Abstract
PTEN loss is a promising prognostic and predictive biomarker in prostate cancer. Because it occurs most commonly via PTEN gene deletion, we developed a clinical-grade, automated, and inexpensive immunohistochemical assay to detect PTEN loss. We studied the sensitivity and specificity of PTEN immunohistochemistry relative to four-color fluorescence in situ hybridization (FISH) for detection of PTEN gene deletion in a multi-institutional cohort of 731 primary prostate tumors. Intact PTEN immunostaining was 91% specific for the absence of PTEN gene deletion (549/602 tumors with two copies of the PTEN gene by FISH showed intact expression of PTEN by immunohistochemistry) and 97% sensitive for the presence of homozygous PTEN gene deletion (absent PTEN protein expression by immunohistochemistry in 65/67 tumors with homozygous deletion). PTEN immunohistochemistry was 65% sensitive for the presence of hemizygous PTEN gene deletion, with protein loss in 40/62 hemizygous tumors. We reviewed the 53 cases where immunohistochemistry showed PTEN protein loss and FISH showed two intact copies of the PTEN gene. On re-review, there was ambiguous immunohistochemistry loss in 6% (3/53) and failure to analyze the same tumor area by both methods in 34% (18/53). Of the remaining discordant cases, 41% (13/32) revealed hemizygous (n=8) or homozygous (n=5) PTEN gene deletion that was focal in most cases (11/13). The remaining 19 cases had two copies of the PTEN gene detected by FISH, representing truly discordant cases. Our automated PTEN immunohistochemistry assay is a sensitive method for detection of homozygous PTEN gene deletions. Immunohistochemistry screening is particularly useful to identify cases with heterogeneous PTEN gene deletion in a subset of tumor glands. Mutations, small insertions, or deletions and/or epigenetic or microRNA-mediated mechanisms may lead to PTEN protein loss in tumors with normal or hemizygous PTEN gene copy number.
Similar content being viewed by others
Main
PTEN is the most commonly lost tumor suppressor gene in prostate cancer1, 2, 3, 4, 5 and is a promising prognostic biomarker for poor clinical outcomes.6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18 As the PTEN gene is almost always lost by genomic deletion of the entire gene in prostate tumors, fluorescence in situ hybridization (FISH) has traditionally been the gold standard assay to detect in situ PTEN loss in tumor tissue. However, the relatively recent availability of reliable rabbit monoclonal antibodies for detection of PTEN protein has enabled the development of highly validated immunohistochemistry protocols to detect PTEN loss in prostate cancer.9, 19 Immunohistochemistry-based detection of PTEN loss in prostate cancer is less expensive and less time-consuming than FISH for the routine screening of prostate tumor specimens, making it easier to adapt to the current pathology workflow for risk assessment in prostate cancer. In addition, because PTEN loss is commonly subclonal and heterogeneous in primary prostate tumors,9, 20, 21, 22 detection of PTEN gene deletion by FISH can be technically challenging in some cases and screening for focal loss may be more easily accomplished by immunohistochemistry. Finally, there is emerging evidence that in addition to genetic deletion, PTEN protein levels may be compromised by mutations in the gene or microRNA- or epigenetic-regulated mechanisms that would not be detectable by FISH.9, 23, 24, 25
We previously optimized and validated a PTEN immunohistochemistry assay for the detection of PTEN loss in prostate cancer specimens,9 and PTEN loss by this assay correlated with increased risk of biochemical recurrence in a case–control cohort of patients undergoing radical prostatectomy12 and with risk of progression and metastasis in two high risk surgical cohorts (though the latter was not significant in multivariate analyses).9, 11 Though originally performed manually, we have recently transferred this assay to a clinical-grade automated immunostaining platform that may be run in any CLIA-certified pathology laboratory. Using this assay, we recently reported that PTEN loss is associated with reduced recurrence-free survival in multivariable models in a multi-institutional cohort of surgically treated patients26 and with higher risk of lethal prostate cancer in a large population-based cohort.18 PTEN gene deletion by FISH has also been recently reported in a subset of the multi-institutional cohort and correlated with recurrence-free survival.17 Here, to analytically validate our clinical-grade PTEN immunohistochemistry assay, we compared the performance of the automated immunohistochemistry assay with PTEN FISH in this cohort, one of the largest multi-institutional cohorts to be studied by both techniques. We demonstrate that our immunohistochemistry assay shows robust sensitivity and specificity for the detection of homozygous PTEN gene deletion.
Materials and methods
Subject Selection and Tissue Microarray Design
The Canary Foundation Retrospective Prostate Tissue Microarray Resource has been described in detail elsewhere.27 Briefly, it is a multicenter, retrospective prostate cancer tissue microarray created as a collaborative effort with radical prostatectomy tissue from six academic medical centers: Stanford University, University of California San Francisco, University of British Columbia, University of Washington (including tissues from University of Washington and a separate cohort from the Fred Hutchinson Cancer Research Center), University of Texas Health Science Center at San Antonio, and Eastern Virginia Medical School. Tumor tissue from 1275 patients was selected for the tissue microarray using a quota sampling plan, from radical prostatectomy specimens collected between 1995 and 2004. A starting date of 1995 was selected to enrich for cases occurring after the implementation of PSA screening. There was no central pathology review in this cohort. The tissue microarray included samples from men with (i) recurrent prostate cancer; (ii) nonrecurrent prostate cancer; and (iii) unknown outcome due to inadequate follow-up time (ie, censoring). Recurrent cases of Gleason score 3+3=6 and 3+4=7 were relatively over-sampled as well as nonrecurrent cases with Gleason score 4+4=8, in order to improve power to detect biomarkers providing prognostic information independent of Gleason score.
Each site built five tissue microarrays, each containing tumor tissue from 42 patients (210 patients from each contributing site). Each tumor was sampled in triplicate, utilizing 1-mm cores and an additional core of histologically benign peripheral zone tissue was included for each patient as a control. Recurrent and nonrecurrent patients were distributed randomly across all tissue microarrays.
Immunohistochemistry Assays
PTEN immunohistochemistry was performed on the CFRPTMR cohort as recently reported.26 Briefly, the protocol uses the Ventana automated staining platform (Ventana Discovery Ultra, Ventana Medical Systems, Tucson, AZ, USA) and a rabbit anti-human PTEN antibody (Clone D4.3 XP; Cell Signaling Technologies, Danvers, MA, USA). We previously validated a manual version of this assay using the same antibody in genetically characterized cell lines and prostate tumor tissue, showing strong correlation of the immunohistochemistry with PTEN gene copy number by two-color FISH and high resolution SNP array analysis9 and good correlation with four-color FISH in a small cohort of needle biopsy specimens.28 To prove equivalence between the manual and automated assays, we also examined a test tissue microarray containing 50 prostate cancer cases with known PTEN protein status (including more than 30 with PTEN protein loss) by manual staining and found 100% concordance between the PTEN protein status on the manual and automated platforms.
Immunohistochemistry Scoring
After staining for PTEN, all tissue microarrays were scanned at × 20 magnification (Aperio, Leica Microsystems, Buffalo Grove, IL, USA) and segmented into TMAJ for scoring (http://tmaj.pathology.jhmi.edu/). PTEN protein status was blindly and independently scored by two trained pathologists (TLL and CLM) using a previously validated scoring system (see below). Overall, there was 'very good' agreement between independent reviewers, with 96% agreement over 2783 cores scored by both reviewers (κ=0.905; 95% CI=0.887–0.923).26
A tissue core was considered to have PTEN protein loss if the intensity of cytoplasmic and nuclear staining was markedly decreased or entirely negative across >10% of tumor cells compared with surrounding benign glands and/or stroma, which provide internal positive controls for PTEN protein expression.9 If the tumor core showed PTEN protein expressed in >90% of sampled tumor glands, the tumor was scored as PTEN intact. If PTEN was lost in <100% of the tumor cells sampled in a given core, the core was annotated as showing heterogeneous PTEN loss in some, but not all, cancer glands (focal loss). Alternatively, if the core showed PTEN loss in 100% of sampled tumor glands, the core was annotated as showing homogeneous PTEN loss. Finally, a small percentage of cores were scored as having ambiguous PTEN immunohistochemistry results. This occurred when the intensity of the tumor cell staining was light or absent in the absence of evaluable internal benign glands or stromal staining. The percent of tissue cores with ambiguous scoring for PTEN immunohistochemistry was fairly constant across six of the seven institutions included in the Canary tissue microarray cohort and varied from 0.7 to 5.3%.26
For statistical analysis, each patient’s tumor sample was scored for the presence or absence of PTEN loss by summarizing the scores of each individual sampled core from that tumor. A patient’s tumor was designated as having heterogeneous PTEN loss if at least one tumor core showed heterogeneous PTEN loss, or alternatively, if at least one core showed heterogeneous or homogeneous PTEN loss and at least one core showed PTEN intact in tumor cells. A patient’s tumor was scored as showing homogeneous PTEN loss if all sampled tumor cores showed homogeneous PTEN loss. Finally, a patient’s tumor was scored as having PTEN intact if all sample tumor cores showed intact PTEN in sampled tumor glands.
Initial Blinded Analysis of PTEN FISH
PTEN FISH was performed as previously described for a subset of this cohort.17 Briefly, the PTEN Del-TECT FISH utilizes a four-color probe combination as described. Probes were supplied by CymoGen Dx LLC (New Windsor, NY, USA) as follows: centromeric copy control probe–CYMO-Pink; WAPAL–CYMO-Green; PTEN–CYMO-Red; and FAS–CYMO-Aqua. We have shown previously that use of the probes bracketing PTEN improves the fidelity of assessments of PTEN loss.29 The two probes WAPAL and FAS on either side of PTEN provide information about the size of larger deletions and also allow recognition of background artifactual losses of PTEN due to histologic sectioning. Artifacts in assessing PTEN loss can arise when histologic sectioning cuts away part of the nucleus containing the PTEN locus in cells in the section while leaving the centromere in place. The latter is a result of the long distance between the centromere and the PTEN locus on chromosome 10.
PTEN FISH analysis was performed entirely independently of PTEN immunohistochemistry, using 5-micron tissue microarray sections stained with DAPI (4′,6-diamidino-2-phenylindole, dihydrochloride) in tumor areas selected by a pathologist who was not involved in PTEN immunohistochemistry scoring (TJ) using an immediately adjacent section stained with hematoxylin and eosin. PTEN copy number was evaluated by counting spots for all four probes using SemRock filters appropriate for the excitation and emission spectra of each dye in 50–100 non-overlapping, intact, interphase nuclei per tumor tissue microarray core. For the initial blinded analysis of each case, two tumor-containing cores were scored based on the overall quality of FISH hybridization. In cases where different clonal deletions were present, all three cores were analyzed and more cells were analyzed. Hemizygous (single copy) PTEN loss was assigned when >50% of nuclei exhibited either interstitial loss of PTEN or concomitant loss of adjacent genes (PTEN and WAPAL and/or FAS). Homozygous deletion was defined by a simultaneous lack of both PTEN locus signals in 30% of scored nuclei.
Immunohistochemistry-Guided Re-analysis of Cases with Discrepant Results by Immunohistochemistry and FISH
Fifty-three cases showed PTEN protein loss by immunohistochemistry with two copies of PTEN gene present by initial FISH analysis (see Results, below). Two cases showed PTEN protein intact by immunohistochemistry with homozygous PTEN deletion by PTEN FISH. To analyze the cause of these discrepancies, we re-examined both the immunohistochemistry and FISH data in these cases. A digitally scanned photomicrograph of the most representative core with immunohistochemistry loss was selected to guide FISH re-analysis of the identical core from each case. As the majority (85%) of these discrepant cases showed only focal immunohistochemistry loss in a subset of glands, the FISH re-analyses concentrated on determining the PTEN gene copy number within these small areas guided by the immunohistochemistry staining. Because only 50–100 cells from the best two of the three tumor-containing cores were initially analyzed for each case by PTEN FISH,29 this more extensive analysis could include tissue microarray cores and regions of tissue microarray sections that had not been studied by FISH during initial blinded analysis.
Immunohistochemistry and FISH on Standard Tissue Sections
To examine possible effects of tumor heterogeneity on immunohistochemistry and FISH interpretation in the setting of tissue microarray cores, we additionally examined 20 cases of varying PTEN status (enriched for discordance between immunohistochemistry and FISH) by FISH and immunohistochemistry on standard tissue sections. Immunohistochemistry and FISH interpretation of these sections were performed blinded to the results of the tissue microarray analysis and the results of the other methodology.
Results
Data for PTEN FISH and immunohistochemistry in a subset of the CFRPTMR cohort were separately reported previously.17, 26 Briefly, of the 1275 patients with tissue sampled for the tissue microarrays, 86% (1095/1275) had evaluable PTEN status by immunohistochemistry and 14% (180/1275) had missing data (Supplementary Table S1). Of these, 17% (30/180) were missing because of ambiguous immunostaining results and 83% (150/180) had absence of tumor tissue present on the tissue microarray slides. Of the tumors with evaluable staining, 24% (258/1095) showed any PTEN protein loss, with 14% (150/1095) showing heterogeneous PTEN loss (in some but not all sampled tumor glands, best exemplified by case #10 in Figure 4a), and 10% (108/1095) showing homogeneous PTEN loss (in all sampled tumor glands). The remaining 76% (837/1095) of cases had intact PTEN protein by immunohistochemistry in all sampled tumor glands. PTEN FISH results were evaluable in 64% of the cases sampled on the tissue microarray (810/1275). Of the evaluable cases, PTEN FISH showed any PTEN deletion in 18% of cases, with 9% (70/810) of cases showing hemizygous deletion and 9% (75/810) of cases showing homozygous PTEN deletion. The remaining 82% (665/810) of cases showed two intact PTEN alleles.
(a) Prostate cancer cases with discordant PTEN immunohistochemistry and FISH results on initial review. Case #9: PTEN immunohistochemistry demonstrates very weak cytoplasmic immunostaining with loss of nuclear immunostaining and thus was called negative on initial review, though in retrospect, it may be better classified as ambiguous because of weak staining and absence of benign glands for comparison (left). Four-color FISH image from an adjacent section that is representative of all examined cores in this tissue microarray (right) indicates that the PTEN gene does not have a detectable deletion by FISH. The enlarged inset shows that the centromeres, WAPAL, PTEN, and FAS gene probes are each present as two copies. Case #10: PTEN immunohistochemistry image (left) shows heterogeneous PTEN loss in some tumor glands (arrow), but PTEN protein is expressed by the majority of other tumor glands in this core. FISH image from an adjacent section (right) was initially read as PTEN intact, but shows a focal area with hemizygous PTEN deletion recognized on re-examination guided by immunohistochemistry. The enlarged inset shows that there is only one copy of the red PTEN gene probe (one red signal) and loss of both aqua FAS gene probes. Case #11: PTEN immunohistochemistry image (left) demonstrates heterogeneous PTEN loss in some tumor glands (arrows) but not in others (arrowheads). FISH image from an adjacent section (right) shows the small area of the section that had a homozygous PTEN deletion on re-examination. The enlarged inset shows that there are no copies of the red PTEN gene probe and one copy of the aqua FAS gene probe, but retention of the adjacent WAPAL and centromere probes. (b) Prostate cancer cases with discordant PTEN immunohistochemistry and FISH results on initial review. Case #12: PTEN immunohistochemistry image (left) shows heterogeneous loss of PTEN protein in some tumor glands (arrow) but not in others (arrowhead). A FISH image from an adjacent section that is representative of all examined cores in this tissue microarray (right) indicates that the PTEN gene does not have a detectable deletion by FISH. The enlarged inset shows that the centromeres, WAPAL, PTEN, and FAS gene probes are each present as two copies. The heterogeneous loss in this case may have resulted in different tumor areas sampled in slides for immunohistochemistry and that for FISH. Case #13: PTEN immunohistochemistry image (left) shows predominantly intact/light immunostaining in tumor glands (arrowhead) and benign glands (b) with a very focal area of tumor with PTEN loss identified on re-review after FISH analysis (arrowhead, inset). FISH analysis of an adjacent section to the immunohistochemistry indicates a homozygous PTEN deletion. The enlarged inset shows that there are no copies of the red PTEN gene probe and loss of one green WAPAL gene probe but retention of both the FAS and the centromere probes.
PTEN immunohistochemistry results were available on 90% of cases with interpretable PTEN FISH results (731/810). The rates of PTEN gene and PTEN protein loss were quite similar in the subset with both FISH and immunohistochemistry results compared with the entire evaluable cohort for each assay reported separately. Overall, 22% (158/731) of cases with interpretable immunohistochemistry and FISH results showed PTEN protein loss, with 13% (96/731) showing heterogeneous loss and 8% (62/731) showing homogeneous loss. Similarly, 17% (129/731) of cases with interpretable immunohistochemistry and FISH results showed PTEN gene deletion (8% hemizygous and 9% homozygous).
Overall, intact PTEN immunohistochemistry was 91% specific for lack of underlying PTEN gene deletion. Of cases with two copies of the PTEN gene by FISH analysis, 549/602 showed intact PTEN protein (Figure 1, Tables 1 and 2). Notably, 85% (45/53) of the discrepant cases (loss of PTEN protein expression by immunohistochemistry and two copies of PTEN gene by FISH analysis) showed heterogeneous PTEN protein loss in some, but not all, sampled tumor glands, suggesting the possibility that a small area with PTEN deletion may have been missed in the initial FISH analysis (see below). PTEN immunohistochemistry loss was 65% sensitive for the detection of underlying hemizygous PTEN gene deletion because 40/62 of cases with hemizygous PTEN gene deletion by FISH showed PTEN protein loss by immunohistochemistry (Figure 2). Of these cases, 65% (26/40) showed heterogeneous PTEN loss in some but not all sampled tumor glands. PTEN immunohistochemistry loss was 97% sensitive for homozygous PTEN gene deletion. Of cases with homozygous gene deletion by FISH, 65/67 showed PTEN protein loss by immunohistochemistry (Figure 3). Only 37% (25/67) of the cases with homozygous PTEN gene deletion and PTEN protein loss had heterogeneous loss of PTEN protein by immunohistochemistry. The fraction of tumors with underlying homozygous PTEN gene deletion differed by the extent of PTEN protein loss observed: 26% (25/96) tumors with heterogeneous PTEN protein loss had an underlying homozygous PTEN deletion compared with 64% (40/62) of tumors with homogeneous PTEN protein loss (P<0.0001 by Fisher’s exact test).
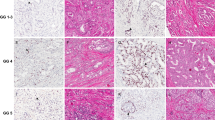
Prostate cancer cases showing intact PTEN protein with two intact PTEN gene alleles. Cases #1 and 2: PTEN immunohistochemistry demonstrates intact PTEN protein (left), while four-color FISH image from adjacent section (right) shows two intact PTEN alleles (see enlarged inset–two red signals) with two intact copies flanking genes WAPAL (green) and FAS (aqua) as well as chromosome 10 centromeres (pink).
Prostate cancer cases showing variable PTEN protein expression with hemizygous PTEN gene deletion. Case #3: PTEN immunohistochemistry demonstrates intact PTEN protein (left), with four-color FISH image from an adjacent section showing a hemizygous PTEN deletion with loss of one PTEN gene (see enlarged inset-one red signal). As both centromeres (pink) and the WAPAL (green) and FAS (aqua) probes that flank either side of PTEN are retained, it is likely that this hemizygous deletion is interstitial and restricted to the PTEN region. Case #4: PTEN immunohistochemistry image shows homogeneous loss of PTEN protein (left) while FISH image from an adjacent section (right) shows a hemizygous PTEN deletion (see enlarged inset–one red signal). Concurrent hemizygous deletion of the adjacent FAS gene probe (one aqua signal missing) but retention of two copies of the centromere and WAPAL gene probes indicates the deletion includes both the PTEN and FAS genes. Case #5: PTEN immunohistochemistry image shows somewhat light, but intact immunostaining for PTEN protein (left) while the FISH image from an adjacent section (right) shows a hemizygous PTEN deletion (see enlarged inset–one red signal). Because there was concurrent loss of the WAPAL, PTEN, and FAS gene probes (green, red, and aqua, respectively), but retention of both centromeres (pink), this hemizygous deletion extends outside the PTEN region in both directions.
Prostate cancer cases showing absence of PTEN protein expression with homozygous PTEN gene deletion. Case #6: PTEN immunohistochemistry image (left) shows loss of PTEN protein in tumor glands. Intraductal spread of tumor is present in this case and retention of PTEN protein is seen in benign basal and luminal cells of duct containing tumor (arrowhead). Four-color FISH image from an adjacent section (right) shows a homozygous deletion with loss of both PTEN genes (see enlarged inset–no red signals). The retention of the centromeres (pink) and both WAPAL genes (green), but the presence of only one copy of the FAS gene (aqua) indicates that one of the deletions involved both the PTEN and FAS genes. Case #7: PTEN immunohistochemistry image (left) shows loss of PTEN protein in tumor glands, with retention in entrapped benign gland (B). FISH image from an adjacent section (right) shows a homozygous PTEN deletion (see enlarged inset–no red signals). The retention of the centromeres (pink) but concurrent loss of one WAPAL (green) and one FAS gene (blue) indicates the deletions extend outside the PTEN region. Case #8: PTEN immunohistochemistry image (left) shows loss of PTEN protein in tumor glands, with retention in adjacent benign gland (B) and nearby endothelial cells (arrowhead). (FISH image from an adjacent section (right) shows a homozygous PTEN deletion (see enlarged inset–no red signals). The retention of the centromeres and both the WAPAL genes (green), but the concurrent loss of both FAS (blue) and PTEN (red), indicates that both copies of chromosome 10 have deletions involving these genes.
The negative predictive value for intact PTEN immunohistochemistry was 96% (549/573) for lack of any gene deletion and 99.6% (571/573) for lack of homozygous PTEN deletion (Table 2). The positive predictive value of PTEN immunohistochemistry loss for presence of any PTEN gene deletion (homozygous or hemizygous) was 66% (105/158) overall, or 53% (51/96) for heterogeneous PTEN protein loss and 87% (54/62) for homogeneous PTEN protein loss (Table 2).
Next, we re-examined cases where there was a discrepancy between the PTEN immunohistochemistry and FISH. Overall, 53 cases with PTEN protein loss had two intact copies of PTEN by FISH, of which 85% (45/53) showed heterogeneous PTEN protein loss. Because only 50–100 tumor cells from two of the three tumor cores from each cases were initially evaluated by FISH, it is possible that focal tumor areas with PTEN gene deletion by FISH were missed or not analyzed in this blinded analysis. To examine this and other possible explanations for the immunohistochemistry-FISH discrepancy, each of these 53 discordant cases were re-reviewed for immunohistochemistry and FISH staining. Immunohistochemistry-guided FISH re-analysis in these cases revealed borderline immunohistochemistry loss in 6% (3/53) cases (Figure 4a, Case #10) and failure to analyze the identical tumor core or area by both immunohistochemistry and FISH in 34% (18/53) cases. Of the remaining discrepant cases where the immunohistochemistry result was convincing and the identical tumor area was analyzed by both methods, 41% (13/32) revealed hemizygous (n=8, Figure 4a, Case #11) or homozygous (n=5, Figure 4a, Case #12) deletion that was focal in 94% (11/13) cases and thus likely missed on initial FISH analysis. The remaining 59% (19/32) of these cases showed two copies of PTEN, thus representing truly discordant cases. One explanation for these cases is the presence of a small deletion and/or mutation undetectable by FISH at one or both PTEN alleles. Another possibility is that even though the same core was evaluated by both methods in these cases, there may be heterogeneity within the core such that different levels of the core sampled on the FISH and immunohistochemistry slide may have been truly heterogeneous (Figure 4b, Case # 13). Of the two discrepant cases with homozygous PTEN deletion and intact PTEN protein, different tumor areas were analyzed in one case. In the other case, a minute focus of tumor with PTEN loss by immunohistochemistry that was initially missed was observed on re-examination (Figure 4b, Case # 14).
Finally, to further assess the effects of tumor heterogeneity on PTEN immunohistochemistry and FISH results and to determine whether this might account for discordance in some cases, we blindly studied a subset of 20 cases from the tissue microarray using standard tissue sections and compared results of immunohistochemistry and FISH on standard sections with one another and with those obtained for the tissue microarray cores of the same cases (Table 3). Cases chosen for this analysis were relatively enriched for discordance between tissue microarray-based immunohistochemistry and FISH results. In cases where the immunohistochemistry and FISH were concordant on the tissue microarray cores, results were generally highly concordant using standard sections as well. For example, in three cases where there was heterogeneous PTEN loss by immunohistochemistry and homozygous PTEN loss by FISH in the tissue microarray cores, two of these tumors had clonal homozygous PTEN deletions, and the third tumor had a region with homozygous loss surrounded by a larger area with PTEN hemizygous loss. Similarly, in four cases that were PTEN intact by both immunohistochemistry and FISH on tissue microarray cores, three showed PTEN intact by FISH on standard sections (the fourth case failed to hybridize) and three showed PTEN intact by immunohistochemistry on standard sections (the fourth case showed focal PTEN loss). In some cases where there was discordance between the immunohistochemistry and FISH results on tissue microarray cores, more detailed analysis of standard sections suggested that tumor heterogeneity may be the underlying cause. Of eight cases with heterogeneous PTEN loss by immunohistochemistry and intact PTEN by FISH on tissue microarray, four showed either hemizygous or homozygous PTEN loss by FISH on standard sections. Another case with homogeneous PTEN loss by immunohistochemistry and intact PTEN by FISH on tissue microarray revealed hemizygous PTEN loss by FISH on analysis of standard sections. Overall, these results support the possibility that underlying tumor heterogeneity is one potential cause of PTEN immunohistochemistry-FISH discordance. Despite this, tissue microarray-based evaluation of tumor PTEN status appears to be highly concordant with standard section analysis in most cases.
Discussion
There is an increasing need for validated prognostic and predictive biomarkers in prostate cancer at both ends of the clinical spectrum. Developing prognostic biomarkers to help select patients who are appropriate for active surveillance as well as predictive biomarkers to guide the application of targeted therapy in metastatic disease remain major areas of unmet clinical need. PTEN has long been a promising marker in both regards, however, until relatively recently, the lack of well-validated antibodies to detect PTEN loss has made it challenging to incorporate into routine pathologic risk assessment protocols or clinical trials of PI3K-targeted agents in prostate cancer. Owing to this difficulty, FISH has historically been used to assess whether PTEN is an effective prognostic biomarker by testing the association of PTEN gene deletion with prostate cancer progression. The results from these studies have consistently shown that PTEN gene deletion is associated with increased Gleason grade and stage in prostate cancer.6, 8, 10, 17, 30, 31 In addition, PTEN gene deletion is associated with prostate cancer progression and death in multivariable models.6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16 Though many of these previous studies have used two-color FISH, there is increasing evidence that four-color probes are better suited to distinguish true gene deletions from sectioning artifacts in interphase FISH studies (Yoshimoto et al, in preparation). Accordingly, our group recently demonstrated that homozygous PTEN deletion by four-color FISH is associated with decreased recurrence-free survival in a subset of the prostate tumor cohort examined in the current study.17
Despite these compelling data, PTEN FISH has not been widely implemented in clinical prostate cancer risk stratification protocols to date for a number of reasons. First, FISH to detect gene deletions is technically challenging, requiring careful probe design29 and rigorous cutoffs to ensure that sectioning artifacts do not result in false calls of deletion. Detection of hemizygous deletions can be particularly challenging when nuclei are overlapping or have been distorted during preparation. Depending on tissue quality and fixation, there may also be difficulties with optimizing protease digestion such that as many as 30–40% of cases cannot be evaluated on the first attempt when using tissue microarrays, though this may be less of an issue for biopsies.17 In large part because it is so technically challenging, FISH is relatively expensive compared with immunohistochemistry, and it has been harder to integrate the daily workflow of pathology laboratories as a reflexive test. Finally, though PTEN is most commonly lost via larger genomic deletions in prostate cancer, as many as 10–20% of cases may have mutations, small insertions, or deletions that are not detectable by FISH, in addition to potential epigenetic and miRNA-mediated mechanisms of PTEN loss.1, 2, 3, 4, 5, 32 To address these challenges, several groups have developed immunohistochemistry assays to query PTEN status in tissue.9, 19, 33 Although a number of such assays have been published, for the most part, these assays have largely been compared with two-color FISH in only small-scale studies with around 100 tumors each.23, 24, 34, 35 In the only large studies to compare immunohistochemistry and FISH, there was only weak (κ=0.5)14 or no significant correlation13 between the assays, suggesting a failure of the immunohistochemistry and/or FISH assay to analytically validate.
We used a commercially available rabbit monoclonal antibody to develop an immunohistochemistry assay to assess PTEN protein loss in prostate cancer and showed that this assay is reasonably sensitive for the detection of PTEN gene deletion by two-color FISH or high density SNP array in prostate cancer samples and shows minimal inter-observer variability in interpretation.9 Similarly, the assay performed well vs four-color FISH in a small cohort of needle biopsy specimens.28 Using this assay, our group previously demonstrated that PTEN protein loss is associated with an increased risk of recurrence and progression in surgically treated cohorts of prostate cancer patients.11, 12
To facilitate clinical use of the assay, we adapted it to the automated Ventana staining platform with clinical-grade reagents suitable for in vitro diagnostic use. This assay was clinically validated in a recent study showing that PTEN loss is associated with increased risk of lethal prostate cancer in a large population-based cohort in multivariable models.18 Despite a 4-category scoring system, the assay has shown high inter-observer reproducibility in a number of cohorts (including the current one), with κ values exceeding 0.9.18, 26 In the current study, we analytically validated this automated assay by comparing it to four-color PTEN FISH across a large multi-institutional cohort of prostate cancer patients. Remarkably, we found that the automated immunohistochemistry assay was 91% specific for two intact copies of the PTEN gene and 97% sensitive for homozygous PTEN gene deletions. This is by far the highest sensitivity and specificity reported for a PTEN immunohistochemistry assay relative to FISH. This improved sensitivity and specificity is in part because of the improved specificity of the automated immunohistochemistry assay vs the manual assay and also because of the improved four-color FISH assay, which uses two PTEN gene-flanking probes, in addition to a centromeric control and a PTEN probe to detect PTEN gene deletions. Surprisingly, the immunohistochemistry assay was also 65% sensitive for detection of hemizygous PTEN gene deletion, suggesting that there is complete protein loss in a large fraction, perhaps even a majority, of apparently hemizygous cases. This is most likely due to truncating mutations (nonsense, frameshift, and splice site mutations) or epigenetic modifications at the second allele that are undetectable by FISH yet lead to protein loss.1, 3, 5, 36 Interestingly, though the prevalence of such mutations in PTEN is below 5% in most prostate tumors, many of these mutations are truncating alterations occurring in cases with hemizygous deletions that would lead to protein loss detectable by immunohistochemistry.1, 2, 3, 4, 5
In addition to the potential increased sensitivity of immunohistochemistry vs FISH for detecting combinations of events including copy loss, point mutations, small insertions and deletions, and epigenetic modifications leading to PTEN inactivation, immunohistochemistry is also very useful for screening for areas of focal PTEN loss. By necessity, PTEN FISH is analyzed at high magnification, examining 50–100 nuclei, which may miss small areas of loss within the sampled tumor. In contrast, immunohistochemistry can be easily screened at low magnification and still afford a nearly cell-by-cell resolution image of PTEN expression. In the current study, in over 40% of cases where PTEN immunohistochemistry detected loss and PTEN FISH was initially read as two copies in the identical tumor core, rescreening the FISH guided by areas of immunohistochemistry loss resulted in detection of small areas with PTEN deletion, initially missed or beneath the cutoff for the FISH scoring. This result, in addition to the high negative predictive value of intact immunohistochemistry for lack of deletion strongly suggests that immunohistochemistry screening for PTEN loss is likely to be an efficient and cost-effective strategy to ascertain PTEN status in tissue sections.
Akin to HER2 assessment in breast, it is ultimately likely that the best protocol will be to perform reflexive FISH on a subset of prostate tumors after initial immunohistochemistry screening. Clearly, in cases with ambiguous immunohistochemistry results (<5%), FISH will have an important role. However, there may also be a role for FISH in cases with heterogeneous loss of PTEN by immunohistochemistry. As in previous cohorts,12 in the current cohort, we found that homogeneous PTEN immunohistochemistry loss was more strongly associated with decreased recurrence-free survival compared with heterogeneous PTEN protein loss in both univariate and multivariate analyses.26 The explanation for why focal PTEN loss is a less potent prognostic indicator than homogeneous loss remains unclear. Homogeneous PTEN loss may be a surrogate indicator for expansion of a single, dominant clone of tumor cells. Alternatively, perhaps loss of PTEN in a larger number of cells increases risk of tumor progression for stochastic reasons. Finally, this result may also be related to the higher prevalence of homozygous PTEN deletion among the cases with homogeneous immunohistochemistry loss, compared with the cases with heterogeneous immunohistochemistry loss (64 vs 26%; P<0.0001 by Fisher’s exact test). Indeed, in the subset of the current cohort where PTEN FISH was correlated with disease outcomes, only homozygous but not hemizygous PTEN loss was associated with decreased recurrence-free survival in multivariate models.17 Thus, it may be that tumors with heterogeneous PTEN protein loss and underlying homozygous PTEN gene deletion have outcomes roughly equivalent to cases with homogeneous PTEN protein loss (the majority of which have homozygous deletion). Though larger case numbers than were included in the current study will be required to formally address this hypothesis, this would suggest that it may be useful to perform reflexive FISH in the case of heterogeneous PTEN protein loss by immunohistochemistry (14% of total cases in current cohort) to determine whether there is underlying homozygous PTEN gene deletion. The FISH could be guided by the immunohistochemistry to focus on areas with protein loss, increasing the sensitivity of the assay in this way.
There are a number of limitations of the current study. Though both FISH and immunohistochemistry were performed on the same tissue microarrays, analysis of all tissue microarray cores was not technically feasible for both methods in all cases and correlation between the two assays was carried out on a tumor-by-tumor rather than core-by-core basis for most cases. Thus, some of the disagreements between FISH and immunohistochemistry likely came about because of tumor heterogeneity, where different areas of the same tumor were being analyzed by each assay, and standard section analysis of a subset of cases largely bears this out. In addition, the gold standard for assessing PTEN gene status is not clear at this point. Though FISH can detect larger deletions, which are the most common mechanism of loss in prostate cancer, it will miss smaller deletions, as well as indels and missense mutations, which may inactivate the gene. Thus, in cases where the same tumor tissue was analyzed, it is impossible to know the true cause of the apparent discrepancies between FISH and immunohistochemistry without using a third methodology such as sequencing to examine for gene alterations that would be missed by FISH (these studies are ongoing in separate cohorts currently). Finally, owing to the relatively small numbers of discordant cases overall, it was not feasible to do a meaningful analysis comparing FISH and immunohistochemistry for prediction of prognosis in these cases, to determine which assay is a better prognostic tool.
In conclusion, in a large multi-institutional cohort of prostate tumors, our immunohistochemistry assay for PTEN loss shows the highest specificity and sensitivity for PTEN gene deletion reported for an immunohistochemistry assay to date. These data strongly suggest that immunohistochemistry is a cost-efficient method to screen for PTEN loss in prostate tumors, requiring ~$100 and a single 4 μm tumor section for assay performance. In cases with ambiguous PTEN immunohistochemistry results or heterogeneous PTEN protein loss, reflexive PTEN FISH may be a useful confirmatory test. This inexpensive, automated, and analytically validated immunohistochemistry assay has already been used to demonstrate the association of PTEN loss with lethal prostate cancer in a large population-based cohort in multivariable models.18 Ultimately, its portability will enable the performance of clinical validation studies on a large number of additional cohorts, credentialing PTEN as a prognostic and potentially predictive biomarker in diverse clinical settings.
References
Berger MF, Lawrence MS, Demichelis F et al. The genomic complexity of primary human prostate cancer. Nature 2011;470:214–220.
Barbieri CE, Baca SC, Lawrence MS et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet 2012;44:685–689.
Grasso CS, Wu YM, Robinson DR et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 2012;487:239–243.
Beltran H, Yelensky R, Frampton GM et al. Targeted next-generation sequencing of advanced prostate cancer identifies potential therapeutic targets and disease heterogeneity. Eur Urol 2013;63:920–926.
Robinson D, Van Allen EM, Wu YM et al. Integrative clinical genomics of advanced prostate cancer. Cell 2015;161:1215–1228.
Yoshimoto M, Cunha IW, Coudry RA et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br J Cancer 2007;97:678–685.
McCall P, Witton CJ, Grimsley S, Nielsen KV, Edwards J . Is PTEN loss associated with clinical outcome measures in human prostate cancer? Br J Cancer 2008;99:1296–1301.
Yoshimoto M, Joshua AM, Cunha IW et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod Pathol 2008;21:1451–1460.
Lotan TL, Gurel B, Sutcliffe S et al. PTEN protein loss by immunostaining: analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin Cancer Res 2011;17:6563–6573.
Krohn A, Diedler T, Burkhardt L et al. Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am J Pathol 2012;181:401–412.
Antonarakis ES, Keizman D, Zhang Z et al. An immunohistochemical signature comprising PTEN, MYC, and Ki67 predicts progression in prostate cancer patients receiving adjuvant docetaxel after prostatectomy. Cancer 2012;118:6063–6071.
Chaux A, Peskoe SB, Gonzalez-Roibon N et al. Loss of PTEN expression is associated with increased risk of recurrence after prostatectomy for clinically localized prostate cancer. Mod Pathol 2012;25:1543–1549.
Steurer S, Mayer PS, Adam M et al. TMPRSS2-ERG fusions are strongly linked to young patient age in low-grade prostate cancer. Eur Urol 2014;66:978–981.
Cuzick J, Yang ZH, Fisher G et al. Prognostic value of PTEN loss in men with conservatively managed localised prostate cancer. Br J Cancer 2013;108:2582–2589.
Liu W, Xie CC, Thomas CY et al. Genetic markers associated with early cancer-specific mortality following prostatectomy. Cancer 2013;119:2405–2412.
Mithal P, Allott E, Gerber L et al. PTEN loss in biopsy tissue predicts poor clinical outcomes in prostate cancer. Int J Pathol 2014;21:1209–1214.
Troyer DA, Jamaspishvili T, Wei W et al. A multicenter study shows PTEN deletion is strongly associated with seminal vesicle involvement and extracapsular extension in localized prostate cancer. Prostate 2015;75:1206–1215.
Ahearn TU, Pettersson A, Ebot EM et al. A prospective investigation of PTEN loss and ERG expression in lethal prostate cancer. J Natl Cancer Inst 2016;108.
Sangale Z, Prass C, Carlson A et al. A robust immunohistochemical assay for detecting PTEN expression in human tumors. Appl Immunohistochem Mol Morphol 2011;19:173–183.
Gumuskaya B, Gurel B, Fedor H et al. Assessing the order of critical alterations in prostate cancer development and progression by IHC: further evidence that PTEN loss occurs subsequent to ERG gene fusion. Prostate Cancer Prostatic Dis 2013;16:209–215.
Bismar TA, Yoshimoto M, Duan Q et al. Interactions and relationships of PTEN, ERG, SPINK1 and AR in castration-resistant prostate cancer. Histopathology 2012;60:645–652.
Krohn A, Freudenthaler F, Harasimowicz S et al. Heterogeneity and chronology of PTEN deletion and ERG fusion in prostate cancer. Mod Pathol 2014;27:1612–1620.
Han B, Mehra R, Lonigro RJ et al. Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod Pathol 2009;22:1083–1093.
Verhagen PC, van Duijn PW, Hermans KG et al. The PTEN gene in locally progressive prostate cancer is preferentially inactivated by bi-allelic gene deletion. J Pathol 2006;208:699–707.
Poliseno L, Salmena L, Zhang J et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010;465:1033–1038.
Lotan TL, Wei W, Morais CL et al. PTEN loss as determined by clinical-grade immunohistochemistry assay is associated with worse recurrence-free survival in prostate cancer. Eur Urol Focus 2016 (in press).
Hawley S, Fazli L, McKenney JK et al. A model for the design and construction of a resource for the validation of prognostic prostate cancer biomarkers: the Canary Prostate Cancer Tissue Microarray. Adv Anat Pathol 2013;20:39–44.
Lotan TL, Carvalho FL, Peskoe SB et al. PTEN loss is associated with upgrading of prostate cancer from biopsy to radical prostatectomy. Mod Pathol 2015;28:128–137.
Yoshimoto M, Ludkovski O, DeGrace D et al. PTEN genomic deletions that characterize aggressive prostate cancer originate close to segmental duplications. Genes Chromosomes Cancer 2012;51:149–160.
Sircar K, Yoshimoto M, Monzon FA et al. PTEN genomic deletion is associated with p-Akt and AR signalling in poorer outcome, hormone refractory prostate cancer. J Pathol 2009;218:505–513.
Reid AH, Attard G, Ambroisine L et al. Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at low and high risk of death from prostate cancer. Br J Cancer 2010;102:678–684.
Poliseno L, Salmena L, Riccardi L et al. Identification of the miR-106b~25 microRNA cluster as a proto-oncogenic PTEN-targeting intron that cooperates with its host gene MCM7 in transformation. Sci Signal 2010;3:ra29.
Ugalde-Olano A, Egia A, Fernandez-Ruiz S et al. Methodological aspects of the molecular and histological study of prostate cancer: focus on PTEN. Methods 2015;77-78:25–30.
Yoshimoto M, Cutz JC, Nuin PA et al. Interphase FISH analysis of PTEN in histologic sections shows genomic deletions in 68% of primary prostate cancer and 23% of high-grade prostatic intra-epithelial neoplasias. Cancer Genet Cytogenet 2006;169:128–137.
Ferraldeschi R, Nava Rodrigues D, Riisnaes R et al. PTEN protein loss and clinical outcome from castration-resistant prostate cancer treated with abiraterone acetate. Eur Urol 2015;67:795–802.
Taylor BS, Schultz N, Hieronymus H et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 2010;18:11–22.
Acknowledgements
Funding for this research was provided in part by the Canary Foundation, the CDMRP Transformative Impact Award (W81XWH-12-PCRP-TIA), a CDMRP Idea Award (W81XWH-13-1-0271) a Prostate Cancer Foundation Young Investigator Award, and a generous gift from Mr. David H Koch. The PTEN Del-TECT™ four-color fluorescent probe used in this study was generously donated by Biocare Medical, Concord, CA. JAS was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) in Brazil.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
TLL has received research funding from Ventana Medical Systems and JAS has consulted for CymoGen Dx, LLC.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Lotan, T., Wei, W., Ludkovski, O. et al. Analytic validation of a clinical-grade PTEN immunohistochemistry assay in prostate cancer by comparison with PTEN FISH. Mod Pathol 29, 904–914 (2016). https://doi.org/10.1038/modpathol.2016.88
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.88
This article is cited by
-
High throughput assessment of biomarkers in tissue microarrays using artificial intelligence: PTEN loss as a proof-of-principle in multi-center prostate cancer cohorts
Modern Pathology (2021)
-
PTEN status assessment in the Johns Hopkins active surveillance cohort
Prostate Cancer and Prostatic Diseases (2019)
-
Precise definition of PTEN C-terminal epitopes and its implications in clinical oncology
npj Precision Oncology (2019)
-
Distinct subtypes of genomic PTEN deletion size influence the landscape of aneuploidy and outcome in prostate cancer
Molecular Cytogenetics (2018)
-
Clinical implications of PTEN loss in prostate cancer
Nature Reviews Urology (2018)