Abstract
Breast cancer is a heterogeneous disease comprising a diversity of tumor subtypes that manifest themselves in a wide variety of clinical, pathological, and molecular features. One important subset, luminal breast cancers, comprises two clinically distinct subtypes luminal A and B each of them endowed with its own genetic program of differentiation and proliferation. Luminal breast cancers were operationally defined as follows: Luminal A: ER+, PR+, HER2-, Ki-67<14% and Luminal B: ER+ and/or PR+, HER2-,Ki-67≥14% or, alternatively ER+ and/or PR+, HER2+, any Ki-67. There is currently a need for a clinically robust and validated immunohistochemical assay that can help distinguish between luminal A and B breast cancer. MCM2 is a family member of the minichromosome maintenance protein complex whose role in DNA replication and cell proliferation is firmly established. As MCM2 appears to be an attractive alternative to Ki-67, we sought to study the expression of MCM2 and Ki-67 in different histological grades and molecular subtypes of breast cancer focusing primarily on ER-positive tumors. MCM2 and Ki-67 mRNA expression were studied using in silico analysis of available DNA microarray and RNA-sequencing data of human breast cancer. We next used immunohistochemistry to evaluate protein expression of MCM2 and Ki-67 on tissue microarrays of invasive breast carcinoma. We found that MCM2 and Ki-67 are highly expressed in breast tumors of high histological grades, comprising clinically aggressive tumors such as triple-negative, HER2-positive and luminal B subtypes. MCM2 expression was detected at higher levels than that of Ki-67 in normal breast tissues and in breast cancers. The bimodal distribution of MCM2 scores in ER+/HER2- breast tumors led to the identification of two distinct subgroups with different relapse-free survival rates. In conclusion, MCM2 expression can help sorting out two clinically important subsets of luminal breast cancer whose treatment and clinical outcomes are likely to diverge.
Similar content being viewed by others
Main
According to the last annual report of the American Cancer Society, it is estimated that 246 660 new cases of invasive breast cancer will be diagnosed among women in the United States in 2016 resulting in an estimated 40 450 breast cancer deaths. Only lung cancer exacts a larger toll from women in terms of mortality.1 Breast cancer is a heterogeneous disease that manifests itself with a variety of clinical and pathological features, and is characterized by variable responses to treatment.2, 3
Immunohistochemistry analysis of tumor samples identifies three major breast tumor types based on expression of two main therapeutic targets, ER and HER2. Tumors expressing ER (ER+) are candidates for hormonal therapies, while those expressing ERBB2/HER2 (HER2+) are candidates for therapies targeting this receptor. Tumors negative for both markers are called triple-negative, a classification by default that is likely heterogeneous. Classification of breast tumors by gene expression profiling can further refine these broad categories of tumors, based on gene expression patterns, identifying subtypes of tumors with characteristic disease phenotypes and significant differences in outcomes.4, 5 Molecular subtypes include estrogen receptor positive (ER+) breast cancer (luminal A and luminal B) and estrogen receptor negative (ER-) breast cancer (HER2-positive, basal-like and a so-called normal breast-like breast cancer subtype).6 Luminal A and luminal B breast cancers are significantly distinct at the molecular level. In our study, luminal breast cancers were operationally defined as follows: Luminal A: ER+, PR+, HER2-, Ki-67 <14% Luminal B: ER+ and/or PR+, HER2-, Ki-67 ≥14% or, alternatively ER+ and/or PR+, HER2+, any Ki-67.7, 8, 9
The proliferation signature of tumor cells, comprised of cell cycle-regulated genes whose expression correlates with rapid cell division,10, 11 has been detected in different types of cancer such as breast, prostate, liver, and lung.12, 13, 14, 15 There is now growing evidence that the proliferative capacity of breast cancer impacts on the prognosis, clinical behavior and aggressiveness of the tumor suggesting that accurate measurement of cell proliferation may help to select a more appropriate treatment.16, 17, 18 Indeed, proliferation is considered having the highest weight component in the Oncotype DX recurrence score.19 While DNA microarray is an ideal way to measure multi-gene expression proliferation signature in a quantitative and automated manner, the need for a histological based proliferation test is crucial for pathologist routine clinical assessment of breast cancer.10, 20 Currently, mitotic index and immunohistochemical assay of proliferation-associated antigen such as Ki-67, cyclins, minichromosome maintenance (MCM) proteins, and topoisomerase IIα are the most common practical histological means to assess cell proliferation.17, 21
Ki-67 is the single most important cell proliferation related genes and, as such, it has been widely investigated as a proliferative marker in breast cancer with the hope of distinguishing luminal A from luminal B breast cancer.22, 23, 24 Indeed, an immunohistochemical panel of four markers: ER, PR, HER2, and Ki-67 have been proposed as a surrogate classification method for breast tumor subtypes.7, 22, 25 However, Ki-67 assessment in breast cancer has failed to impose itself as an accepted proliferative biomarker due to its lack of reproducibility and the difficulty finding an appropriate cutoff (ie, 10%, 14% and 20%).9, 22, 26, 27, 28, 29 Indeed, neither the American Society of Clinical Oncology nor the National Comprehensive Cancer Network has included Ki-67 in its recommended list of routine breast cancer biomarkers.30, 31 Furthermore, the IMPAKT 2012 working group pointed out that there was not enough evidence to support using a Ki-67 labeling index of 14% to identify clinically relevant subtypes of breast cancer.32 In this context, identification of a robust breast cancer proliferation marker would be of the utmost interest.
MCM2, whose role in initiating DNA replication is now firmly established,33 may represent an appealing alternative to Ki-67. MCM2 belongs to the MCM protein complex or pre-replicative complex, which consists of six highly conserved proteins (MCM2-MCM7) collectively interacting to bring about initiation of DNA replication.34 This complex is considered to be the converging point that connects the growth signaling pathways with the initiation of DNA replication.35 MCMs are expressed in actively proliferating cells and non-cycling cells with proliferative potential.36 Recently, it has been recognized that cancers arising in different anatomic sites such as stomach, colon, and skin are associated with MCM2 overexpression.37, 38, 39 Furthermore, recent reports support the notion that MCM2 is a promising proliferative marker in many other types of cancer including thyroid, rectal and breast cancers.40, 41, 42
Our goal was to a compare the level of expression of MCM2 and Ki-67 as they relate to breast cancers of different histological grades and molecular subtypes, focusing primarily on ER-positive tumors. More specifically, we sought to assess the potential clinical usefulness of these markers to assess cellular proliferation in breast cancer. We also wanted to compare the prognostic value of MCM2 and Ki-67 in predicting the overall survival, incidence of relapse and survival after relapse. To achieve these aims, we performed both an in silico analysis on available human breast cancer expression data sets (expression microarray and RNA-sequencing), and immunohistochemical staining of tissue microarrays constructed from two different cohorts of breast cancer patients, a discovery cohort (n=554) and a validation cohort (n=266). In brief, our data strongly support the both MCM2 and Ki-67 are highly expressed in breast cancer of high histological grades that comprise clinically aggressive tumors such as triple-negative, HER2-positive and luminal B subtypes of breast cancer.
Material and methods
In silico Analysis of MCM2 and MKI67 mRNA Expression Levels According to PAM50 Subtypes
The web application bc-GenExMiner43 was used for correlation analysis of MCM2 and MKI67 gene expression on a microarray data set collection comprising over 3063 samples, out of which 1260 could be positively ascribed to one out of the five different molecular subtypes of breast cancer. Gene expression values were used to define three groups of equal size (low, intermediate, and high expression) in order to study their distribution in the different molecular subtypes.
To validate results obtained from bc-GenExMiner application, cohorts from the TCGA (RNAseq data of 756 breast cancer patients) and The University of California Santa Cruz Cancer Genomics Browser (Gene expression array of 597 patients (AgilentG4502A_07_3 array)) were also used.44 The heat map (Supplementary Figure 1) produced from the University of California Santa Cruz cancer genomic browser displays the expression of different genes using distinct sets of colors, red representing data values>0, green values<0 and black value=0. BreastMark, a web-based application, was used to evaluate MCM2 and MKI67 genes association along with survival in breast cancer and its molecular subtypes. This algorithm uses gene expression data from 2652 samples in 26 different data sets, and detailed clinical data to correlate the outcome with gene expression levels. Cox regression analysis was used to calculate hazard ratios. All calculations were carried out in the R statistical environment.45
Patients and Tissue Samples
Discovery cohort
All specimens in this cohort were from Saint-Sacrement Hospital in Québec city. Specimens were obtained from mastectomies and segmental resections that were performed at the Centre des Maladies du Sein Deschênes-Fabia at the Saint-Sacrement Hospital in Québec, Québec, Canada, between February 2011 and April 2012 (Table 1). Specimens were fixed with 10% neutral buffered formalin, embedded in paraffin, cut in to 4 μm tissue sections, stained with hematoxylin and eosin, and used for routine pathological evaluation. The study population consisted of 554 consecutive cases of invasive breast carcinoma with a tumor size on histological slides of at least ~1 cm none of which received chemotherapy before surgery. All eligible women provided written informed consent. Ethical approval of the study was obtained from the Research ethics committee of the Quebec Research Center (No. 2016-2688).
Validation cohort
This cohort consisted of 266 female breast cancer patients comprising tumors of different histological grades. None of those patients received neo-adjuvant chemotherapy. Representative tumor blocks of formalin-fixed, paraffin-embedded were collected after surgery (lumpectomy or mastectomy; Table 1). We also added 21 normal breast tissues from healthy women undergoing plastic surgery to serve as internal controls. Tumor grades were confirmed using the Modified Scarff-Bloom-Richardson-Elston-Ellis grading system.46 Normal human brain was included as negative control for MCM2.47 In addition, a number of extraneous tissues such as colon, thyroid, and placenta were included. A complete set of follow-up data from this cohort of patients, including the onset of relapse and death was available for review. All samples were obtained from Centre Hospitalier de l’Université de Montréal after granting the approval of the research ethical committee (SL 05.019). Since all donor blocks remained anonymous no individual patient consent was required.
Tissue Microarray
Sections (4 μm) from each paraffin donor block were stained with hematoxylin and eosin and examined by two independent pathologists. Core punches, 0.6 mm (for the discovery cohort), 1mm (for the validation cohort) in diameter, were plucked from representative areas contained within each formalin-fixed, paraffin-embedded tumor blocks. Each core was realigned in duplicate or triplicate into recipient blocks according to the intended design of the map using a Manual Tissue Arrayer I (Beecher Instruments). Blocks were next inverted and incubated overnight in the oven over a glass slide. Tissue microarray blocks were allowed to cool until they could easily detach from the glass slide. Tissue sections from each tissue microarray were prepared and one slide from each block was stained with hematoxylin and eosin to review the diagnoses and histological grades on all tissue samples. Additional representative sections from each block were submitted to immunohistochemical staining.
Immunohistochemistry
Immunohistochemical assays were performed on formalin-fixed, paraffin-embedded tissues obtained from each tissue microarray. These assays were carried out according to manufacturer recommendations on an automated immunostainer (Discovery XT system, Ventana Medical Systems, Tucson, AZ, USA). Immunohistochemical analysis of MCM2 (monoclonal; #12079, dilution 1/200, citrate buffer, Cell Signaling) was carried out. Specificity of the anti-MCM2 monoclonal antibody was confirmed using normal colonic mucosa48 and normal cerebral cortex as positive and negative controls, respectively (Supplementary Figure 2), based on the expression data in the Human Protein Atlas database.49 In addition, immunohistochemical analysis of estrogen receptor (ER; monoclonal, clone SP1, Ready to use, citrate buffer, Ventana Medical Systems), progesterone receptor (PR; monoclonal, clone 1E2, Ready to use, citrate buffer, Ventana Medical Systems), HER2 (monoclonal, clone 4B5, Ready to use, citrate buffer, Ventana Medical Systems), and Ki-67 (monoclonal, clone SP6, dilution 1/100, pretreated citrate buffer, BioCare medical) was carried out as previously described.50
Scoring of Stained Slides
The scoring systems used for each antibody are listed in Supplementary Table 1. The expression of MCM2 and Ki-67 in breast epithelium was studied by calculating the percentage of positively stained nuclei.22, 51 Two different approaches of scoring were used, a visual method using conventional light microscopy and computer-assisted automated scoring method using Visiomorph and Tissuemorph Digital Pathology software. Visual scoring of MCM2 and Ki-67 staining reactions is labor-intensive, time consuming and is subjected to intra-observer and inter-observer variability. To increase reproducibility in the visual scoring system, evaluation was performed by two independent observers. The visual scoring was used for assessment of both cohorts of patients. Automated MCM2 and Ki-67 scorings were also carried out resulting in a much more rapid and accurate readout. Visiomorph Digital Pathology has the distinct advantage of leaving out stromal cells from the analysis retaining only cancer cells in the Region Of Interest. As for Tissuemorph Digital Pathology, it allows accurate counting of the positive and negative nuclei in the region of interest. This automated scoring system was used only with the validation patient cohort. ER, PR, HER2, and Ki-67 were used as surrogate markers to sub-classify breast cancers into different molecular subtypes as listed in Supplementary Table 2.3, 32
Statistical Analyses
All statistical analyses were carried out using XLSTAT (http://www.xlstat.com/en/) and different packages of the R language (http://www.R-project.org/). The allocation of MCM2- and Ki-67-positive cases among different histological grades and molecular subtypes is depicted using boxplots and scatter plots for both visual and automated cell counts. The overall relationship between MCM2 and Ki-67 scores and both histological grades and molecular subtypes was evaluated using the χ2-test and Mann–Whitney test /two-tailed test. Correlation analysis for immunohistochemical expression levels was carried out using Spearman’s ρ correlation coefficient for visual scores and Pearson correlation coefficient for automated score. Concordance of automated and visual scoring was also achieved using Intra-class Correlation Coefficient.52
The receiver-operating characteristic (ROC) curve was used to detect the optimal cutoff point, which simultaneously reached maximum sensitivity and specificity values for MCM2 score. Using this cutoff point, continuous variables could then be treated as dichotomous variables (low and high MCM2 expression).53 Distribution of MCM2 and Ki-67 in luminal breast cancer was displayed using histograms. Kaplan–Meier plots were drawn for the overall survival and survival after relapse of patient groups with low or high levels of MCM2 expression. Cox regression was used for multivariate survival analysis. Statistical significance was considered for P-values lower than 0.05.
Results
MCM2 is Highly Expressed at the MRNA Level in Basal-Like, Luminal B and HER2-Positive but not Luminal a Breast Tumors
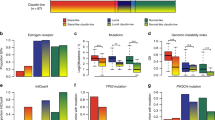
The web application bc-GenExMiner was used to compare the mRNA levels of MCM2 and MKI67 within each breast cancer molecular subtype on an aggregated data set comprising 1260 tumor RNA expression profiles.43 As shown in Figures 1a and b and Supplementary Figure 1, basal-like, luminal B and HER2-positive breast cancers show higher expression of MCM2 and MKI67 mRNA levels in comparison with both luminal A and normal-like breast cancers (P<0.0001). Notably, there is no overlap between individual boxes in the boxplot when luminal A were compared with luminal B breast cancers (Figures 1a and b). The percentage of patients with low, intermediate and high levels of MCM2 and MKI67 expression in each molecular subtype of breast cancer is depicted in Figure 1c. In this microarray data set, 74% of basal-like (total no.=388), 62% of luminal B (total no.=116) and 49% of HER2-positive (total no.=103) breast cancer patients show high level of MCM2 mRNA expression. In contrast, only 7% of luminal A (total no.=443) and 5% of normal-like (total no.=210) breast cancer show high level of MCM2 mRNA expression. Similar relative levels of MKI67 mRNA expression were observed in the different breast cancer molecular subtypes (Figure 1c).
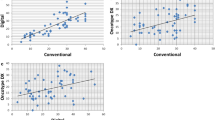
In silico analysis of MCM2 and MKI67 mRNA expression in PAM50 molecular subtypes of breast cancer. (a and b) Boxplots depicting higher levels of MCM2 and MKI67 mRNA levels in basal-like, luminal B and HER2-positive breast cancers in comparison to luminal A and normal-like breast cancers (P<0.0001). (c) Quantification of the number of patients with low, intermediate and high MCM2 and MKI67 expression in each molecular subtype of breast cancer in the bc-GenExMiner database. Elevated expression of MCM2 and MKI67 is observed in basal-like, luminal B and HER2-positive breast cancer. (d) Two-way scatter plots illustrating the correlation between mRNA expression levels ofMCM2,MKI67,andESR1in breast tumors from The Cancer Genome Atlas (TCGA). Correlation between MCM2 and MKI67 is positive (r=0.73), whereas those between ESR1 and both MCM2 and MKI67 are negative (r=−0.33, r=−0.38, respectively).
MCM2 expression levels were next correlated with those of MKI67 and ESR1 in breast tumors based on RNA-sequencing data derived from The Cancer Genome Atlas (TCGA). As predicted, a strong positive correlation could be detected between MCM2 and MKI67 (r=0.73), whereas a negative correlation was observed between ESR1 and both MCM2 and MKI67 (r =−0.33, r=−0.38, respectively) (Figure 1d). This close similarity between the in silico profiles of MCM2 and MKI67 prompted us to pursue assessment of MCM2 protein expression in breast carcinoma tissue microarrays along with cognate normal breast tissue.
A 40% Index Cutoff Point Can Distinguish Between Two Distinct Subgroups of Low and High Expression of MCM2 in Breast Cancer
After confirming the sensitivity and specificity of the MCM2 immunohistochemical assay (Ab #12079, Cell Signaling), we carried out high-throughput immunohistochemical reactions on the discovery cohort tissue microarrays. A visual scoring system was used to calculate the percentage of positive nuclei of MCM2 and Ki-67 in the stained sections. A the ROC curve was used to determine the optimal cutoff point based on the MCM2 data in the discovery cohort. When the accuracy and the sum of sensitivity and specificity were taken into account, the optimal cutoff point corresponded to a value of 40%. Using this cutoff value, the sensitivity of MCM2 index was found to be 87% and specificity 96%. We next evaluated the accuracy of the test by measuring the area under the curve as it provides clear indication on how well the ROC curve separates out the cohort into two distinct subgroups. When our scores were considered, the Area Under the Curve was found to be 0.95 (P<0.0001). This implies that a 40% cutoff point can be considered an 'excellent' discriminating value in separating low and high MCM2 expression levels.54 In other words, tumors with scores ranging from 0 to 40% could be considered to have low MCM2 expression while those with scores exceeding 40% were considered to have high MCM2 expression.
These results were further expanded using a second independent cohort of patients (Validation cohort). In this cohort, two different approaches were used to calculate the percentage of positive nuclei of MCM2 in the stained sections, a visual scoring and an automated quantifying method. It was confirmed that 40% is the optimal cutoff point that can discriminate between low and high MCM2 expression. The sensitivity of the MCM2 index was found to be 69%, the specificity 91% and the approximate area under the curve was found to be 0.81 (P<0.0001) based on the automated scores.
High Concordance Between Automated and Visual Scores
We also used both a visual scoring and an automated quantifying method to calculate the percentage of nuclei positive for Ki-67 in the stained sections of the validation cohort of patients. Intra-class correlation coefficient was calculated to compare visual and automated scores for both MCM2 and Ki-67 and to evaluate their relative performance.52 Intra-class correlation coefficient was obtained for the 365 pairs of automated and visual scores of MCM2 and Ki-67. Our results demonstrated that Intra-class Correlation Coefficients for MCM2 automated versus visual scoring were 0.94 (95%CI=0.93–0.95, P<0.0001) for single measures and 0.97 (95% CI= 0.96–0.97, P<0.0001) for average measures. Intra-class Correlation Coefficients for Ki-67 automated versus visual scoring were 0.90 (95%CI=0.87–0.91) for single measures and 0.94 (95%CI=0.93–0.95) for average measures. Therefore, there is an excellent agreement (intra-class correlation coefficient >0.90) between automated and visual scorings for both MCM2 and Ki-67.52 Taken together, our results suggest that both methods can be used to reliably assess MCM2 and Ki-67 expressions in breast cancer. Interestingly, automated scoring of immunohistochemistry stained tissue microarray can be considered as an effective high-throughput method to evaluate protein expression.
Two Distinct Patterns of MCM2 Expression Were Detected in Normal Breast Tissues
Normal breast tissues were stained with either MCM2 or Ki-67 antibodies (Figure 2). Of note, there was no MCM2 or Ki-67 labeling either in the myoepithelial cell layer or in the surrounding stromal cells. Also, we never observed cytoplasmic or membrane staining in the labeled cells. Ki-67 staining was localized to only a few scattered epithelial cells lining the terminal duct lobular units (Figure 2a). The majority (62%) of normal breast tissue samples exhibit a similar pattern upon MCM2 staining (Figure 2b). However, as many as 36% of normal breast tissues demonstrated a strong MCM2 nuclear labeling in most if not all of the luminal cells lining the normal breast ducts (Figure 2c). In these tissues Ki-67 staining was similar to that of the remaining samples in the cohort, pointing to a difference in the detection patterns of these markers in breast epithelial cells. Although a positive correlation could be established between MCM2 and Ki-67 scores in normal breast tissue (r=0.45, P=0.04), MCM2 scores were higher in normal breast tissue compared with that of Ki-67 (mean difference between two scores=40%, 95%CI= 22.21–56.94, P<0.0001).
Patterns of MCM2 and Ki-67 expression in normal breast tissue. (a) Staining of normal breast tissues with Ki-67 demonstrating nuclear labeling in only a few scattered luminal cells lining the terminal duct lobular units in 100% of normal breast tissue. (b) Staining of normal breast tissues with MCM2 displaying nuclear labeling in only a few scattered luminal cells in the terminal duct lobular units; this labeling pattern is representative of the majority (62%) of normal breast tissue samples (c) Staining of normal breast tissues with MCM2 displaying strong nuclear labeling in most luminal cells lining the normal breast ducts; this pattern is representative of ~36% of normal breast tissues. In all cases (a–c), no labeling was detected in the myoepithelial cell layer or in the surrounding stromal cells. Magnification × 20.
Higher Levels of Expression of MCM2 are Associated With Breast Cancers of High Histological Grades
The expression of MCM2 and Ki-67 was then examined in breast cancers of various histological grades. MCM2 and Ki-67 reactivity was restricted to the nuclei of cancer cells, neither of the two markers being detected in the adjacent stromal cells. On the basis of the previously defined 40% cutoff, high expression of MCM2 was closely associated with high-grade breast tumors (P<0.0001) in the validation cohort (Figure 3a). Similarly, high expression of Ki-67 was significantly associated with high-grade breast cancer (P<0.0001) in the same group of patients (Figure 3b). To further investigate this finding, we next assessed the distribution of low- versus high-expression groups in different grades of breast tumors in the validation cohort for both MCM2 and Ki-67 using the pre-determined cutoff values. Our results demonstrated that 97% of grade I breast cancer expressed low levels of MCM2. On the other hand, high levels of MCM2 expression were detected in 29% of grade II (P<0.0001) and 74% of grade III breast cancer (P<0.0001), which is statistically significant when compared with grade I breast cancer (Figure 3c). We also found significantly higher levels of expression of Ki-67 in grade II (P<0.014) and grade III breast cancers (P<0.0001) in comparison with grade I tumors (Figure 3d). As expected, we found a positive correlation between MCM2 and Ki-67 in breast cancer of different histological grades (Grade I r=0.69, Grade II r=0.73, Grade III r=0.91, P<0.0001). However, MCM2 scores in breast cancers exceeded those of Ki-67 (mean difference between two scores=25%, 95%CI=20.12-29.54, P<0.0001) (Figures 3e and g).
Overexpression of MCM2 is associated with high histological grade of breast cancer. (a) Diagram showing that high expression of MCM2 is closely associated with high grades in breast tumors (P<0.0001) in our validation cohort. (b) Diagram showing that high expression of Ki-67 is significantly associated with high grades in breast tumors (P<0.0001) in the same cohort. (c) Histograms displaying the percentages and numbers of patients with low and high MCM2 expression in breast tumors of different histological grades. High levels of MCM2 expression were detected in 29% of grade II and 74% of grade III breast cancer, which is statistically significant (P<0.0001) when compared with grade I breast cancer. (d) Histograms displaying the percentages and numbers of patients with low and high Ki-67 expression in breast tumors of different histological grades. Significant higher levels of Ki-67 expression were detected in grade II (P<0.014) and grade III breast cancers (P<0.0001) in comparison to grade I tumors. (e) Staining of grade I, (f) grade II, (g) grade III breast tumors with MCM2 (upper panels) or Ki-67 antibodies (lower panels). Note the overexpression of MCM2 and Ki-67 in breast tumors of higher histological grades. Levels of MCM2 staining (% of cells) characteristically exceeded those with Ki-67 in different grades of breast cancers (pictures were taken from the same area of adjacent slides).
Increased Expression of MCM2 is Associated With Luminal B, Triple-Negative, and HER2-Positive Breast Cancer
We next examined whether the association between luminal B, triple-negative and HER2-positive status and high expression levels of MCM2 observed at the RNA level in the in silico analysis is reproduced at the protein level using the above-described immunohistochemical assays for MCM2 and Ki-67 in the discovery and validation cohorts. Molecular subtypes were operationally defined using the following the surrogate immunohistochemical signature consisting of ER, PR, HER2 and Ki-67 as documented in Supplementary Table 2.32 As shown in Table 2, high levels of MCM2 expression were detected in luminal B (P<0.0001), HER2-positive (P<0.0001) and triple-negative (P<0.0001) in both cohorts of patients. In contrast, most luminal A breast tumors expressed low levels of MCM2, and luminal A tumors displayed significant differences when compared all other subtypes of breast cancer (Figure 4a). Likewise, significant differences could be detected between Ki-67 expression in luminal B, HER2-positive and triple-negative breast cancer (P<0.0001) when compared with luminal A subtype (Figure 4b).
Overexpression of MCM2 is associated with triple-negative, luminal B and HER2-positive breast cancer. (a) Levels of MCM2 expression in luminal B, HER2-positive and triple-negative breast cancer in validation cohort. Significant differences (P<0.0001) were detected between these three subtypes and luminal A breast cancer. (b) Significant differences (P<0.0001) were also detected between Ki-67 expression in luminal B, HER2-positive and triple-negative breast cancer when compared with luminal A tumors in the same cohort. (c) Luminal A, (d) Luminal B, (e) HER2-positive, (f) Staining of triple-negative breast tumors with MCM2 (upper panel), or Ki-67 (lower panel). Overexpression of MCM2 and Ki-67 is associated with luminal B, HER2-positive and triple-negative breast cancer. MCM2 staining is consistently higher than that of Ki-67 in different molecular subtypes of breast cancer. (Pictures were taken from adjacent sections of the same punches). Magnification × 20 (c-f).
In further agreement with the in silico correlation analyses, we found a negative correlation between MCM2 and ER (r=−0.54, P<0.0001) and MCM2 and PR (r=−0.49, P<0.0001) in breast cancer at the level of protein expression. In addition, there was a positive correlation between MCM2 and Ki-67 in different molecular subtypes of breast cancer (luminal A r=0.55 (P<0.0001), luminal B r=0.46 (P=0.001), HER2-positive r=0.65 (P<0.0001), triple-negative r=0.62 (P<0.0001)). However, as mentioned earlier, MCM2 scores were significantly and consistently higher than those of Ki-67 for each molecular subtype. The mean difference between the two scores was 13.10% (95%CI= 0.01–16.02, P<0.0001) in the luminal A subtype, 18.98% (95%CI=0.008–30.11, P=0.001) in the luminal B subtype, 24.99% (95%CI=0.015–34.67, P<0.0001) in HER2-positive breast tumors and 18.67% (95%CI=0.01–25.65, P<0.0001) in triple-negative breast tumors (Figures 4c and f).
MCM2 Scores in Luminal Breast Cancer Follow a Bimodal Frequency Distribution
The frequency distribution of MCM2 scores for all luminal breast tumors (defined by positive expression of ER and/or PR, lack of HER2 expression and 14% labeling index of Ki-67) was further assessed. In the discovery cohort, we observed a bimodal distribution of MCM2 scores in the luminal subtype (data not shown), suggesting that MCM2 can separate out two distinct subgroups among hormonally responsive luminal breast cancers. Among 753 samples of luminal breast cancer, 65% showed scores ranging from 0 to 40%, representing luminal breast cancer with low proliferation. On the other hand, approximately 35% of tumors showed MCM2 scores higher than 40%, defining a class of luminal breast cancer endowed with high proliferation.
The same bimodal distribution pattern has been confirmed in the validation cohort (Figure 5a). Among the 156 samples of luminal breast cancer, 70% showed scores ranging from 0 to 38.4%, representing luminal breast cancer with low proliferation. On the other hand, ~30% of patients had MCM2 scores ranging from 38.5–100%, defining a subgroup of luminal breast tumors endowed with high proliferation. This bimodal distribution pattern, detected in both cohorts, readily justifies the usefulness of a 40% threshold as a cutoff point to distinguish between MCM2 high and low expression. In contrast, when the same type of analysis was applied to Ki-67 scores using the same set of 156 luminal breast cancer tissue cores, we failed to observe a similar bimodal distribution in luminal breast cancers (Figure 5b).
A bimodal frequency distribution of MCM2 scores is detected in luminal breast cancer. (a) Frequency distribution analysis illustrating the bimodal distribution of MCM2 scores among luminal breast tumors. In validation cohort, MCM2 labeling helps to distinguish two distinct subgroups in luminal breast tumors. Tumors with scores lower than 38.4% are considered to represent tumors with low proliferative potential, while tumors with higher scores are considered have a high proliferative potential. (b) Distribution of Ki-67 scores in the same cohort. Note the absence of a clear bimodal distribution pattern.
As shown in Figure 5a, one can notice that some of the breast tumors defined by low Ki-67 (<14%, putative luminal A subgroup), express high levels of MCM2, while some of the high Ki-67 expressers (putative luminal B subgroup), have low MCM2 expression. In the discovery cohort of patients, if applying a 40% index of MCM2 instead of 14% index of Ki-67 would result in roughly 6.6% of tumors (50 out of 753) being reclassified. In the validation cohort, 10% of the tumors (15 out of 156) would have been reassigned to the other subtype of breast cancer based on MCM2 as a proliferation marker instead of Ki-67.
High Levels of MCM2 are associated with a Shorter Latency to Relapse
The association between MCM2 and MKI67 gene expression and survival in breast cancer was investigated in silico with BreastMark using a data set comprising gene expression profiles from 2652 tumors. High expression of MCM2 and MKI67 is significantly associated with shorter survival rate in breast cancer (For MCM2: Hazard ratio=1.467, P=1.504e–10) (For MKI67: hazard ratio=1.432, P=1.948e–09) (Figures 6a and b).
High levels of MCM2 are associated with a shorter latency to relapse. (a and b) Kaplan–Meier curves for the overall survival of groups with high or low expression of MCM2 or Ki-67 in breast cancer using the BreastMark web application. High expression of MCM2 and MKI67 is significantly associated with shorter survival rates in breast cancer (For MCM2: hazard ratio=1.467, P=1.504e–10) (For MKI67: hazard ratio=1.432, P=1.948e–09). (c and d) Elevated levels of MCM2 and Ki-67 expression are associated with a shorter latency to clinical relapse in our validation cohort. The association between MCM2 and relapse (P=0.0003) is tighter than that of Ki-67 (P=0.0458). (e and f) No significant association between either MCM2 (P=0.0631) or Ki-67 (P=0.453) levels of expression and overall survival could be identified.
We next reviewed the clinical charts of 200 patients from the validation cohort for the period extending from 2000 to 2013 to assess the association between MCM2 and Ki-67 expression with clinical relapse. Our results demonstrate that enhanced expression of both of MCM2 and Ki-67 is associated with a shorter latency to clinical relapse in a statistically significant manner. However, the association between MCM2 expression and relapse (P=0.0003) is tighter than that of Ki-67 (P=0.0458) (Figures 6c and d). We next conducted univariate Cox regression analysis on our data to determine the role of a number of factors including MCM2 expression levels, histological grade, histological subtype, and molecular subtype on the incidence of relapse for patients. Results confirmed the association of MCM2 overexpression with a higher risk of relapse (HR=6.98, 95%CI=2.00–24.40, P=0.002). Moreover, as expected the luminal A subtype, defined by the surrogate immunohistochemical signature, had a significantly lower risk to develop relapse than HER2-positive breast cancer (HR=0.11, 95%CI=0.02–0.56, P=0.01) (Table 3). However, no other factor was found to be significant in multivariate analysis.
On the other hand, the Kaplan–Meier overall survival curve obtained from the same cohort of patients showed no significant association between both MCM2 and Ki-67 levels of expression and overall survival (Figures 6e and f). The log-rank test demonstrated that the survival intervals are not significantly different in both groups at 5% level of significance, for MCM2 (P=0.0631) and for Ki-67 (P=0.453). It is interesting however to note that the difference in survival after 5.76 years is almost 10% lower in patients with high MCM2 expression. The difference remains the same even after 10 years, indicating that patients who survive after 5.76 years maintain a constant risk of dying. Finally, no significant difference could be identified between survival after relapse for patients having high or low levels of MCM2 (P=0.0817) or of Ki-67 expression (P=0.355). Univariate and multivariate analyses confirmed the association between triple-negative breast cancer and risk of death (HR=0.23, 95%CI=0.07–0.78, P=0.0185) and survival after relapse (HR=0.25, 95%CI=0.07–0.85, P=0.03) in patients with high MCM2 expression. No other factors such as histological grades, histological subtypes, and other molecular subtypes appeared to be significant in case of overall survival and survival after relapse (Tables 3 and 4).
Discussion
In the present study, we have compared the levels of expression of both MCM2 and Ki-67 as a means to assess cellular proliferation in a large cohort of patients with breast tumors of different grades and molecular subtypes. We also compared the prognostic value of these two markers in predicting the overall survival, incidence of relapse and survival after relapse. Our findings confirmed that both MCM2 and Ki-67 are highly expressed in higher histological grade tumors especially in clinically aggressive breast cancers such as triple-negative, HER2-positive and luminal B subtypes tumors. Importantly, in contrast to Ki-67, MCM2 had a bimodal distribution in luminal breast tumors, leading to the identification of two distinct subgroups among luminal breast cancer (ER+/HER2−) using a MCM2 labeling index of 40%.
Using tissue microarrays comprising normal breast tissue and human breast cancers of different grades and subtypes, we found that MCM2 is more readily detected than Ki-67 both in normal breast tissue and in different types of breast cancer. This finding has also been obtained using whole sections of breast cancer. This is consistent with previous observations by others reporting higher expression of MCM2 when compared with Ki-67 in normal breast tissues and breast carcinoma at the protein level using immunohistochemistry.51, 55 The fact that MCM2 labeling was observed at high levels in tissues that are labeled at low levels by Ki-67 suggests that MCM2 expression can detect subsets of proliferating mammary epithelial cells that are not detected by Ki-67 alone. This is in line with the findings of Lopez et al.56 who confirmed that Ki-67 is completely absent in the initial G1 phase of the cell cycle. Alternatively, one might surmise that Ki-67 protein is present inside the cells but cannot be detected due to pre-analytical condition such as fixation or because of altered biological properties of Ki-67 such as conformational changes or stable interactions with other proteins57 or because of low sensitivity of Ki-67 antibody used in immunohistochemistry.
The dual pattern of MCM2 expression in normal breast tissue is intriguing. Although 62% of normal breast tissues (n=21) showed MCM2 nuclear labeling in only a few scattered cells lining the terminal duct units, a significant proportion (36%) of normal tissues (n=21) displayed MCM2 expression in the vast majority of normal breast cells. The biological significance of these distinct patterns of MCM2 expression is presently unknown and needs further clarification. The highly proliferative group may reflect the state of hormonal stimulation in a given patient at the time of surgery. Whether or not this higher MCM2 staining in normal tissues results in a higher susceptibility to neoplastic transformation is currently an unresolved question.58 Clearly, more investigations are needed with larger cohorts patients without breast cancer with long-term follow-up to clarify this issue.
Our results also indicate that both MCM2 and Ki-67 are highly expressed in high histological grade breast cancer. This is consistent with two different studies that reported a significant correlation between proliferative markers including both MCM2 and Ki-67 and breast cancer grades.59, 60 Our findings coming from both in silico analyses and immunohistochemistry also support the notion that, similar to Ki-67,61, 62, 63, 64 MCM2 is more highly expressed both at the mRNA and protein levels in triple-negative, HER2-positive as well as luminal B subtypes than in luminal A breast tumors. Although to our knowledge, no previous report in the literature specifically correlated MCM2 expression with individual breast cancer molecular subtypes, our findings are in agreement with Kowk et al. and Joshi et al., who reported an association between MCM2 overexpression and a negative ER status of breast tumors.60, 65
On the basis of Ki-67 labeling distribution, it has been previously proposed that ER-positive breast tumors form a continuum rather than segregating into distinct subtypes.25 The data presented herein clearly challenge this interpretation by showing that two distinct subgroups of hormonally responsive breast cancers (ER+/HER2-) can be identified based on the estimated percentage of MCM2-positive tumor cells using a threshold of 40% set in accordance with the ROC curve determined in our discovery cohort. Hence, ER+/HER2- tumors with MCM2 expression ranging from 0 to 40% are considered to be luminal breast cancer with low proliferative potential, while ER+/HER2- tumors with overexpression of MCM2 (>40%) are considered to represent luminal breast cancer with high proliferative potential. This approach is similar to the one used to determine the cutoff of Ki-67 to distinguish luminal A from luminal B breast cancer.22 The fact that we observed a bimodal distribution of MCM2 scores in luminal breast cancer suggests that differences in the patterns of gene expression between the two markers. This conclusion is supported by a recent gene expression study reporting that MCM2 but not Ki-67 belongs to a class of breast cancer genes with bimodal distribution.66
Two different approaches of immunohistochemical scoring were used, a visual method using conventional light microscopy and computer-assisted automated scoring method using Visiomorph, Tissuemorph Digital Pathology software. Our results demonstrate that there was an excellent concordance between the two scoring system, which suggest that both scoring methods can be used reliably to assess MCM2 and Ki-67 expression in breast cancer. This is consistent with Joshi et al. who demonstrate that automated scoring can be used effectively as a high-throughput method to evaluate immunohistochemical expression.60
Lastly, we sought to correlate the expression profiles of MCM2 and Ki-67 with clinical outcome characteristics such as tumor relapse and survival rates. Our results confirmed that overexpression of both markers is associated with shorter latency to clinical relapse. However, the association between MCM2 and relapse (P=0.0003) is more tightly correlated than that of Ki-67 (P=0.0458). Our data strongly support the role of Ki-67 and MCM2 as prognostic markers in breast cancer,22, 51, 65 compatible with an altered function of the DNA replication licensing system, MCM2-MCM7 leading to uncontrolled cell proliferation.67 In future studies, it will be of interest to evaluate whether other components of this system provide additional information or represent redundant markers.
There are several reasons to believe that MCM2 would outperform Ki-67 as a tool to assess cell proliferation in breast cancer. Firstly, in contrast to Ki-67, MCM2 can label all proliferative cells during the active phases of the cell cycle, disappearing only when cells are quiescent.68 Indeed, Ki-67 labeling occurs only in a fraction of proliferating cells entering the G1 phase while sustained Ki-67 expression takes place only in late G1.56, 69 One plausible explanation to the variability of Ki-67 staining may be related to the normal variation of Ki-67 concentration during different phases of the cell cycle, with accumulation during G2 and S phases and a nadir during anaphase and telophase of the mitotic cycle.70, 71, 72 Accordingly, Ki-67 has a short half-life of ~90 min.73 On the other hand, MCM2 exerts its function through reversible binding to the chromatin instead of undergoing alternating cycles of synthesis and degradation like Ki-67.34 However, quiescent cells have low levels of MCM proteins compared with proliferating cells (PMID: 11493639; PMID: 20440261). Importantly, eukaryotic cells cannot proliferate in the absence of MCM2.34 Although Ki-67 plays an important role in cell division its mechanism of action inside the cell is poorly understood.30 Notably, cells that are depleted of Ki-67, maintain a normal cell cycle profile albeit their nuclei remain smaller.74
An intrinsic limitation to our study is that our definition of luminal A and luminal B tumors in our discovery and validation cohorts was based on a surrogate immunohistochemical marker set, ie, ER, PR, HER2 and 14% index of Ki-67. Assessment of MCM2 expression in these two groups defined by mRNA expression profiles should clearly be performed in follow-up studies, which will also include additional sets of patients to independently confirm our findings. Finally, scoring of whole tissue sections rather than tissue microarray cores will better enable to evaluate the robustness of MCM2, especially in regions found to be heterogeneous using Ki-67 staining.
In conclusion, our results indicate that MCM2 outperforms Ki-67 as a proliferative and prognostic marker of breast cancer. Although Ki-67 does represent an independent prognostic marker, many practical issues limit its clinical use: tumor heterogeneity often observed in cancers, the unwieldy 14% threshold, and the fact that some proliferating cells escape Ki-67 labeling. In contrast, MCM2 detection may overcome many of these shortcomings as it labels proliferating cells that are not detected by Ki-67. It also provides related prognostic information more likely to be clinically applicable because of a better-defined cutoff point (40%) to classify ER+/HER2- breast cancer into two subsets of hormone sensitive tumors with diverging proliferative status, which should respond differently to hormonal and chemotherapeutic treatments.
References
American Cancer Society Cancer Facts and Figures. American Cancer Society: Atlanta, GA, USA, 2016.
Viale G . The current state of breast cancer classification. Ann Oncol 2012; 23: 207–210.
Caldarella A, Puliti D, Crocetti E et al, Biological characteristics of interval cancers: a role for biomarkers in the breast cancer screening. J Cancer Res Clin Oncol 2013; 139: 181–185.
Perou CM, Sorlie T, Eisen MB et al, Molecular portraits of human breast tumours. Nature 2000; 406: 747–752.
Sihto H, Lundin J, Lehtimaki T et al, Molecular subtypes of breast cancers detected in mammography screening and outside of screening. Clin Cancer Res 2008; 14: 4103–4110.
Weigelt B, Baehner FL, Reis-Filho JS . The contribution of gene expression profiling to breast cancer classification, prognostication and prediction: a retrospective of the last decade. J Pathol 2010; 220: 263–280.
Prat A, Cheang MC, Martin M et al, Prognostic significance of progesterone receptor-positive tumor cells within immunohistochemically defined luminal A breast cancer. J Clin Oncol 2013; 31: 203–209.
Tran B, Bedard PL . Luminal-B breast cancer and novel therapeutic targets. Breast Cancer Res 2011; 13: 221.
Goldhirsch A, Winer EP, Coates AS et al, Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol 2013 24: 2206–2223.
Whitfield ML, George LK, Grant GD et al, Common markers of proliferation. Nat Rev Cancer 2006; 6: 99–106.
Ganguly A, Shields CL . Differential gene expression profile of retinoblastoma compared to normal retina. Mol Vis 2010; 16: 1292–1303.
Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature 2012; 490: 61–70.
Cuzick J, Swanson GP, Fisher G et al, Prognostic value of an RNA expression signature derived from cell cycle proliferation genes in patients with prostate cancer: a retrospective study. Lancet Oncol 2011; 12: 245–255.
Andrisani OM, Studach L, Merle P . Gene signatures in hepatocellular carcinoma (HCC). Semin Cancer Biol 2011; 21: 4–9.
Wistuba II, Behrens C, Lombardi F et al, Validation of a proliferation-based expression signature as prognostic marker in early stage lung adenocarcinoma. Clin Cancer Res 2013; 19: 6261–6271.
Aleskandarany MA, Green AR, Benhasouna AA et al, Prognostic value of proliferation assay in the luminal, HER2-positive, and triple-negative biologic classes of breast cancer. Breast Cancer Res 2012; 14: R3.
van Diest PJ, van der Wall E, Baak JP . Prognostic value of proliferation in invasive breast cancer: a review. J Clin Pathol 2004; 57: 675–681.
Dai H, van't Veer L, Lamb J et al, A cell proliferation signature is a marker of extremely poor outcome in a subpopulation of breast cancer patients. Cancer Res 2005; 65: 4059–4066.
Klein ME, Dabbs DJ, Shuai Y et al, Prediction of the Oncotype DX recurrence score: use of pathology-generated equations derived by linear regression analysis. Mod Pathol 2013; 26: 658–664.
Chibon F . Cancer gene expression signatures - the rise and fall? Eur J Cancer 2013; 49: 2000–2009.
Beresford MJ, Wilson GD, Makris A . Measuring proliferation in breast cancer: practicalities and applications. Breast Cancer Res 2006; 8: 216.
Cheang MC, Chia SK, Voduc D et al, Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst 2009; 101: 736–750.
Colozza M, Azambuja E, Cardoso F et al, Proliferative markers as prognostic and predictive tools in early breast cancer: where are we now? Ann Oncol 2005; 16: 1723–1739.
Yerushalmi R, Woods R, Ravdin PM et al, Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 2010; 11: 174–183.
Geyer FC, Rodrigues DN, Weigelt B et al, Molecular classification of estrogen receptor-positive/luminal breast cancers. Adv Anat Pathol 2012; 19: 39–53.
Colozza M, Sidoni A, Piccart-Gebhart M . Value of Ki67 in breast cancer: the debate is still open. Lancet Oncol 2010; 11: 414–415.
Bevilacqua P, Verderio P, Barbareschi M et al, Lack of prognostic significance of the monoclonal antibody Ki-S1, a novel marker of proliferative activity, in node-negative breast carcinoma. Breast Cancer Res Treat 1996; 37: 123–133.
Aleskandarany MA, Rakha EA, Macmillan RD et al, MIB1/Ki-67 labelling index can classify grade 2 breast cancer into two clinically distinct subgroups. Breast Cancer Res Treat 2011; 127: 591–599.
Veronese SM, Gambacorta M, Gottardi O et al, Proliferation index as a prognostic marker in breast cancer. Cancer 1993; 71: 3926–3931.
Jonat W, Arnold N . Is the Ki-67 labelling index ready for clinical use? Ann Oncol 2011; 22: 500–502.
Allred DC, Carlson RW, Berry DA et al, NCCN Task Force Report: estrogen receptor and progesterone receptor testing in breast cancer by immunohistochemistry. J Natl Compr Canc Netw 2009; 7(Suppl 6):S1–S21.
Guiu S, Michiels S, Andre F et al, Molecular subclasses of breast cancer: how do we define them? The IMPAKT 2012 Working Group Statement. Ann Oncol 2012; 23: 2997–3006.
Reena RM, Mastura M, Siti-Aishah MA et al, Minichromosome maintenance protein 2 is a reliable proliferative marker in breast carcinoma. Ann Diagn Pathol 2008; 12: 340–343.
Laskey R . The Croonian Lecture 2001 hunting the antisocial cancer cell: MCM proteins and their exploitation. Philos Trans R Soc Lond B Biol Sci 2005; 360: 1119–1132.
Wharton SB, Hibberd S, Eward KL et al, DNA replication licensing and cell cycle kinetics of oligodendroglial tumours. Br J Cancer 2004; 91: 262–269.
Stoeber K, Tlsty TD, Happerfield L et al, DNA replication licensing and human cell proliferation. J Cell Sci 2001; 114: 2027–2041.
Yang C, Wen Y, Li H et al, Overexpression of minichromosome maintenance 2 predicts poor prognosis in patients with gastric cancer. Oncol Rep 2012; 27: 135–142.
Liu Y, He G, Wang Y et al, MCM-2 is a therapeutic target of Trichostatin A in colon cancer cells. Toxicol Lett 2013; 221: 23–30.
Abdou AG, Elwahed MG, Serag El-Dien MM et al, Immunohistochemical expression of MCM2 in nonmelanoma epithelial skin cancers. Am J Dermatopathol 2014; 36: 959–964.
Cho Mar K, Eimoto T, Nagaya S et al, Cell proliferation marker MCM2, but not Ki67, is helpful for distinguishing between minimally invasive follicular carcinoma and follicular adenoma of the thyroid. Histopathology 2006; 48: 801–807.
Guzinska-Ustymowicz K, Pryczynicz A, Kemona A et al, Correlation between proliferation markers: PCNA, Ki-67, MCM-2 and antiapoptotic protein Bcl-2 in colorectal cancer. Anticancer Res 2009; 29: 3049–3052.
Wojnar A, Kobierzycki C, Krolicka A et al, Correlation of Ki-67 and MCM-2 proliferative marker expression with grade of histological malignancy (G) in ductal breast cancers. Folia Histochem Cytobiol 2010; 48: 442–446.
Jezequel P, Frenel JS, Campion L et al, bc-GenExMiner 3.0: new mining module computes breast cancer gene expression correlation analyses. Database (Oxford) 2013; 2013: bas060.
Rosenbloom KR, Armstrong J, Barber GP et al, The UCSC Genome Browser database: 2015 update. Nucleic Acids Res 2015; 43: D670–D681.
Madden SF, Clarke C, Gaule P et al, BreastMark: an integrated approach to mining publicly available transcriptomic datasets relating to breast cancer outcome. Breast Cancer Res 2013; 15: R52.
Tavassoli FA, Devilee P (eds).World health organization classification of tumours. In: Pathology and Genetics Tumours of the Breast and Female Genital Organs. IARC Press: Lyon, France, 2003, pp19–23.
Zhao S, Ma W, Zhang M et al, High expression of CD147 and MMP-9 is correlated with poor prognosis of triple-negative breast cancer (TNBC) patients. Med Oncol 2013; 30: 335.
Wang Y, Zhou ZG, Xia QJ et al, Expression of minichromosome maintenance protein 2 in colonic adenocarcinoma, adenoma and normal colonic mucosa and its clinical significance. Zhonghua Wei Chang Wai Ke Za Zhi 2008; 11: 465–468.
The Human Protein Atlas. Available at:http://www.proteinatlasorg.
Yousef EM, Tahir MR, St-Pierre Y et al, MMP-9 expression varies according to molecular subtypes of breast cancer. BMC Cancer 2014; 14: 609.
Gonzalez MA, Pinder SE, Callagy G et al, Minichromosome maintenance protein 2 is a strong independent prognostic marker in breast cancer. J Clin Oncol 2003; 21: 4306–4313.
Malhotra A, Younes M, Kuna ST et al, Performance of an automated polysomnography scoring system versus computer-assisted manual scoring. Sleep 2013; 36: 573–582.
Singh G . Determination of Cutoff Score for a Diagnostic Test. Internet J Lab Med 2006; 2: 1–4.
Zhu W, Zeng N, Wang N . Sensitivity, specificity, accuracy, associated confidence interval and ROC analysis with practical SAS® implementations. In: NESUG Proceedings: Health Care and Life Sciences, Baltimore, MD,2010, pp1–9.
Shetty A, Loddo M, Fanshawe T et al, DNA replication licensing and cell cycle kinetics of normal and neoplastic breast. Br J Cancer 2005; 93: 1295–1300.
Lopez F, Belloc F, Lacombe F et al, Modalities of synthesis of Ki67 antigen during the stimulation of lymphocytes. Cytometry 1991; 12: 42–49.
Scholzen T, Gerdes J . The Ki-67 protein: from the known and the unknown. J Cell Physiol 2000; 182: 311–322.
Alison MR, Hunt T, Forbes SJ . Minichromosome maintenance (MCM) proteins may be pre-cancer markers. Gut 2002; 50: 290–291.
Ali HR, Dawson SJ, Blows FM et al, Aurora kinase A outperforms Ki67 as a prognostic marker in ER-positive breast cancer. Br J Cancer 2012; 106: 1798–1806.
Joshi S, Watkins J, Gazinska P et al, Digital imaging in the immunohistochemical evaluation of the proliferation markers Ki67, MCM2 and Geminin, in early breast cancer, and their putative prognostic value. BMC Cancer 2015; 15: 546.
Keam B, Im SA, Lee KH et al, Ki-67 can be used for further classification of triple negative breast cancer into two subtypes with different response and prognosis. Breast Cancer Res 2011; 13: R22.
Ribelles N, Perez-Villa L, Jerez JM et al, Pattern of recurrence of early breast cancer is different according to intrinsic subtype and proliferation index. Breast Cancer Res 2013; 15: R98.
Li FY, Wu SG, Zhou J et al, Prognostic value of Ki-67 in breast cancer patients with positive axillary lymph nodes: a retrospective cohort study. PLoS One 2014; 9: e87264.
Ferguson NL, Bell J, Heidel R et al, Prognostic value of breast cancer subtypes, Ki-67 proliferation index, age, and pathologic tumor characteristics on breast cancer survival in Caucasian women. Breast J 2013; 19: 22–30.
Kwok HF, Zhang SD, McCrudden CM et al, Prognostic significance of minichromosome maintenance proteins in breast cancer. Am J Cancer Res 2015; 5: 52–71.
Bessarabova M, Kirillov E, Shi W et al, Bimodal gene expression patterns in breast cancer. BMC Genomics 2010; 11: S8.
Blow JJ, Hodgson B . Replication licensing—defining the proliferative state? Trends Cell Biol 2002; 12: 72–78.
van Dierendonck JH, Keijzer R, van de Velde CJ et al, Nuclear distribution of the Ki-67 antigen during the cell cycle: comparison with growth fraction in human breast cancer cells. Cancer Res 1989; 49: 2999–3006.
Gerdes J, Lemke H, Baisch H et al, Cell cycle analysis of a cell proliferation-associated human nuclear antigen defined by the monoclonal antibody Ki-67. J Immunol 1984; 133: 1710–1715.
Bruno S, Darzynkiewicz Z . Cell cycle dependent expression and stability of the nuclear protein detected by Ki-67 antibody in HL-60 cells. Cell Prolif 1992; 25: 31–40.
du Manoir S, Guillaud P, Camus E et al, Ki-67 labeling in postmitotic cells defines different Ki-67 pathways within the 2c compartment. Cytometry 1991; 12: 455–463.
Starborg M, Gell K, Brundell E et al, The murine Ki-67 cell proliferation antigen accumulates in the nucleolar and heterochromatic regions of interphase cells and at the periphery of the mitotic chromosomes in a process essential for cell cycle progression. J Cell Sci 1996; 109: 143–153.
Heidebrecht HJ, Buck F, Haas K et al, Monoclonal antibodies Ki-S3 and Ki-S5 yield new data on the 'Ki-67' proteins. Cell Prolif 1996; 29: 413–425.
Booth DG, Takagi M, Sanchez-Pulido L et al, Ki-67 is a PP1-interacting protein that organises the mitotic chromosome periphery. Elife 2014; 3: e01641.
Acknowledgements
We thank Mrs Julie Hinsinger for help with Visiomorph, Tissuemorph Digital Pathology software, Mrs Micheline Fortin for expert technical assistance and Mr Romain Sabina of the bio banking facility. We are also grateful to Dr. Simon Jacob for the confirmation of the molecular subtypes of the Discovery cohort. This work was supported by funds from the Canadian Cancer Research Institute to LG and SM, the Institute for Research in Immunology and Cancer (IRIC) to LG, the Faculty of Graduate and Postdoctoral Studies of Université de Montreal and The Egyptian Supreme Council of Universities to EY. The above-mentioned parties had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Discovery specimens were provided by the Fondation du cancer du sein du Québec and the Banque de tissus et de données of the Réseau de recherche sur le cancer of the FRQS, which is affiliated with the Canadian Tumour Repository Network.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Yousef, E., Furrer, D., Laperriere, D. et al. MCM2: An alternative to Ki-67 for measuring breast cancer cell proliferation. Mod Pathol 30, 682–697 (2017). https://doi.org/10.1038/modpathol.2016.231
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.231
This article is cited by
-
RANKL inhibition reduces lesional cellularity and Gαs variant expression and enables osteogenic maturation in fibrous dysplasia
Bone Research (2024)
-
FAM83D acts as an oncogene by regulating cell cycle progression via multiple pathways in synovial sarcoma: a potential novel downstream target oncogene of anlotinib
Discover Oncology (2024)
-
Analysis of AT7519 as a pro-resolution compound in an acetaminophen-induced mouse model of acute inflammation by UPLC-MS/MS
Journal of Inflammation (2023)
-
A differential diagnosis between uterine leiomyoma and leiomyosarcoma using transcriptome analysis
BMC Cancer (2023)
-
A novel nomogram based on cell cycle-related genes for predicting overall survival in early-onset colorectal cancer
BMC Cancer (2023)