Abstract
Specialized stromal tumors of the prostate encompass stromal sarcoma and stromal tumors of uncertain malignant potential (STUMP). The molecular signature associated with stromal sarcoma and STUMP has not been unraveled. The study was conducted to detect the chromosomal imbalances in stromal sarcoma and STUMP by using array comparative genomic hybridization (aCGH). The study consisted of two cases of stromal nodule, eight cases of STUMP (three degenerative atypia type, three myxoid type, one hypercellular type, and one phyllodes type), and four cases of stromal sarcoma, including a distant metastasis developed metachronously after a primary stromal sarcoma of the prostate. DNA was extracted from the representative paraffin-embedded formalin-fixed specimens and was submitted for aCGH. All stromal sarcomas and seven STUMPs revealed chromosomal aberrations. Overall, the most common alteration was loss of chromosome 13 (10 cases), followed by loss of chromosome 14 (9 cases), and loss of chromosome 10 (7 cases). Except one stromal sarcoma, which showed a distinct chromosomal profile of multiple amplifications, other stromal sarcomas showed a similar pattern to those of STUMP. Stromal sarcoma and STUMP shared similar profiles of chromosomal imbalances. From a molecular genetic perspective, the recurrent chromosomal alterations support the concept of specialized stromal tumors of the prostate as a distinctive tumor entity.
Similar content being viewed by others
Main
Prostatic stromal tumors arising from the specialized prostatic stroma are rare and distinct tumors with diverse histological patterns.1, 2 In the past, these tumors have been reported under a variety of terms including atypical stromal (smooth muscle) hyperplasia, phyllodes type of atypical stromal hyperplasia, and cystic epithelial–stromal tumors.3, 4, 5, 6 In 1998, a classification of these stromal lesions was proposed, identifying (1) prostatic stromal sarcoma and (2) stromal tumors of uncertain malignant potential (STUMP).1 Implicit in this classification is that all such atypical stromal proliferations are neoplastic. Four distinct histological patterns, namely, degenerative atypia pattern, hypercellular pattern, myxoid pattern, and phyllodes-type growth pattern, were recognized. Stromal sarcomas are subtyped as low grade and high grade on the basis of cellular pleomorphism and degree of cellularity. Follow-up of these patients revealed that these neoplasms are associated with unique local morbidity and malignant potential.2 STUMP can recur frequently, occur at a younger age, predominantly involve the peripheral zone where they can adhere to the rectum, requiring its removal, and dedifferentiate to or be concurrently associated with stromal sarcoma. Low-grade stromal sarcoma can locally invade and high-grade sarcoma has the potential to metastasize.
Although the histological and immunohistochemical patterns of STUMP and stromal sarcoma have been characterized, the cytogenetics of these tumors are largely unknown.7, 8 In this study, we aimed to delineate the chromosomal profile of STUMP and stromal sarcomas using array comparative genomic hybridization (aCGH). This molecular genetic study is helpful to clarify the neoplastic nature of the disease. The series recruited nine patients from the pathology consult file of the Johns Hopkins Hospital and two patients from the surgical pathology file of Taipei Veterans General Hospital. The study abided by the regulations of the institutional review board of Taipei Veterans General Hospital.
Materials and methods
Sample Collection
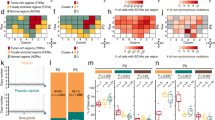
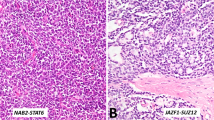
Initially, we collected tumor samples from 13 patients with STUMP and 4 patients with stromal sarcoma from Johns Hopkins Hospital, MD, and 2 patients with stromal sarcoma from Taipei Veterans General Hospital, Taiwan. The cases of the Johns Hopkins Hospital were reported in previous studies. One patient of the Taipei Veterans General Hospital had samples of both primary and metastatic tumors (a distant metastasis in abdominal cavity 2 years after prostatectomy) for analysis. The patients’ ages ranged from 42–66 (mean, 56) years. The tumors were subtyped histologically as previously described (Figure 1).2 The samples were fixed in formalin and embedded in paraffin. The paraffin blocks were punctured at the most suitable site for DNA extraction corresponding to the most cellular area of the tumor based on hematoxylin and eosin-stained slides. The tumor cells occupied at least 80% of the selected areas. Cases where tumor cells were too infiltrative to be separated from non-neoplastic prostatic glands were excluded. Finally, 12 (eight STUMPs and four stromal sarcomas) of these samples had DNA of sufficient quality, and were used for subsequent analysis. Two cases of hyperplastic stromal nodule were run in parallel.
Histology of (a) degenerative atypia type, (b) hypercellular type, (c) myxoid type and (d) phyllodes type of stromal tumor of uncertain malignant potential, (e) low-grade stromal sarcoma, (f) high-grade stromal sarcoma, and (g) the metastatic high-grade stromal sarcoma. One peculiar high-grade stromal sarcoma (h) composed of spindle cells and myxoid areas showed distinct chromosomal profile.
DNA Extraction
Fomalin-fixed paraffin-embedded tissues were deparaffinized using xylene and washed in ethanol. A 1 M solution of sodium thiocyanate was added to the dry pellet and incubated at 37 °C overnight to remove crosslinks. After removal of the sodium thiocyanate, a QIAamp (Qiagen, Foster City, CA) extraction procedure was performed according to the manufacturer’s instruction. After adsorbed onto a QIAamp silica membrane, the DNA was washed and eluted. The quantity (OD 260 nm) and quality (OD 260 nm/OD 280 nm) of DNA in the obtained solution were measured using an ND-1000 spectrophotometer (Nanodrop Technology, Wilmington, DE). The integrity of the DNA was determined using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA). DNA samples with a length profile showing a sufficient amount of single-stranded DNA of over 500 bp in length were used of aCGH analysis.
aCGH analysis
Genomic DNA (2 μg) isolated from the tissue samples and DNA from healthy tissue (Promega Corporation, Madison, WI) were labeled with Cy5 and Cy3, respectively, using an Oligo aCGH Labeling Kit (Agilent Technologies). The Cy5-labeled and Cy3-labeled samples were purified using Agilent-KREApure columns (Agilent Technologies). The appropriately purified and labeled samples were processed according to the manufacturer’s protocol. Before hybridization to the array, the hybridization mixtures were denatured at 95 °C for 3 min and incubated at 37 °C for 30 min, and the samples were then applied to the Agilent Genome CGH Microarray 244 K (Agilent Technologies). Hybridization was carried out for 40 h at 65 °C. After washing, the slides were scanned on an Agilent microarray scanner. The images were analyzed using Feature Extraction software, version 10.5.1.1 (Agilent Technologies) and Agilent Genomic Workbench software, version 7.0.4.0 (Agilent Technologies) using the ADM-2 algorithm with the following settings: 2 Mb moving average; threshold, 6; filter, 3 probes; and 0.4 log2 ratio.
Results
The histological patterns, tissue sources, and alterations in copy number are listed in Table 1. Seven of the eight STUMPs and all stromal sarcomas revealed chromosomal aberrations. One myxoid STUMP (case 4) showed no change in copy number. Overall, the most common alteration was loss of chromosome 13 (10 cases), followed by loss of chromosome 14 (9 cases), and partial or complete loss of chromosome 10 (7 cases). Less frequent losses were noted on chromosomes 1p (5 cases), 22 (4 cases) and 16q (3 cases). Chromosomal gains were uncommon in STUMPs but more common in stromal sarcomas. The occurrence of chromosomal alteration was observed in all histological subtypes of STUMP. One low-grade stromal sarcoma (case 9) revealed a similar pattern to those of STUMP, involving a total of six chromosomes. The total aberrations involved 19 chromosomes in one high-grade stromal sarcoma (case 10). The metastatic tumor (case 10 m) of this case manifested almost an identical aCGH profile as that of the primary tumor. The other high-grade stromal sarcoma (case 11) showed a distinct chromosomal profile of multiple amplification affecting chromosomes 8q, 12, 18, 19, 20, X, and Y. Morphologically, this peculiar sarcoma was composed of a mixture of spindle cells and myxoid areas (Figure 1h). The tumor cells were positive for CD34 immunohistochemically, but negative for CD117, S100 protein, and actin. A diagnosis of high-grade prostatic stromal sarcoma was rendered on the basis of histology and immunoprofile, as well as its prostatic location without other primary sites. The patient developed liver metastasis 8 months later and died of the disease 10 months after the initial diagnosis. Examples of aCGH profiles of STUMP and stromal sarcoma are shown in Figures 2 and 3, respectively. The accumulative penetrance plot is demonstrated in Figure 4. The two cases of stromal nodule showed balanced dosages of chromosomes.
One low-grade stromal sarcoma (case 9) reveals a chromosomal profile comparable to stromal tumor of uncertain malignant potential. One high-grade stromal sarcoma (case 10) reveals multiple chromosomal aberrations including losses of chromosomes 10, 13, and 14. Note that the chromosomal profiles of case 10 and its metastasis (case 10 m) are generally the same. The other stromal sarcoma (case 11) shows different chromosomal changes manifested as multiple gains.
Discussion
The genetic profiles of STUMP and stromal sarcoma have not been delineated. One study employing loss of heterozygosity analyses on six phyllodes tumor of the prostate8 and one report of aCGH on a single case of phyllodes tumor were published.7 In the former study, McCarthy et al.8 demonstrated that both epithelium and stroma in phyllodes tumor are clonal proliferations, and they appear to have different clonal origins. In the latter report, Fujii et al.7 did not identify genome-wide chromosomal changes. No molecular genetic or cytogenetic studies were performed on stromal sarcoma and other subtypes of STUMP.
In this study, we found recurrent chromosomal changes in both the STUMP and stromal sarcoma. This finding further confirms the clonal nature of this disease, and these lesions should be interpreted as being neoplastic rather than hyperplastic. It should be noted that these chromosomal alterations were not specific for a certain histological subtype, but were observed in all of them. The presence of similar chromosomal changes in the ‘degenerative atypia’ subtype as in other subtypes indicates that such lesion is a true neoplasm rather than a non-neoplastic degenerative process. The detection of similar aberration in one phyllodes tumor (case 8) supports the inclusion of this subtype into the category of STUMP.
It is intriguing that stromal sarcoma shared common chromosomal alterations with STUMP. This observation implies that stromal sarcoma and STUMP belong to the same family of neoplasm, and probably represent a spectrum. This finding is concordant with the coexistence of STUMP and stromal sarcoma observed in some previous series. Given the more ominous histological features and aggressive behavior of the stromal sarcoma compared with STUMP,2 there plausibly exist yet unraveled molecular events in the progression from STUMP to stromal sarcoma. One may intuitively speculate that stromal sarcomas bear more extensive genetic abnormalities. One of the high-grade stromal sarcomas in this series had wider aberrations involving numerous chromosomes. It is noted that this disordered chromosomal pattern was highly preserved in the metastasis that occurred 2 years after the primary tumor. On the contrary, the low-grade stromal sarcoma showed comparable changes with those of STUMP. However, the case number is too low to draw any conclusion at present.
Chromosomal regions that were most frequently lost included 13, 14 and 10. Complete or partial losses of these chromosomes have been reported in various types of cancers.9, 10, 11, 12, 13, 14, 15 Losses in 10q and 13q are frequently detected in the leiomyosarcomas and malignant fibrous histiocytomas.13, 14, 15 However, those soft tissue sarcomas usually reveal complex genomic profiles without consistent aberration pattern. Loss of chromosome 14 appears to be a relatively unique anomaly for prostatic stromal neoplasms, as chromosome 14 is not commonly deleted in other soft tissue sarcomas.13
The regions delineated by our assay are too broad to allow the identification of specific candidate genes. Target tumor suppressor genes that could be inactivated through 13q deletion include the retinoblastoma gene16 and the BRCA-2 gene,17 on 13q14, and 13q12, respectively. KLF6, located on 10p15, has been demonstrated as a candidate tumor suppressor gene often mutated in prostate cancer.18, 19 Three tumor suppressor genes residing on chromosome 10q (ERCC6, PTEN/MMAC1, and DMBT1) have been implicated in the development and progression of several types of neoplasms.20, 21, 22 High frequency of allelic loss on chromosome 10q23, where PTEN/MMAC1 gene locates was also reported in phyllodes tumor of the prostate.8 No tumor suppressor genes have been well defined on chromosome 14, but putative tumor suppressor loci had been mapped by using microsatellite allotyping.23 Whether these genes participate in the oncogenesis of prostatic stromal tumors requires further investigations.
One STUMP of myxoid pattern did not exhibit gross chromosomal imbalances. Morphologically, this case could not be discerned from other myxoid STUMPs harboring chromosomal changes. aCGH detects only large structural or numeric changes of chromosomes; therefore, we could not exclude the possibility of small genetic mutations in that case. One high-grade stromal sarcoma showed an entirely disparate aCGH profile from other cases. That singular case was characterized by multiple chromosomal over-representation rather than under-representation. There likely exists a minor subset of stromal neoplasms that possess different somatic genetics.
Our study included two samples of stromal nodule of nodular hyperplasia. No gene dosage changes were discerned. The absence of genetic alteration in hyperplasia is compatible with the non-neoplastic nature of the lesion, and lends further credence to the hypothesis that stromal neoplasms and stromal hyperplasia are two fundamentally divergent processes.
A limitation of the present study relates to the clonal nature of epithelial component of STUMP. The present assay cannot determine whether the epithelial cells are of the same clone as the stromal component. As aCGH requires larger amount of DNA and epithelial cells is a minor population compared with stromal cells, it is difficult to purify sufficient DNA of epithelial cells from the paraffin sections. We chose the areas where neoplastic stromal cells accounted for more than 80% of the lesion; consequently, the aCGH profiles mostly reflect the clonality of stromal cells.
In conclusion, prostatic specialized stromal neoplasms share similar profiles of chromosomal imbalances (mainly deletions of chromosomes 13, 14, and 10). The chromosomal aberrations can be observed in all subtypes of STUMP and stromal sarcoma. From a molecular genetic perspective, the common chromosomal alterations support the concept of specialized stromal tumors of the prostate as a distinctive tumor entity.
References
Gaudin PB, Rosai J, Epstein JI . Sarcomas and related proliferative lesions of specialized prostatic stroma: a clinicopathologic study of 22 cases. Am J Surg Pathol 1998;22:148–162.
Herawi M, Epstein JI . Specialized stromal tumors of the prostate: a clinicopathologic study of 50 cases. Am J Surg Pathol 2006;30:694–704.
Attah EB, Powell ME . Atypical stromal hyperplasia of the prostate gland. Am J Clin Pathol 1977;67:324–327.
Bostwick DG, Hossain D, Qian J et al. Phyllodes tumor of the prostate: long-term followup study of 23 cases. J Urol 2004;172:894–899.
Kevwitch MK, Walloch JL, Waters WB et al. Prostatic cystic epithelial-stromal tumors: a report of 2 new cases. J Urol 1993;149:860–864.
Sauder KJ, MacLennan GT . Phyllodes tumor of the prostate. J Urol 2005;174:2373.
Fujii T, Shimada K, Tanaka N et al. Phyllodes tumor of the prostate. Pathol Int 2012;62:204–208.
McCarthy RP, Zhang S, Bostwick DG et al. Molecular genetic evidence for different clonal origins of epithelial and stromal components of phyllodes tumor of the prostate. Am J Pathol 2004;165:1395–1400.
Jiang F, Moch H, Richter J et al. Comparative genomic hybridization reveals frequent chromosome 13q and 4q losses in renal carcinomas with sarcomatoid transformation. J Pathol 1998;185:382–388.
Petersen S, Wolf G, Bockmuhl U et al. Allelic loss on chromosome 10q in human lung cancer: association with tumour progression and metastatic phenotype. Br J Cancer 1998;77:270–276.
Shaughnessy J, Tian E, Sawyer0. J et al. High incidence of chromosome 13 deletion in multiple myeloma detected by multiprobe interphase FISH. Blood 2000;96:1505–1511.
Williams RD, Al-Saadi R, Natrajan R et al. Molecular profiling reveals frequent gain of MYCN and anaplasia-specific loss of 4q and 14q in Wilms tumor. Genes Chromosomes Cancer 2011;50:982–995.
Gibault L, Perot G, Chibon F et al. New insights in sarcoma oncogenesis: a comprehensive analysis of a large series of 160 soft tissue sarcomas with complex genomics. J Pathol 2011;223:64–71.
Larramendy ML, Kaur S, Svarvar C et al. Gene copy number profiling of soft-tissue leiomyosarcomas by array-comparative genomic hybridization. Cancer Genet Cytogenet 2006;169:94–101.
Larramendy ML, Gentile M, Soloneski S et al. Does comparative genomic hybridization reveal distinct differences in DNA copy number sequence patterns between leiomyosarcoma and malignant fibrous histiocytoma? Cancer Genet Cytogenet 2008;187:1–11.
Cance WG, Brennan MF, Dudas ME et al. Altered expression of the retinoblastoma gene product in human sarcomas. N Engl J Med 1990;323:1457–1462.
Venkitaraman AR . Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell 2002;108:171–182.
Fukuhara H, Maruyama T, Nomura S et al. Functional evidence for the presence of tumor suppressor gene on chromosome 10p15 in human prostate cancers. Oncogene 2001;20:314–319.
Narla G, Heath KE, Reeves HL et al. KLF6, a candidate tumor suppressor gene mutated in prostate cancer. Science 2001;294:2563–2566.
Aghi MK, Batchelor TT, Louis DN et al. Decreased rate of infection in glioblastoma patients with allelic loss of chromosome 10q. J Neurooncol 2009;93:115–120.
Li J, Yen C, Liaw D et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997;275:1943–1947.
Sasaki H, Betensky RA, Cairncross JG et al. DMBT1 polymorphisms: relationship to malignant glioma tumorigenesis. Cancer Res 2002;62:1790–1796.
Bandera CA, Takahashi H, Behbakht K et al. Deletion mapping of two potential chromosome 14 tumor suppressor gene loci in ovarian carcinoma. Cancer Res 1997;57:513–515.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Pan, CC., Epstein, J. Common chromosomal aberrations detected by array comparative genomic hybridization in specialized stromal tumors of the prostate. Mod Pathol 26, 1536–1543 (2013). https://doi.org/10.1038/modpathol.2013.99
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.99
Keywords
This article is cited by
-
Re-evaluating tumors of purported specialized prostatic stromal origin reveals molecular heterogeneity, including non-recurring gene fusions characteristic of uterine and soft tissue sarcoma subtypes
Modern Pathology (2021)
-
Reexamining the molecular findings in specialized stromal tumors of the prostate
Modern Pathology (2021)
-
In Response to “Reexamining the molecular findings in specialized stromal tumors of the prostate”
Modern Pathology (2021)
-
Clinicopathological features to distinguish malignant solitary fibrous tumors of the prostate from prostatic stromal tumors
Virchows Archiv (2021)
-
Mesenchymal tumors of the prostate
Modern Pathology (2018)