independent high-risk factor.
Similar content being viewed by others
Main
Multiple myeloma is a neoplasm of plasma cells that is heterogeneous at the clinicopathological and genetic/genomic levels. Patients can have an overall survival that ranges from months to many years. Cytogenetics has emerged as an important tool to stratify multiple myeloma patients and to facilitate a precision medicine approach.1 Several recurrent cytogenetic abnormalities are known to impact patient outcome. Adverse prognostic findings include t(4;14)/FGFR3/MMSET-IGH or t(14;16)/MAF-IGH and deletion of chromosome 17p13/TP53.2 RB1 deletion and t(11;14)/MYEOV/CCND1-IGH do not have independent prognostic value,3 whereas a hyperdiploid karyotype, defined as gains of at least two odd chromosomes, is reported to predict a better outcome.4 Hypodiploidy, defined as chromosome number count <44, or a complex karyotype, defined as the presence of three or more chromosomal aberrations commonly involving both numerical and structural aberrations with a chromosome count of 44–47, have been described to have a poor prognosis.5 Most of these cytogenetic subgroups can be easily identified by traditional chromosome analyses when there are dividing plasma cells. However, in clinical practice, most often the plasma cells are non-dividing6 and therefore, detection of recurrent cytogenetic aberrations largely has relied on interphase fluorescence in situ hybridization (FISH), especially on CD138-enriched plasma cells,7, 8, 9 to yield a higher detection rate.10 However, FISH-based testing cannot fully capture tumor heterogeneity as this method is focused on specific loci instead of being a genome-wide assessment.11 Therefore, integrated chromosome and FISH analyses have been utilized in clinical practice. Recent advances in technology have shown a much improved analytic resolution at the genomic level.12
To better predict the prognosis of multiple myeloma patients, the International Myeloma Working Group established an International Staging System that integrates clinical and laboratory parameters to assess patient risk as a part of clinical management. Most recently, the Revised International Staging System introduced in 2015 incorporated high serum lactate dehydrogenase level and chromosomal abnormalities for a more accurate risk stratification, by including translocations t(4;14)/FRFR3/MMSET-IGH or t(14;16)/MAF-IGH and deletion of chromosome 17p13/TP53 that are usually detected by interphase FISH in high-risk groups. With this update, the Revised International Staging System considered all other recurrent cytogenetic subgroups as standard-risk groups that are largely not further characterized.13 However, in the most recently proposed model of cytogenetic risk stratification by the International Myeloma Working Group,14 chromosome 1q gain, t(14;20) and non-hyperdiploid karyotype, and 13q deletion have also been included as ‘high-risk’. Nevertheless, even within these cytogenetically high-risk subgroups, overall survival can vary greatly depending on the presence of tumor heterogeneity or genomic complexity.15
A hyperdiploid karyotype is present in ~45% of multiple myeloma cases as assessed by FISH. Although a hyperdiploid karyotype is known to be associated with a relatively better outcome, there has been controversy regarding its clinical impact.16, 17, 18, 19 Some studies show that specific trisomies, such as trisomy 3 or 15, may correlate with a better outcome and may even overcome some high-risk features.16 Other studies have suggested that a hyperdiploid karyotype does not modify the prognosis in high-risk multiple myeloma patients.16 Moreover, within the hyperdiploid karyotype group, there is a wide variability in overall survival, which may be attributable heterogeneous additional–structural–chromosomal aberrations.16, 17, 18, 19, 20
In this study, we retrospectively reviewed the clinicopathological and cytogenetic/FISH data of all hyperdiploid karyotype multiple myeloma patients tested and/or treated at our institution to better understand and/or to further assess if the number of additional–structural–chromosomal aberrations in hyperdiploid karyotype multiple myeloma impacts patient outcome.
Materials and methods
Patient Inclusion
We retrospectively reviewed the cytogenetic data for all multiple myeloma patients diagnosed and/or treated at The University of Texas MD Anderson Cancer Center from January 1998 to December 2014. All patients with cytogenetic evidence of a hyperdiploid karyotype were included with the exception of those cases with additional 14q32/IGH that were analyzed in another study. Clinicopathological, laboratory, and clinical data were also reviewed. The study was approved by the Institutional Review Board.
Cytogenetic and Fluorescence In Situ Hybridization Analyses
Chromosome analysis was performed on unstimulated bone marrow cultures (24 hour and 48 hour) following standard procedures in the Clinical Cytogenetics laboratory. Karyotypes were described according to the International System for Human Cytogenetic Nomenclature (ISCN 2009 and 2013). A hyperdiploid karyotype was defined in this study as a chromosome count of 48–65 and with a gain of at least two odd chromosomes. Additional–structural aberrations include apparent chromosomal translocations, deletions, duplications, additions, inversions, insertions, and marker chromosomes etc at the level of resolution of conventional cytogenetic analysis. Interphase FISH studies were conducted on cultured bone marrow cells using four dual color FISH probe sets targeting CDKN2C/CKS1B, MYEOV/CCND1-IGH, RB1/13q34, and TP53/CEN17 (Abbott, Des Plaines, IL, USA) according to the manufacturer’s instructions and laboratory procedures. At least 200 cells were counted for each probe set.
Statistical Analysis
Statistical analyses were performed with the GraphPad Prism 6 software (GraphPad Software, La Jolla, CA, USA). We defined overall survival as the lapse time between the diagnosis and the death or last follow-up of the patient. Kaplan–Meier curves for overall survival were plotted and long-rank test was applied with a P-value <0.05 being considered as statistically significant. Multivariate analysis was performed using SPSS version 9.3 (SPSS Institute, Chicago, IL, USA).
Results
Clinicopathological Data
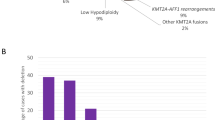
A total of 3489 patients with multiple myeloma and cytogenetic data were reviewed. Of these, 318 (9.1%) had a hyperdiploid karyotype based on chromosome analysis and patients (n=34) with additional 14q32/IGH translocations detected either by FISH or chromosome analysis were excluded to be analyzed in another study. To accurately analyze the data, we divided the remainder 284 patients based on the number of the additional–structural–chromosomal aberrations into four groups: group 1 with no additional–structural–chromosomal aberrations (n=35, 12%), group 2 with one additional–structural–chromosomal aberration (n=46, 16%), group 3 with two additional–structural–chromosomal aberrations (n=39, 13%), and group 4 with three or more additional–structural–chromosomal aberrations (n=164, 59%; Table 1 and Figure 1a). The median follow-up of these patients was 55 months (range, 3–221 months).
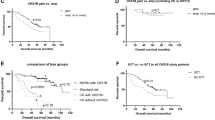
(a) Proportion of patients in each group. (b) Comparison of the median overall survival among the four groups. (c) Impact of additional–structural–chromosomal aberrations (ASAs) on the overall survival of patients with multiple myeloma in all the four groups. The Kaplan–Meier curve that represents the group 4, with three or more additional–structural–chromosomal aberrations have a significant shorter survival than the groups with one or no additional–structural–chromosomal aberrations. (d) Comparison of the overall survival between group 1 and a cytogenetic control group consisted of patients with a normal karyotype and normal FISH for CKS1B, MYEOV/CCND1-IGH, RB1, and TP53.
All demographic and laboratory data are summarized in Table 1. The median age was 60 years (range, 36–90), and the male:female ratio was 2. The median white blood cell count, platelet count, serum creatinine, beta-2 microglobuline, lactic dehydrogenase, serum calcium, paraprotein level, and bone marrow plasma cell count did not show significant differences among all the four groups, except serum albumin and hemoglobin levels were higher in group 1 (P=0.031 and P=0.045, respectively).
As we excluded all the 14q32/IGH rearranged cases, we used TP53 deletion as the high-risk chromosomal abnormality for staging, following the Revised International Scoring System. The distribution of stages did not reach significant differences among the four groups as the following: in group I, the stages were 33% stage I, 22% stage II, and 45% stage III; in group 2, stages were 20% stage I, 20% II, and 60% III; in group 3, 33% stage I, 18% II, and III 49%; and in group 4, 25% stage I, 18% II, and 57% III. The percentages of patients with newly diagnosed vs relapsed and refractory disease at the time of the cytogenetic analysis were 60% newly diagnosed vs 31% relapsed vs 9% refractory in group 1; 65% vs 15% vs 20% in group 2; 51% vs 28% vs 21% in group 3; and 49% vs 35% vs 16% in group 4, respectively with no significantly different distributions observed (Table 2).
Clinical Outcomes among the Four Groups
Almost all (97.9%) patients were diagnosed and/or treated after 2002 and received standard clinical management with immunomodulatory drugs and proteasome inhibitors, followed by stem cell transplantation in many patients.21 During the follow-up interval, 5 of 35 (14%) patients in group 1, 12 of 46 (26%) in group 2, 13 of 39 (33%) in group 3, and 77 of 164 (47%) patients in group 4 had died with group 4 showing a significantly higher mortality rate (P<0.0001). Among the newly diagnosed patients in group 1 (n=21), group 2 (n=30), group 3 (n=20), and group 4 (n=81), the death rate was 14% (3/21), 33.3% (10/30), 40% (8/20), and 52% (42/81), respectively (P=0.002) (Supplementary Table 1).
The group 1 had the longest median follow-up of 74 months and the group 4 had the shortest median follow-up of 42 months (P<0.0001; Table 2). The median overall survival for patients in group 1 was 98 vs 76 months in group 2 (P=0.119), 61 months in group 3 (P=0.0004), and 48 months in group 4 (P=0.0003) as shown in Figure 1b and c (P<0.0001). These data show that high genomic complexity has an adverse clinical impact in patients with hyperdiploid karyotype multiple myeloma (P<0.0001; Figure 1b and c). To assess the impact of additional–structural–chromosomal aberrations in newly diagnosed multiple myeloma only, we compared the overall survival among these four groups. There were 21, 30, 20, and 81 newly diagnosed patients for group 1, 2, 3, and 4 and showing an overall survival of 97, 87, 48, and 48 months, respectively (P=0.0026), all were consistent with the entire study (Supplementary Figure 1a; Supplementary Table 1).
To further assess the clinical impact of a hyperdiploid karyotype in multiple myeloma, we subsequently included the overall survival data of 35 multiple myeloma patients who had a normal karyotype and were negative for CKS1B gain, MYEOV/CCND1-IGH rearrangement, RB1, or TP53 deletions by FISH analyses as cytogenetically/FISH normal or baseline controls. Compared with this baseline group, the overall survival of patients in group 1 was 98 vs 150 months for the control group (P=0.273), indicating that even a simple hyperdiploid karyotype did not achieve an assumed better clinical outcome (Figure 1d) compared with the control group.
Clinical Impact of Additional RB1 and TP53 Deletions, and CKS1B Gain in Group 4
A FISH panel targeting CDKN2C/CKS1B, RB1/13q34, and TP53/CEN17 was performed mostly on group 4 patients (Table 3). RB1 deletion was found in 41 of 105 (39%) patients tested in group 4. The overall survival was 45 months, which was lower than the 62 months in those patients (n=64) with a normal RB1, but was not statistically significant (P=0.645; Figure 2a). TP53 deletion was found in 18 of 100 (18%) patients tested and also added no additional impact on overall survival (P=0.60; Figure 2b). There were only 27 cases in group 4 tested for CDKN2C/CKS1B and 23 (85%) were positive for CKS1B gain, which was consistent with the chromosome analysis showing the structural aberrations of 1q gain in most of the cases in this group. The overall survival of this patient group was 43 months, which was significantly worse than the 93 months overall survival in the four patients with a normal CKS1B, however it did not reach statistical significance (P=0.99; Figure 2c).
Kaplan–Meier curves comparing the survival between patients with positive and negative FISH results in group 4. (a) RB1 deletion vs no RB1 deletion. (b) TP53 deletion vs no TP53 deletion. (c) CKS1B gain vs no CKS1B gain. (d) Comparison of survival among patients in group 3 and 4 with no TP53 deletion and patients with a normal karyotype and TP53 (NLKP/TP53-del) deletion by FISH.
Groups 3 and 4 have an Overall Survival Similar to Patients with TP53 Deletion and are a High-risk Cytogenetic Group
To further assess the independent clinical impact of complex additional–structural–chromosomal aberrations, we analyzed 21 patients who were known to be at high risk per current Revised International Staging System, showing an apparently normal karyotype, but with a TP53 deletion (termed NLKP/TP53-del) detected by FISH testing. We compared the overall survival of patients in groups 3 and 4 with this patient subset. The overall survival in the NLKP/TP53-del was 62 months, almost the same as group 3 patients (62 vs 61 months) and better than group 4 patients (62 vs 48 months, P=0.387; Figure 2d). These data indicate that a high complexity hyperdiploid karyotype in multiple myeloma patients (groups 3 and 4) has an independent adverse clinical impact, even without TP53 deletion.
Trisomy 3 or 15, Monosomy 13 or 13q Deletion, and Monosomy 14 Show Independent Clinical Impact in Group 4
We examined the clinical impact of trisomies of 3, 5, 15, and 21, as well as monosomy 13 or 13q deletion and monosomy 14 in each group, particularly in group 4 as it contained most of the cases in the study (Table 4; Figure 3), and since the literature reports on their impact are controversial.16, 17 There were 91 (55%) cases with trisomy 3 in group 4 and 73 (45%) without trisomy 3; the presence of trisomy correlated with a better overall survival (62 vs 43 months, P=0.007; Figure 3a). There were 95 (58%) cases in group 4 with trisomy 5 and 69 (42%) without trisomy 5, however, the overall survival comparison was not significantly different (P=0.114; Figure 3b). Trisomy 15 was found in 110 (67%) cases and the overall survival overall survival was 59 months compared with an overall survival of 30 months in those (n=54) without trisomy 15 in the same group 4 (P<0.0001; Figure 3c), indicating that this specific trisomy might overcome some of the adverse impact in group 4. Trisomy 21 was found in 88 (54%) cases in group 4, but showed no impact on overall survival (P=0.755; Figure 3d).
There were 60 (37%) cases in group 4 with apparent monosomy 13 or 13q deletion by chromosome analysis, and these patients showed a worse overall survival comparing those without monosomy 13 or 13q deletion (n=104) (36 vs 56 months, P=0.047; Figure 3e). Monosomy 14 was observed only in 11 patients who had an overall survival of 30 months compared with the overall survival of 48 months observed in those 153 cases without monosomy 14 (P=0.012; Figure 3f). These results show independent clinical impact of trisomy 3, trisomy 15, and monosomy 13 or 13q deletion and monosomy 14 in group 4 patients, respectively. Additional analysis of these aberrations in groups 1–3 did not show a consistent impact on overall survival (data not shown), perhaps due to limited overall cases numbers analyzed in these groups.
Of 81 newly diagnosed cases in group 4, 48 (59.3%) had trisomy 3, 48 (59.3%) had trisomy 5, 56 (69.1%) had trisomy 15, 46 (56.8%) had trisomy 21, 29 (35.8%) had monosomy 13 or visible 13q deletion, and 7 (8.6) had monosomy 14. We found statistical differences showing a longer median overall survival in patients with trisomy 5 (P=0.0218), trisomy 15 (P=0.0144); and a shorter overall survival in patients with monosomy 13 or 13q deletion (P=0.0303), and a shorter overall survival in patients with monosomy 14 (P=0.0259). However we found no statistical differences on overall survival in patients with trisomy 21 (P=0.384) or trisomy 3 (P=0.0556). (Supplementary Figure 1b–1f).
Stem Cell Transplantation did not Improve Overall Survival in Patients with Hyperdiploid Karyotype Multiple Myeloma
Most patients (n=192) in this study eventually received stem cell transplantation in addition to chemotherapy and/or radiation therapy. When comparing the overall survival between transplanted vs non-transplanted patients in all the four groups, we found no significant difference in survival (P=0.142; Figure 4a–d).
As the patient number for each group is limited, more studies are needed in order to better assess the response of multiple myeloma patients to stem cell transplantation, especially in those who have a highly complex hyperdiploid karyotype as observed in our group 4 (median overall survival 56 months with stem cell transplantation vs 34 months without stem cell transplantation, P=0.053). When we further assessed the clinical impact of stem cell transplantation in group 4 among patients with new diagnosis (n=81), refractory (n=26) and relapsed (n=57) disease respectively, we found stem cell transplantation significantly improved the overall survival in patients with refractory disease (50.5 vs 26 months, P=0.0084) whereas, no impact was seen in new diagnosis (P=0.7275) or relapse (P=0.591) (Supplementary Table 2; Supplementary Figure 2).
Multivariate Analysis on Overall Survival
Additional multivariate analysis was performed for all 284 patients using the SPSS (Table 5). We observed that male gender (HR 0.58, P<0.0001), two additional–structural–chromosomal aberrations (HR 3.3, P<0.0001), three or more additional–structural–chromosomal aberrations (HR 4.2, P<0.0001), disease stage II (HR 0.73, P=0.02), stage III (HR 0.68, P=0.002) and relapsed disease (HR 0.68, P=0.009) significantly impacted overall survival. Age (using the median age of 60 years old, HR 0.90, P=0.321) and stem cell transplantation (HR 1.25, P=0.055), were insignificant. These data further confirm the independent risk prediction of additional–structural–chromosomal aberrations in patients with hyperdiploid karyotype multiple myeloma.
Discussion
Patients with multiple myeloma are currently stratified into cytogenetically standard- and high-risk groups based on the presence or absence of del(17p), t(4;14)/t(14;16)/t(14;20), 1q gain, and non-hyperdiploidy, according to the Revised International Staging System13 and the most recent update by the International Myeloma Working Group.14 In the current study, we further dissected the heterogeneity of the category of multiple myeloma with a hyperdiploid karyotype, a group currently considered to be standard risk. A hyperdiploid karyotype is often characterized by gains of chromosomes 3, 5, 9, 11, and 15 and can be easily detected by conventional cytogenetic and/or FISH analyses.7, 17 With the recognition of the limited potential of plasma cells to divide in short-term bone marrow culture, detection of recurrent cytogenetic aberrations such as gain CKS1B/1q21, t(11;14)(q13;q32), RB1/13q, and TP53/17p deletions largely rely on interphase FISH studies, preferably performed on CD138-enriched plasma cells.9 A hyperdiploid karyotype is therefore reported in about 45–50% of the multiple myeloma cases by FISH analysis.22 In the current study, we detected ~9% of multiple myeloma cases with a hyperdiploid karyotype by the chromosomal analysis, exactly reflecting the technical limitations. Surprisingly, in this study, only a small fraction (12%) of patients with a hyperdiploid karyotype had simple hyperdiploidy showing no additional–structural–chromosomal aberrations. Instead, most (72%) patients showed two or more additional–structural–chromosomal aberrations. In the literature, a hyperdiploid karyotype in multiple myeloma is thought to have a relatively better clinical outcome compared with other standard-risk cytogenetic subgroups.23 In this study, however, patients with a simple hyperdiploid karyotype with no additional–structural–chromosomal aberrations (group 1) did not have a better clinical outcome compared with patients who had normal karyotype multiple myeloma (Figure 1d), suggesting that a hyperdiploid karyotype may not be associated with a better overall survival, as has been proposed previously. Furthermore, our data indicate that patients with high genomic complexity at the chromosomal level, manifested by two or more additional–structural–chromosomal aberrations, had a significantly poorer overall survival.24 Such complex structural aberrations are less likely to be detected by FISH analysis alone even with the coverage of most commonly involved trisomies in multiple myeloma, as FISH can only target limited loci. These data highlight the value of traditional chromosome analysis for risk stratification, whenever feasible clinically, especially for the laboratories that cannot offer the enriched plasma cell FISH studies or genomic profiling, although traditional chromosomal analysis is not required as standard of care.14
In this study, we did not observe differences in demographic and laboratory data among patients in the four groups, with the exception of hemoglobin and albumin levels, which were higher in group 1 patients. In addition, although disease stages or disease status (new diagnosis, relapse, or refractory) were not significantly different among patients in all the four groups, the death rate or fatality was highly associated with the higher level of the chromosomal complexity (Table 2).25 These data indicate that multiple myeloma patients with an apparent hyperdiploid karyotype do not represent a uniform clinicopathological subgroup.
Deletion of 17p13/TP53 is well established as an adverse prognostic factor in multiple myeloma patients. Regardless of the presence of other chromosomal aberrations and/or laboratory results, FISH testing for TP53 has been fully utilized for risk stratification. In the highly complex hyperdiploid karyotype group 4, although most (68.5%) patients were tested for TP53 deletion by FISH, only 18% had TP53 deletion and this finding did not further worsen the overall survival,26 indicating that TP53 deletion may not have independent adverse impact in multiple myeloma patients with a highly complex hyperdiploid karyotype. On the other hand, when comparing the overall survival of patients in groups 3 and group 4 with the group of cases known to be high risk per Revised International Staging System showing TP53 deletion by FISH, but with a normal karyotype (NLKP/TP53-del), group 3 patients showed almost an identical median overall survival, whereas group 4 showed an even worse median overall survival, indicating that genomic complexity in hyperdiploid karyotype multiple myeloma has an independent adverse impact. Therefore, a complex hyperdiploid karyotype should be considered as a high-risk factor in multiple myeloma patients.
Gains of chromosome 1q21/CKS1B gene, although not mentioned in the 2015 Revised International Staging System, have been included as high risk.14 1q21 gain, encoding CKS1B and PSMB4, likely alters control of the cell cycle and promotes cell proliferation and resistance to proteasome inhibitors.27, 28 In group 4 patients, although only 27 patients were tested for CDKN2C/CKS1B by FISH, 23 (85.1%) showed gains of CKS1B, which is consistent with the karyotypic finding of apparent structural aberrations of 1q gain in most of the cases in this group. RB1 deletion was another recurrent finding that has been reported in about 50% of the multiple myeloma cases.29 In 105 cases tested in group 4, about 40% showed RB1 deletion, but this finding had no additional impact,30, 31, 32, 33 as most of them were not visible by conventional chromosome analysis.34 When further evaluated overall survival in those with apparent monosomy 13 or cytogenetically visible 13q deletion in the same group 4, we did observe the negative impact (P=0.047; Figure 3e). To better analyze our data set, we purposely excluded all the hyperdiploid karyotype cases with apparent IGH translocations detected by chromosome and/or FISH analyses, as different IGH fusion partners may yield variable clinical outcomes (ie, adverse vs non-adverse IGH rearrangements) that may have confounded assessment of the clinical impact of hyperdiploid karyotypes with additional–structural–chromosomal aberrations in this study.
Recent studies have suggested that the presence of certain trisomies or monosomy 14 correlates with a more favorable or worse prognosis, respectively, in patients with hyperdiploid karyotype multiple myeloma.7, 16, 17 In this study, we found a significant difference in survival between patients with and without trisomy 3, but only in the highly complex hyperdiploid karyotype group 4, however, not in newly diagnosed patients (Supplementary Figure 1) and in contrast, trisomy 5 showed a longer overall survival in new patients only in group 4 (P=0.0218; Supplementary Figure 1). We also found a difference in median overall survival between patients with vs without trisomy 15 in groups 3 and 4 (P=0.02 and P<0.0001), as well as in new diagnosis in group 4 (P=0.0144); moreover patients with monosomy 14 in group 4 had a shorter overall survival compared with those without (P=0.012; Figure 3f). However, we did not find a difference in overall survival between those cases with or without trisomy 21 in this study, unlike earlier reports.16, 17 Our results indicate that trisomy 3, 5, 15, and monosomy 13 or 13q deletion or monosomy 14 are independent risk factors in the subset of patients with hyperdiploid karyotype multiple myeloma with high complexity at the chromosomal level.
Over 95% of patients in this study were treated after 2002 when the current standard clinical treatment regimen entered widespread use. Of these, 192 (67.6%) patients eventually received a stem cell transplantation. The overall survival between transplanted vs non-transplanted patients showed no significant difference in this study (P=0.142),14, 35, 36, 37 although patients with stem cell transplantation in group 4 seem to show a much better outcomes (overall survival of 56 months with stem cell transplantation vs 34 months without stem cell transplantation, P=0.053), and no overall survival advantage was observed in new diagnosis vs relapse; however stem cell transplantation did improve the overall survival in patients with refractory disease in group 4 (Supplementary Figure 2). More data are needed as stem cell transplantation would have improved the overall survival as previously reported.14
In conclusion, hyperdiploid karyotype multiple myeloma is a cytogenetically heterogeneous group and highly associated with the presence of additional–structural–chromosomal aberrations. The assessment of additional–structural–chromosomal aberrations by traditional chromosomal analysis remains important for risk stratification, whenever clinically feasible, although this is not considered as a ‘routine test’ by the current consensus from the International Myeloma Working Group.14 However, given the technical limitations of traditional chromosome in the analysis of plasma cells, in addition to FISH testing, DNA- or RNA-based genomic profiling,38 gene expression profiling,39 and/or next-generation sequencing40 will facilitate better characterization of this cytogenetic subgroup at the molecular level for accurate risk stratification and personalized clinical management. Meanwhile, we suggest that a hyperdiploid karyotype with 2 or more additional–structural–chromosomal aberrations should be classified as a high-risk parameter in multiple myeloma patients, and should be treated with strategies tailored to the high-risk phenotype.14
References
Sawyer JR . The prognostic significance of cytogenetics and molecular profiling in multiple myeloma. Cancer Genet 2011; 204: 3–12.
Teoh PJ, Chng WJ . p53 abnormalities and potential therapeutic targeting in multiple myeloma. Biomed Res Int 2014; 2014: 1–9.
Prideaux SM, Conway O'Brien E, Chevassut TJ . The genetic architecture of multiple myeloma. Adv Hematol 2014; 2014: 1–16.
Chng WJ, Kumar S, Vanwier S et al, Molecular dissection of hyperdiploid multiple myeloma by gene expression profiling. Cancer Res 2007; 67: 2982–2989.
Li Y, Wang X, Zheng H et al, Classify hyperdiploidy status of multiple myeloma patients using gene expression profiles. PLoS ONE 2013; 8: e58809.
Facon T, Lai JL, Nataf E et al, Improved cytogenetic analysis of bone marrow plasma cells after cytokine stimulation in multiple myeloma: a report on 46 patients. Br J Haematol 1993; 84: 743–745.
Hebraud B, Magrangeas F, Cleynen A et al, Role of additional chromosomal changes in the prognostic value of t(4;14) and del(17p) in multiple myeloma: the IFM experience. Blood 2015; 125: 2095–2100.
Ross FM, Avet-Loiseau H, Ameye G et al, Report from the European Myeloma Network on interphase FISH in multiple myeloma and related disorders. Haematologica 2012; 97: 1272–1277.
Lu G, Muddasani R, Orlowski RZ et al, Plasma cell enrichment enhances detection of high-risk cytogenomic abnormalities by fluorescence in situ hybridization and improves risk stratification of patients with plasma cell neoplasms. Arch Pathol Lab Med 2013; 137: 625–631.
Cook JR, Hartke M, Pettay J et al, Fluorescence in situ hybridization analysis of immunoglobulin heavy chain translocations in plasma cell myeloma using intact paraffin sections and simultaneous CD138 immunofluorescence. J Mol Diagn 2006; 8: 459–465.
Schmidt-Hieber M, Gutierrez ML, Perez-Andres M et al, Cytogenetic profiles in multiple myeloma and monoclonal gammopathy of undetermined significance: a study in highly purified aberrant plasmacells. Haematologica 2013; 98: 279–287.
Bolli N, Avet-Loiseau H, Wedge DC et al, Heterogeneity of genomic evolution and mutational profiles in multiple myeloma. Nat Commun 2014; 5: 2997.
Palumbo A, Avet-Loiseau H, Oliva S et al, Revised International Staging System for multiple myeloma: a report from International Myeloma Working Group. J Clin Oncol 2015; 33: 2863–2869.
Sonneveld P, Avet-Loiseau H, Lonial S et al, Treatment of multiple myeloma with high-risk cytogenetics: a consensus of the International Myeloma WorkingGroup. Blood 2016; 127: 2955–2962.
Lopez-Corral L, Mateos MV, Corchete LA et al, Genomic analysis of high-risk smoldering multiple myeloma. Haematologica 2012; 97: 1439–1443.
Kumar S, Fonseca R, Ketterling RP et al, Trisomies in multiple myeloma: impact on survival in patients with high-risk cytogenetics. Blood 2012; 119: 2100–2105.
Chretien ML, Corre J, Lauwers-Cances V et al, Understanding the role of hyperdiploidy in myeloma prognosis: which trisomies really matter? Blood 2015; 126: 2713–2719.
Chng WJ, Santana-Davila R, Van Wier SA et al, Prognostic factors for hyperdiploid-myeloma: effects of chromosome 13 deletions and IgH translocations. Leukemia 2006; 20: 807–813.
Pawlyn C, Melchor L, Murison A et al, Coexistent hyperdiploidy does not abrogate poor prognosis in myeloma with adverse cytogenetics and may precede IGH translocations. Blood 2015; 125: 831–840.
Rajan AM, Rajkumar SV . Interpretation of cytogenetic results in multiple myeloma for clinical practice. Blood Cancer J 2015; 5: e365.
Shank BR, Brown VT, Schwartz RN . Multiple myeloma maintenance therapy: a review of the pharmacologic treatment. J Oncol Pharm Pract 2015; 21: 36–51.
Wuilleme S, Robillard N, Lode L et al, Ploidy, as detected by fluorescence in situ hybridization, defines different subgroups in multiple myeloma. Leukemia 2005; 19: 275–278.
Bergsagel PL, Mateos MV, Gutierrez NC et al, Improving overall survival and overcoming adverse prognosis in the treatment of cytogenetically high-risk multiple myeloma. Blood 2013; 121: 884–892.
Lim JH, Seo EJ, Park CJ et al, Cytogenetic classification in Korean multiple myeloma patients: prognostic significance of hyperdiploidy with 47-50 chromosomes and the number of structural abnormalities. Eur J Haematol 2014; 92: 313–320.
Laubach J, Garderet L, Mahindra A et al, Management of relapsed multiple myeloma: recommendations of the International Myeloma Working Group. Leukemia 2016; 30: 1005–10017.
Rajkumar SV . Myeloma today: disease definitions and treatment advances. Am J Hematol 2016; 91: 90–100.
Nemec P, Zemanova Z, Greslikova H et al, Gain of 1q21 is an unfavorable genetic prognostic factor for multiple myeloma patients treated with high-dose chemotherapy. Biol Blood Marrow Transplant 2010; 16: 548–554.
Nahi H, Vatsveen TK, Lund J et al, Proteasome inhibitors and IMiDs can overcome some high-risk cytogenetics in multiple myeloma but not gain 1q21. Eur J Haematol 2016; 96: 46–54.
Dowd AA, Homeida S, Elkarem HA . Detection of chromosome 13 (13q14) deletion among Sudanese patients with multiple myeloma using a molecular genetics fluorescent in situ hybridization technique(FISH). Malays J Pathol 2015; 37: 95–100.
Boyd KD, Ross FM, Walker BA et al, Mapping of chromosome 1p deletions in myeloma identifies FAM46C at 1p12 and CDKN2C at 1p32.3 as being genes in regions associated with adverse survival. Clin Cancer Res 2011; 17: 7776–7784.
Ouyang J, Gou X, Ma Y et al, Prognostic value of 1p deletion for multiple myeloma: a meta-analysis. Int J Lab Hematol 2014; 36: 555–565.
Weinhold N, Kirn D, Seckinger A et al, Concomitant gain of 1q21 and MYC translocation define a poor prognostic subgroup of hyperdiploid multiple myeloma. Haematologica 2016; 101: 116–119.
Wu KL, Beverloo B, Lokhorst HM et al, Abnormalities of chromosome 1p/q are highly associated with chromosome 13/13q deletions and are an adverse prognostic factor for the outcome of high-dose chemotherapy in patients with multiple myeloma. Br J Haematol 2007; 136: 615–623.
Kaufmann H, Kromer E, Nosslinger T et al, Both chromosome 13 abnormalities by metaphase cytogenetics and deletion of 13q by interphase FISH only are prognostically relevant in multiple myeloma. Eur J Haematol 2003; 71: 179–183.
Martino M, Lemoli RM, Girmenia C et al, Italian consensus conference for the outpatient autologous stem cell transplantation management in multiple myeloma. Bone Marrow Transplant 2016; 51: 1032–1040.
Awan FT, Flynn JM, Jones JA et al, Phase I dose escalation trial of the novel proteasome inhibitor carfilzomib in patients with relapsed chronic lymphocytic leukemia and small lymphocytic lymphoma. Leuk Lymphoma 2015; 56: 2834–2840.
Kazmi SM, Nusrat M, Gunaydin H et al, Outcomes among high-risk and standard-risk multiple myeloma patients treated with high-dose chemotherapy and autologous hematopoietic stem-cell transplantation. Clin Lymphoma Myeloma Leuk 2015; 15: 687–693.
He J, Abdel-Wahab O, Nahas MK et al, Integrated genomic DNA/RNA profiling of hematologic malignancies in the clinical setting. Blood 2016; 127: 3004–3014.
Hermansen NE, Borup R, Andersen MK et al, Gene expression risk signatures maintain prognostic power in multiple myeloma despite microarray probe set translation. Int J Lab Hematol 2016; 38: 298–307.
Ikeda H, Ishiguro K, Igarashi T et al, Molecular diagnostics of a single drug-resistant multiple myeloma case using targeted next-generation sequencing. Onco Targets Ther 2015; 8: 2805–2815.
Acknowledgements
We wish to thank the technologists in the Clinical Cytogenetics Laboratory at MD Anderson Cancer Center for their contribution. RZO acknowledges support from The MD Anderson Cancer Center SPORE in Multiple Myeloma (P50 CA142509), The MD Anderson Cancer Center Support Grant (P30 CA016672), the National Cancer Institute (U10 CA032102, R01 CA184464, CA194264), and the MD Anderson Cancer Center High Risk Multiple Myeloma Moon Shot.
Author contributions
XL designed the study; AAC-Z and QS collected the data; AAC-Z and XL performed the statistical analysis, AAC-Z, XL, PL and LJM wrote the manuscript; RZO and PL provided treatment data. All authors approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Carballo-Zarate, A., Medeiros, L., Fang, L. et al. Additional–structural–chromosomal aberrations are associated with inferior clinical outcome in patients with hyperdiploid multiple myeloma: a single-institution experience. Mod Pathol 30, 843–853 (2017). https://doi.org/10.1038/modpathol.2017.3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.3
This article is cited by
-
The survival impact of CKS1B gains or amplification is dependent on the background karyotype and TP53 deletion status in patients with myeloma
Modern Pathology (2021)
-
MDM2/X inhibitors under clinical evaluation: perspectives for the management of hematological malignancies and pediatric cancer
Journal of Hematology & Oncology (2017)