Abstract
Carcinoma of unknown primary, wherein metastatic disease is present without an identifiable primary site, accounts for ~3–5% of all cancer diagnoses. Despite the development of multiple diagnostic workups, the success rate of primary site identification remains low. Determining the origin of tumor tissue is, thus, an important clinical application of molecular diagnostics. Previous studies have paved the way for gene expression-based tumor type classification. In this study, we have established a comprehensive database integrating microarray- and sequencing-based gene expression profiles of 16 674 tumor samples covering 22 common human tumor types. From this pan-cancer transcriptome database, we identified a 154-gene expression signature that discriminated the origin of tumor tissue with an overall leave-one-out cross-validation accuracy of 96.5%. The 154-gene expression signature was first validated on an independent test set consisting of 9626 primary tumors, of which 97.1% of cases were correctly classified. Furthermore, we tested the signature on a spectrum of diagnostically challenging tumors. An overall accuracy of 92% was achieved on the 1248 tumor specimens that were poorly differentiated, undifferentiated or from metastatic tumors. Thus, we have identified a 154-gene expression signature that can accurately classify a broad spectrum of tumor types. This gene panel may hold a promise to be a useful additional tool for the determination of the tumor origin.
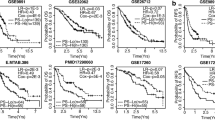
Similar content being viewed by others
Main
Cancer of unknown primary, also known as occult primary tumors, is a heterogeneous group of tumors whose primary site cannot be found when the cancer has metastasized.1 Per 100 000 individuals, the incidence varies from 5 to 7 cases in Europe, 7 to 12 cases in the USA, and 18 to 19 cases in Australia.2 The latest data show that cancer of unknown primary accounts for ~3–5% of all newly diagnosed cancers,2 and it is the fourth leading cause of cancer-related death worldwide.3, 4 Generally, the prognosis of patients with carcinoma of unknown primary site is poor for those receiving empiric chemotherapy. The median survival period is 3–9 months, even when newer combination treatment regiments are administered.5 Hence, cancer of unknown primary remains an important clinical problem that generates frustration among surgeons, oncologists, and pathologists, in addition to the uncertainty and stress it imposes on patients. Identification of the primary site can ease the patient’s anxiety and improve long-term survival with the help of more specific therapy.2, 6
In current clinical practice, patients with carcinoma of unknown primary should inform doctors of their medical history and receive detailed physical examination, laboratory testing, digital imaging, and endoscopic examination. Positron emission tomography–computed tomography, the most efficient imaging test to depict the tumor tissue of origin, can only detect 24–53% of primary lesions of cancer of unknown origin.7 Histological examination, particularly immunohistochemistry, is the cornerstone to identify the tumor of origin. However, even with the best experts and the most advanced technology, the primary site can be identified in only 20–30% of patients with cancer of unknown primary,8 and the results can be subjective.
This clinical need has resulted in a quest for better and more accurate identification of the primary site of tumors. To address this need, several studies have demonstrated that the expression levels of tens to hundreds of genes can be used as a ‘molecular fingerprint’ to classify a multitude of tumor types. Varadhachary et al9 and Talantov et al10 presented a reverse transcription polymerase chain reaction-based method that measures the expression of 10 signature genes among six tumor types. Ma et al11 developed a similar method based on 92 genes to classify 32 tumor types. Tothill et al12 reported a 79-gene panel to discriminate among 13 tumor types. Instead of measuring conventional gene expression, Rosenfeld et al13 analyzed microRNA expression to classify tumor samples.
With the rapid evolution of microarray technology over the last decade, there have been tremendous efforts invested in the field of cancer research using standardized genome-wide microarrays. Considering the large amount of high-quality, publicly available gene expression data sets, the integrative analysis of genomic data, in which data from multiple studies are combined to increase the sample size and avoid laboratory-specific bias, has the potential to yield new biological insights that are not possible from a single study.14
In the present study, we established a comprehensive gene expression database containing the genome-wide expression profiles of more than 16 000 tumor samples representing 22 common human cancer types. By using an innovative analytical method, we aimed to develop a gene expression signature to aid in the identification of tumor origin.
Materials and methods
Sample Collection and Data Curation
The gene expression data sets of 16 674 tumor samples with histologically confirmed origins were collected from public data repositories (eg, ArrayExpress, Gene Expression Omnibus, and The Cancer Genome Atlas Data Portal) and curated to form a comprehensive pan-cancer transcriptome database.
Array-based gene expression profiling of 7048 tumor samples was mainly conducted on three different platforms of Affymetrix oligonucleotide microarray: GeneChip Human Genome U133A Array, U133A 2.0 Array, and U133 Plus 2.0 Array. Data from raw CEL files were pre-processed using the single-channel array normalization method with default parameters. Although different opinions exist concerning data pre-processing, the single-channel array normalization method was considered as most suitable for personalized-medicine workflows. Rather than processing microarray samples as groups, which can introduce biases and present logistical challenges, the single-channel array normalization method can normalize each sample individually by modeling and removing probe- and array-specific background noise using only internal array data.15 We further used the alternative CDF files from BrainArray Resource (http://brainarray.mbni.med.umich.edu/) to summarize the probe level intensities directly to the Entrez gene IDs. Probes mapping to multiple genes and other problems associated with old generations of Affymetrix probe designs were thereby excluded.16
Sequencing-based gene expression profiling of 9626 tumor samples were generated on the Illumina HiSeq 2000 RNA sequencing platform and kindly provided by The Cancer Genome Atlas pan-cancer analysis working group at Synapse website (https://www.synapse.org/).17 The gene expression profile consists of transcriptomic data for 20 501 unique genes. The clinical information for selected samples was retrieved from the ‘Clinical Biotab’ section of the data matrix based on the Biospecimen Core Resource IDs of the patients.
Gene Signature Identification
Gene expression data analysis was performed using R software and packages from the Bioconductor project.18, 19, 20 To identify a gene expression signature, we used the support vector machine—recursive feature elimination algorithm for feature selection and classification modeling.21 For multi-class classification, a one-versus-all approach was used whereby multiple binary classifiers are first derived for each tumor type. The results are reported as a series of probability scores for each of the 22 tumor types. The probability score was estimated as an indicator of the certainty of a classification made by the gene expression signature. The probability score ranges from 0 (low certainty) to 100 (high certainty) and sum to 100 across the 22 primary tumor types. A threshold of probability score equal to 50 was established to indicate the confidence of a single classification. When the probability score fell below 50, the samples were considered ‘unclassifiable cases’. When the probability score was above 50, the tumor type with the highest probability score was considered the tumor of origin. An example of gene expression signature classification is shown in the Supplementary Figure 1.
Signature Performance Assessment
For each specimen, the predicted primary site of the tumor was compared with the reference diagnosis. A true-positive result was indicated when the predicted tumor type matched the reference diagnosis. When the predicted tumor type and reference diagnosis did not match, the specimen was considered a false positive. For each tissue on the panel, sensitivity was defined as the ratio of true-positive results to the total positive samples analyzed, while specificity was defined as the ratio (1−false positive)/(total tested−total positive). The diagnostic odds ratio was calculated as a combination of the sensitivity and specificity as described by Glas et al.22
Results
Establishment of Pan-Cancer Transcriptome Database
To create a cancer transcriptome database for tumor primary site identification, the following issues were primarily considered. First, our database should span the tumor sites to be as large as possible. Second, within each tumor type, all possible histological subtypes should be covered. In addition, to mimic the performance of the candidate gene expression signature to identify the tumor origin in carcinoma of unknown primary, metastatic cancers, poorly differentiated tumors, and undifferentiated tumors should also be included. Thus, a systematic search of major biological data repositories—eg, ArrayExpress, Gene Expression Omnibus, and The Cancer Genome Atlas project—was performed to collect the gene expression profiling data sets of different tumor types.
Overall, we accumulated the gene expression profiles of 16 674 tumor samples to form a comprehensive pan-cancer transcriptome database. The carcinomas originated from 22 major tissue types, including adrenal gland, brain, breast, cervix, colorectal, endometrium, gastroesophagus, head and neck, kidney, liver, lung, lymphoma, melanoma, mesothelioma, neuroendocrine, ovary, pancreas, prostate, sarcoma, testis, thyroid, and urinary. The database also contains patient demographic data and clinical information. To identify a reliable gene expression signature, we adopted a training-validation approach in this study. First, the gene expression profiles of 5800 primary tumors with histologically confirmed origins were retrieved from the database and curated to form a large training set. Next, two independent validation sets were formed: one is composed of sequencing-based gene expression profiles of 9626 tumor specimens with histologically confirmed origins (test set 1) and the other is composed of gene expression profiles of 1248 tumor specimens that were poorly differentiated, undifferentiated or from metastatic tumors (test set 2). Figure 1 depicts three different phases of our study design and Table 1 summarizes the clinical characteristics of the samples in the study.
Gene Selection and Functional Annotation
The training set consisted of 5800 samples covering more than 95% of solid tumors by incidence, with 55–542 specimens per tumor class that encompass a range of intratumor heterogeneity. After data normalization and annotation steps, a matrix of 12 000 unique genes in 5800 samples (≈70 million data points) was prepared for downstream bioinformatics analyses. Extracting a subset of informative genes from such high-dimension genomic data is a critical step for gene expression signature identification. Although many algorithms have been developed, the support vector machine—recursive feature elimination approach is considered one of the best gene selection algorithms. For each tumor type, we used the support vector machine—recursive feature elimination approach to: (1) evaluate and rank the contributions of each gene toward the optimal separation of a specific cancer type from other tumor types; (2) select the top 10-ranked genes as the most differentially expressed genes for this tumor type; and (3) repeat this process for each tumor types, and obtain 22 lists of the top 10 gene set. After removing redundant features, 154 unique genes were obtained. Full list of the 154 candidate genes with respect to each tumor types were provided in Table 2.
We further investigated whether these candidate genes revealed biological features known to be relevant to different cancers. Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis was performed using the GeneCodis bioinformatics tool (http://genecodis.dacya.ucm.es/).23 As shown in Table 3, a diverse group of gene families is represented in the 154-gene list. The most significantly enriched gene categories are those involved in specific biological processes, including tyrosine metabolism, fat digestion and absorption, cytokine–cytokine receptor interaction, extracellular matrix–receptor interaction, and gastric acid secretion. Even more interestingly, genes described in oncogenic pathways such as those of bladder cancer, melanoma, and prostate cancer were also significantly overrepresented, reflecting their differential involvement in a range of tumor classes.
Leave-One-Out Internal Cross-Validation
As an initial step, we assessed the performance of the classifier using leave-one-out cross-validation within the training set. Leave-one-out cross-validation simulates the performance of a classification algorithm on unseen samples. With leave-one-out cross-validation, the algorithm is repeatedly retrained, leaving out one sample in each round and testing each sample on a classifier that was trained without this sample. The 154-gene expression signature showed an overall accuracy of 96.5% (5597 of 5800; 95% CI 96.0 to 97.0%) with notable variation between different cancer types. Sensitivities ranged from 89.7% (endometrium) to 100% (neuroendocrine). Using this internal validation of the training set, these data provide a preliminary estimate of classification performance.
Independent Validation in Primary Tumors Profiled with Next-Generation Sequencing
The final classification model of the 154-gene expression signature was established using the entire training set and then applied to an independent validation set comprising 9626 primary tumor samples profiled with next-generation sequencing (test set 1). Representation from 22 sites ranged from 48 (lymphoma) to 1218 (breast). The 154-gene expression signature estimated 9100 (94.5%) of 9626 samples with probability scores above 50 as ‘valid classification’. Among these 9100 valid cases, the 154-gene expression signature showed 97.1% overall agreement with the reference diagnosis (8839 of 9100; 95% CI 96.8 to 97.5%). Figure 2 shows a matrix of the relationship of the test results compared with the reference diagnoses. Sensitivities for the 22 main cancer types ranged from 84.2% (gastroesophagus) to 100% (prostate). Specificities ranged from 99.4% (gastroesophagus) to 100% (mesothelioma, neuroendocrine and thyroid). The detailed sensitivity and specificity are listed in Table 4. A total of 526 cases (5.5%) were considered ‘unclassifiable’ by the 154-gene expression signature, with probability scores below 50. Cervix, urinary, sarcoma, head and neck, gastroesophagus, and endometrium were the most common biopsy sites among those unclassifiable cases. Diagnostic odds ratios for all the 22 tumor types were significantly >1, indicating that each class reported by the 154-gene expression signature provides significant discrimination and performance.
Confusion matrix by tumor type of the test set 1. Reference diagnoses are shown across the top row, and 154-gene expression signature predictions are shown along the left-hand column. The matrix shows the direct relationship between each adjudicated reference diagnosis versus the molecular classifier prediction, including reproducible patterns of classification and misclassification.
Independent Validation in Metastatic and Undifferentiated Tumors
The 154-gene expression signature was further validated in the test set 2 comprising 1248 tumor specimen samples. For the test set 2, we particularly enriched for tumor metastatic specimens with known primary sites or primary tumors with poor differentiation because these probably reflect the clinical circumstance of carcinoma of unknown primary. Representation from 22 sites ranged from 12 (thyroid) to 216 (sarcoma). The 154-gene expression signature estimated 1077 (86.3%) of 1248 samples with probability scores above 50 as ‘valid classification’. Among these 1077 valid cases, the 154-gene expression signature showed 92% overall agreement with the reference diagnosis (991 of 1077; 95% CI 90.2 to 93.6%). Figure 3 shows a matrix of the relationship of the test results compared with the reference diagnoses. Sensitivities for the 22 main tumor types ranged from 38.9% (pancreas) to 100% (adrenal, brain, head and neck, liver, neuroendocrine, and testis). Specificities ranged from 98.0% (lung) to 100% (adrenal, brain, cervix, mesothelioma, neuroendocrine, pancreas, and prostate). The detailed sensitivity and specificity are listed in Table 4. One hundred seventy-one (13.7%) cases were considered ‘unclassifiable’ by the 154-gene expression signature, with probability scores below 50. Prostate, kidney, pancreas, urinary, adrenal, and melanoma were the most common biopsy sites among those unclassifiable cases. Diagnostic odds ratios for all 22 tumor types were significantly >1.
Confusion matrix by tumor type of the test set 2. Reference diagnoses are shown across the top row, and 154-gene expression signature predictions are shown along the left-hand column. The matrix shows the direct relationship between each adjudicated reference diagnosis versus the molecular classifier prediction, including reproducible patterns of classification and misclassification.
Discussion
Owing to great advancements in high-throughput microarray technologies and the comprehensive efforts of systematic cancer genomics projects, we were able to utilize large genomic data sets for our study. We report here the creation of a pan-cancer gene expression database from more than 160 000 human tumor samples and demonstrate that multi-class tumor classification is feasible by comparing an unknown sample to this reference database. The 154-gene expression signature demonstrated an overall accuracy of 96.5% for 22 tumor types by cross-validation of the training set, and 97.1% in an independent test set of 9626 primary tumors profiled with the next-generation sequencing. Furthermore, we tested the signature on a spectrum of diagnostically challenging tumors. An overall accuracy of 92% was achieved on the 1248 tumor specimens that were poorly differentiated, undifferentiated, or from metastatic tumors.
Several investigations have reported multigene algorithms and results that demonstrate the promise of gene expression-based signatures in tumor origin identification. Unlike many studies in which samples were often dominated by well-differentiated primary cancers, our approach directly exploited undifferentiated metastatic tumor samples for the validation of our 154-gene expression signature. In a clinical scenario, the uncertainty of tumors’ origin usually arises within the context of metastatic and/or poorly differentiated to undifferentiated malignancies, and some of the previously published gene expression-based signatures have shown decreased performance with less-differentiated tumors. In this study, we show that the 154-gene expression signature could reliably identify the tumor origin in 92% of the 1077 tumor samples tested. This accuracy is comparable to other gene expression-based signatures with reported accuracies in the range of 79–91%.24, 25, 26 The performance of this test also compares favorably with current clinical practice standards such as immunohistochemistry, which has shown 75% accuracy in metastatic samples using a predetermined panel of 10 antibodies.27
It is noteworthy that the expression patterns of several genes among the 154-gene panel have been observed previously by other methods to be relatively tissue specific for certain types of carcinomas—eg, KLK3 has been identified as the gene encoding prostate-specific antigen, which has long been known as an important tumor marker used in the diagnosis and monitoring of prostate cancer. Originally, it was thought that prostate-specific antigen was only produced by the cells of the prostate gland. Recently, it has been shown that elevated levels of prostate-specific antigen are also observed in some breast and gynecologic cancers.28, 29 In addition, overexpression of the EGFR gene occurs across a wide range of different cancers, including brain, colorectal, lung, esophageal, cervical cancers, and sarcoma.30, 31, 32, 33, 34, 35 CDH1 and VEGFA have been reported among the highly significant markers in colorectal, gastric, and liver cancers.36, 37, 38, 39, 40, 41
The 154-gene expression signature shows clear promise in identifying the tumor’s origin, but it is not perfect. For diagnostically challenging tumors, systematic errors were noted in the classes of endometrial and pancreatic tumors (58 and 61% misclassified, respectively). Among the seven misclassified endometrial cancers, five were predicted to be ovarian cancer. Given the current controversies over the ontogeny of female genital tract cancers,42, 43, 44, 45 molecular profiling with the 154-gene expression signature may reflect this biologic intersection and provide additional insight into the origin of these tumors. Among the 11 misclassified pancreatic cancers, six were predicted to have originated from the gastroesophagus, and four from the liver. It is known that pancreatic cancer has a complex and heterogeneous genetic base, which is often identified as esophageal cancer.46 Indeed, pancreatic cancer is the most difficult type of carcinoma of unknown primary to identify using our method as well as all published methods.8, 24, 47, 48, 49, 50
Additional research is needed to successfully translate the 154-gene signature from gene expression microarray to real-time reverse transcription polymerase chain reaction assays, thus allowing broader access and utilization in the clinical setting. In routine practice, most diagnostic materials are formalin-fixed and paraffin-embedded; thus, it will be highly interesting to assess the usefulness of the 154-gene signature in formalin-fixed and paraffin-embedded samples. Future translational research should focus on the development and validation of the real-time polymerase chain reaction-based gene expression test using formalin-fixed and paraffin-embedded samples.
In conclusion, this study describes the development and validation of a gene expression-based signature to assist in the identification of the origin of tumor tissue. We foresee its application in cases of poorly differentiated or undifferentiated metastatic tumors and in cases where histology alone fails to suggest a specific primary site of origin. Further studies evaluating the impact of gene expression-based test results on therapy choice and treatment outcome for patients with carcinoma of unknown primary are warranted.
References
Stella GM, Senetta R, Cassenti A et al. Cancers of unknown primary origin: current perspectives and future therapeutic strategies. J Transl Med 2012;10:12.
Richardson A, Wagland R, Foster R et al. Uncertainty and anxiety in the cancer of unknown primary patient journey: a multiperspective qualitative study. BMJ Support Palliat Care 2015;5:366–372.
Pavlidis N, Fizazi K . Cancer of unknown primary (CUP). Crit Rev Oncol Hematol 2005;54:243–250.
Kamposioras K, Pentheroudakis G, Pavlidis N . Exploring the biology of cancer of unknown primary: breakthroughs and drawbacks. Eur J Clin Invest 2013;43:491–500.
Kurahashi I, Fujita Y, Arao T et al. A microarray-based gene expression analysis to identify diagnostic biomarkers for unknown primary cancer. PLoS One 2013;8:e63249.
Hyphantis T, Papadimitriou I, Petrakis D et al. Psychiatric manifestations, personality traits and health-related quality of life in cancer of unknown primary site. Psychooncology 2013;22:2009–2015.
Reske SN, Kotzerke J . FDG-PET for clinical use. Results of the 3rd German Interdisciplinary Consensus Conference, ‘Onko-PET III’, 21 July and 19 September 2000. Eur J Nucl Med 2001;28:1707–1723.
Horlings HM, van Laar RK, Kerst JM et al. Gene expression profiling to identify the histogenetic origin of metastatic adenocarcinomas of unknown primary. J Clin Oncol 2008;26:4435–4441.
Varadhachary GR, Talantov D, Raber MN et al. Molecular profiling of carcinoma of unknown primary and correlation with clinical evaluation. J Clin Oncol 2008;26:4442–4448.
Talantov D, Baden J, Jatkoe T et al. A quantitative reverse transcriptase-polymerase chain reaction assay to identify metastatic carcinoma tissue of origin. J Mol Diagn 2006;8:320–329.
Ma XJ, Patel R, Wang X et al. Molecular classification of human cancers using a 92-gene real-time quantitative polymerase chain reaction assay. Arch Pathol Lab Med 2006;130:465–473.
Tothill RW, Kowalczyk A, Rischin D et al. An expression-based site of origin diagnostic method designed for clinical application to cancer of unknown origin. Cancer Res 2005;65:4031–4040.
Rosenfeld N, Aharonov R, Meiri E et al. MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol 2008;26:462–469.
Rhodes DR, Yu J, Shanker K et al. Large-scale meta-analysis of cancer microarray data identifies common transcriptional profiles of neoplastic transformation and progression. Proc Natl Acad Sci USA 2004;101:9309–9314.
Piccolo SR, Sun Y, Campbell JD et al. A single-sample microarray normalization method to facilitate personalized-medicine workflows. Genomics 2012;100:337–344.
Dai M, Wang P, Boyd AD et al. Evolving gene/transcript definitions significantly alter the interpretation of GeneChip data. Nucleic Acids Res 2005;33:e175.
Omberg L, Ellrott K, Yuan Y et al. Enabling transparent and collaborative computational analysis of 12 tumor types within The Cancer Genome Atlas. Nat Genet 2013;45:1121–1126.
Ihaka R, Robert GR . A language for data analysis and graphics. J Comput Graph Stat 1996;5:299–314.
Reimers M, Carey VJ . Bioconductor: an open source framework for bioinformatics and computational biology. Methods Enzymol 2006;411:119–134.
Chang C, Lin C . LIBSVM: a library for support vector machines. Acm Trans Intell Syst Technol 2011;2:21–27.
Guyon I, Weston J, Barnhill S et al. Gene selection for cancer classification using support vector machines. Mach Learn 2002;46:389–422.
Glas AS, Lijmer JG, Prins MH et al. The diagnostic odds ratio: a single indicator of test performance. J Clin Epidemiol 2003;56:1129–1135.
Tabas-Madrid D, Nogales-Cadenas R, Pascual-Montano A . GeneCodis3: a non-redundant and modular enrichment analysis tool for functional genomics. Nucleic Acids Res 2012;40:W478–W483.
Monzon FA, Lyons-Weiler M, Buturovic LJ et al. Multicenter validation of a 1,550-gene expression profile for identification of tumor tissue of origin. J Clin Oncol 2009;27:2503–2508.
Kerr SE, Schnabel CA, Sullivan PS et al. Multisite validation study to determine performance characteristics of a 92-gene molecular cancer classifier. Clin Cancer Res 2012;18:3952–3960.
Weiss LM, Chu P, Schroeder BE et al. Blinded comparator study of immunohistochemical analysis versus a 92-gene cancer classifier in the diagnosis of the primary site in metastatic tumors. J Mol Diagn 2013;15:263–269.
Park SY, Kim BH, Kim JH et al. Panels of immunohistochemical markers help determine primary sites of metastatic adenocarcinoma. Arch Pathol Lab Med 2007;131:1561–1567.
Mashkoor FC, Al-Asadi JN, Al-Naama LM . Serum level of prostate-specific antigen (PSA) in women with breast cancer. Cancer Epidemiol 2013;37:613–618.
Kucera E, Kainz C, Tempfer C et al. Prostate specific antigen (PSA) in breast and ovarian cancer. Anticancer Res 1997;17:4735–4737.
Devarakonda S, Morgensztern D, Govindan R . Genomic alterations in lung adenocarcinoma. Lancet Oncol 2015;16:e342–e351.
Furnari FB, Cloughesy TF, Cavenee WK et al. Heterogeneity of epidermal growth factor receptor signalling networks in glioblastoma. Nat Rev Cancer 2015;15:302–310.
Giampieri R, Aprile G, Del Prete M et al. Beyond RAS: the role of epidermal growth factor receptor (EGFR) and its network in the prediction of clinical outcome during anti-EGFR treatment in colorectal cancer patients. Curr Drug Targets 2014;15:1225–1230.
Teng HW, Wang HW, Chen WM et al. Prevalence and prognostic influence of genomic changes of EGFR pathway markers in synovial sarcoma. J Surg Oncol 2011;103:773–781.
Li Q, Tang Y, Cheng X et al. EGFR protein expression and gene amplification in squamous intraepithelial lesions and squamous cell carcinomas of the cervix. Int J Clin Exp Pathol 2014;7:733–741.
Li JC, Zhao YH, Wang XY et al. Clinical significance of the expression of EGFR signaling pathway-related proteins in esophageal squamous cell carcinoma. Tumor Biol 2014;35:651–657.
Li YX, Lu Y, Li CY et al. Role of CDH1 promoter methylation in colorectal carcinogenesis: a meta-analysis. DNA Cell Biol 2014;33:455–462.
Jing H, Dai F, Zhao C et al. Association of genetic variants in and promoter hypermethylation of CDH1 with gastric cancer. Medicine (Baltimore) 2014;93:e107.
Liu F, Li H, Chang H et al. Identification of hepatocellular carcinoma-associated hub genes and pathways by integrated microarray analysis. Tumori 2015;101:206–214.
Angelescu C, Burada F, Ioana M et al. VEGF-A and VEGF-B mRNA expression in gastro-oesophageal cancers. Clin Transl Oncol 2013;15:313–320.
Zhang H, Yang R . Resveratrol inhibits VEGF gene expression and proliferation of hepatocarcinoma cells. Hepatogastroenterology 2014;61:410–412.
Kjaer-Frifeldt S, Fredslund R, Lindebjerg J et al. Prognostic importance of VEGF-A haplotype combinations in a stage II colon cancer population. Pharmacogenomics 2012;13:763–770.
Samartzis EP, Noske A, Dedes KJ et al. ARID1A mutations and PI3K/AKT pathway alterations in endometriosis and endometriosis-associated ovarian carcinomas. Int J Mol Sci 2013;14:18824–18849.
Seidman JD, Zhao P, Yemelyanova A . ‘Primary peritoneal’ high-grade serous carcinoma is very likely metastatic from serous tubal intraepithelial carcinoma: assessing the new paradigm of ovarian and pelvic serous carcinogenesis and its implications for screening for ovarian cancer. Gynecol Oncol 2011;120:470–473.
Kurman RJ, Shih IeM . Molecular pathogenesis and extraovarian origin of epithelial ovarian cancer—shifting the paradigm. Hum Pathol 2011;42:918–931.
Wiegand KC, Shah SP, Al-Agha OM et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N Engl J Med 2010;363:1532–1543.
Jones S, Zhang X, Parsons DW et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 2008;321:1801–1806.
Ojala KA, Kilpinen SK, Kallioniemi OP . Classification of unknown primary tumors with a data-driven method based on a large microarray reference database. Genome Med 2011;3:63.
Monzon FA, Medeiros F, Lyons-Weiler M et al. Identification of tissue of origin in carcinoma of unknown primary with a microarray-based gene expression test. Diagn Pathol 2010;5:3.
van Laar RK, Ma XJ, de Jong D et al. Implementation of a novel microarray-based diagnostic test for cancer of unknown primary. Int J Cancer 2009;125:1390–1397.
Dumur CI, Lyons-Weiler M, Sciulli C et al. Interlaboratory performance of a microarray-based gene expression test to determine tissue of origin in poorly differentiated and undifferentiated cancers. J Mol Diagn 2008;10:67–77.
Acknowledgements
The results shown here are, in part, based on data from multiple previously published studies. We acknowledge the investigators and patients who contributed to the acquisition and analysis of the data used in this study. This work was partially supported by research funding from National Natural Science Foundation of China (Grant no. 81472220), Shanghai Science and Technology Development Fund (the Domestic Science and Technology Cooperation Project, No. 14495800300) and Canhelp Genomics. We thank Yang Yang, Xinming Zhang, Yi Cai, and Minzhe Fang for excellent technical and operational assistance.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
QX and JC are employees of Canhelp Genomics. No other potential conflicts of interest were disclosed by the authors.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Xu, Q., Chen, J., Ni, S. et al. Pan-cancer transcriptome analysis reveals a gene expression signature for the identification of tumor tissue origin. Mod Pathol 29, 546–556 (2016). https://doi.org/10.1038/modpathol.2016.60
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.60
This article is cited by
-
Cancer of unknown primary (CUP) through the lens of precision oncology: a single institution perspective
Journal of Cancer Research and Clinical Oncology (2023)
-
Amphicrine carcinoma of the stomach and intestine: a clinicopathologic and pan-cancer transcriptome analysis of a distinct entity
Cancer Cell International (2019)