Abstract
AT-rich interactive domain 1A (ARID1A) is a subunit of switch/sucrose non-fermentable (SWI/SNF) complex. Recently, alterations of ARID1A gene, phosphatidylinositol 3-kinase-protein kinase B (PI3K-Akt) pathway and zinc-finger protein 217 (ZNF217) gene have been identified as frequent molecular genetic changes in ovarian clear cell carcinoma. The relationships between these events have not been studied and integrated in the same cohort. This study was aimed at determining the correlation between these molecular events and other clinicopathological factors, including the prognostic impacts of these clinicopathological factors. A total of 68 ovarian clear cell carcinoma cases were collected and subjected to immunohistochemistry testing for ARID1A, SMARCA2, SMARCA4, SMARCB1 and phosphatase and tensin homolog (PTEN), mutation analysis for phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit alpha (PIK3CA) gene and fluorescence in situ hybridization for ZNF217 amplification. The correlations between ARID1A expression, PI3K-Akt pathway, ZNF217 amplification and other clinicopathological factors were analyzed. Loss of ARID1A expression was present in 35 cases (52%) and loss of SMARCA2 expression occurred in 1 case. SMARCA4 and SMARCB1 expressions were preserved in all cases. PIK3CA mutations were present in 23 cases (34%) and loss of PTEN expression occurred in 8 cases (12%). Alterations in the PI3K-Akt pathway (PIK3CA mutations or loss of PTEN expression) were found in 42 cases (62%). ZNF217 amplification was detected in 21 cases (31%). Loss of ARID1A expression was significantly related to younger patient age (P=0.048), PI3K-Akt pathway activation (P=0.046) and ZNF217 amplification (P=0.028). All of the clinicopathological factors were not prognostic factors for ovarian clear cell carcinoma after multivariate analysis, except International Federation of Gynecology and Obstetrics staging (P=0.001). Our results showed that loss of ARID1A expression usually coexisted with PI3K-Akt pathway activation and/or ZNF217 amplification. Synergic effects of loss of ARID1A and PI3K-Akt pathway activation as well as ZNF217 amplification may be related to the development of ovarian clear cell carcinoma.
Similar content being viewed by others
Main
The prevalence of ovarian clear cell carcinoma in North America and Europe is 1–12%.1 Compared with other epithelial ovarian cancers, patients with clear cell carcinoma respond less favorably to platinum-based first-line chemotherapy.2 Elucidation of common molecular genetic changes is crucial for understanding this disease and new treatment strategies.
AT-rich interactive domain 1A (ARID1A) is one of the ARID family members with non-sequence-specific DNA-binding activity, and is the large subunit of switch/sucrose non-fermentable (SWI/SNF) complex.3 Brahma (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 2, SMARCA2), brahma-related gene 1 (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4, SMARCA4) and integrase interactor 1 (SWI/SNF-related, matrix-associated, actin-dependent regulator of chromatin, subfamily b, member 1, SMARCB1) are other important components involved in SWI/SNF complexes.4 In humans, ARID1A is a tumor-suppressor gene located at 1p36, which is frequently mutated in ovarian, endometrial, breast, urinary bladder and gastric cancers.5, 6, 7, 8, 9 ARID1A mutations are present in 46% of ovarian clear cell carcinomas and 30% of endometrioid carcinomas.5 ARID1A immunohistochemistry (IHC) is closely related to ARID1A mutational status,10 and can be used as a surrogate marker for ARID1A mutation.
In addition to ARID1A, activating mutations in phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit alpha (PIK3CA)11 and zinc-finger protein 217 (ZNF217) amplification12 are common molecular genetic alterations in ovarian clear cell carcinoma. PIK3CA activating mutations and ZNF217 amplification can be found in 40% of ovarian clear cell carcinoma patients.11, 12 Phosphatidylinositol 3-kinase-protein kinase B (PI3K-Akt) pathway activation can be caused by either PIK3CA mutations, phosphatase and tensin homolog (PTEN) mutations or a combination of these alterations.13 ZNF217 encodes a transcriptional repressor protein and forms the ZNF217 complex with CoREST, histone deacetylases HDAC1 and 2, and LSD-1.14
The biological roles of the SWI/SNF complex, PI3K-Akt pathway and ZNF217 amplification in ovarian clear cell carcinoma have been separately investigated by several groups. However, data of relationships between these molecular genetic alterations are limited. Therefore, we analyzed the SWI/SNF complex (including ARID1A, SMARCA2, SMARCA4 and SMARCB1), PI3K-Akt pathway (including PIK3CA mutations and PTEN expression) and ZNF217 amplification in 68 patients with ovarian clear cell carcinoma. We focused on the relationships between these molecular changes, their association to the clinicopathological factors and their prognostic impacts.
Materials and methods
Patients and Tissue Materials
All specimens and clinical data were collected from patients who received debulking surgery at National Taiwan University Hospital. Formalin-fixed paraffin-embedded tissue specimens were obtained from the archives of 1996–2010, at the Department of Pathology in the same institution. All 68 patients were treated surgically and 61 of them also received chemotherapy. The median follow-up period was 46.5 months (11.8–163.6 months), except for the patients who died. The sections stained with hematoxylin and eosin were reviewed and diagnosed as ovarian clear cell carcinoma according to the World Health Organization classification.15 To exclude high-grade serous adenocarcinoma, hepatocyte nuclear factor (HNF)-1beta, estrogen receptor (ER), progesterone receptor (PR), Wilms tumor 1 (WT1) and p53 immunostains were also performed in all cases.16, 17
Immunohistochemistry
Immunohistochemical staining of paraffin tissue sections was performed by the Leica BOND-III autostainer (Leica Microsystems, Buffalo Grove, IL, USA). Commercially available antibodies to ARID1A, SMARCA2, SMARCA4, SMARCB1 and PTEN were used. The manufacturers, clones and antibody dilutions are listed in Table 1. The ARID1A immunoreactivities were divided into undetectable or positive (weakly or strongly) for nuclear staining.18 We used similar criteria for SMARCA2, SMARCA4 and SMARCB1. PTEN immunoreactivity was applied by two-tiered systems (undetectable or positive) and vascular endothelial cells were used as an internal positive control.19
DNA Extraction and Mutation Analysis of the PIK3CA Gene
DNA extraction, polymerase chain reaction (PCR) amplification for PIK3CA gene sequencing and mutation analysis were performed in all 68 samples using approaches previously described.11, 20
Fluorescence In Situ Hybridization (FISH)
ZNF217 amplification was studied by FISH in all cases. The commercial probes (Kreatech, Amsterdam, The Netherlands) were used. The criterion for ZNF217 amplification was an amplification ratio of ≥1.5, enumerated among 100 tumor cells.
Statistical Analysis
The statistical analyses were conducted using PASW Statistics (IBM Corporation, Armonk, NY, USA). The association between ARID1A expression and age was evaluated using the t-test. The association between ARID1A expression and other clinicopathologic parameters was evaluated using the χ2 test or Fisher’s exact test. The influences of clinicopathological parameters on progression-free survival (PFS) and overall survival (OS) were analyzed by Cox proportional hazard model. P-values from Wald’s statistic were recorded. The cutoff of significance level was 0.05.
Results
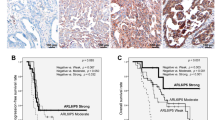
The clinical data and results of IHC, mutation analysis and FISH are listed in Table 2, and association between ARID1A expression and other clinicopathological factors are summarized in Table 3. Loss of ARID1A expression (35/68, 52%; Figures 1a and b), PIK3CA mutations (23/68, 34%) and ZNF217 amplification (21/68, 31%) were common in ovarian clear cell carcinoma. SMARCA4 and SMARCB1 expressions were preserved in all cases, but only one case showed loss of SMARCA2 expression (Figures 1c and d). Loss of PTEN expression was present in 12% of all cases (Figures 1e and f).
ARID1A, SMARCA2 and PTEN expressions in representative ovarian clear cell carcinoma cases. (a) A clear cell carcinoma with diffuse ARID1A immunoreactivity. (b) A clear cell carcinoma without ARID1A immunoreactivity. (c) Positive SMARCA2 immunostain. (d) Negative SMARCA2 immunostain. (e) Intact PTEN immunoreactivity. (f) Loss of PTEN immunoreactivity. Immunoperoxidase immunohistochemical stain, original magnification × 200.
The frequencies of PI3K-Akt pathway (PTEN expression/PIK3CA mutations) activation and ZNF217 amplification in the ARID1A-negative carcinomas were higher than those in the ARID1A-positive carcinomas (P=0.046 and 0.028, respectively). Loss of ARID1A expression was also correlated with younger patient age (P=0.048). PIK3CA mutations and loss of PTEN expression, respectively, were not associated with ARID1A expression (P=0.105 and 0.710, respectively).
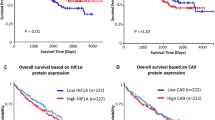
The prognostic effects of clinicopathological factors are summarized in Table 4. International Federation of Gynecology and Obstetrics (FIGO) staging was the most important prognostic predictor for ovarian clear cell carcinoma (P=0.001). Loss of PTEN expression also had impact on PFS and OS in univariate analysis (P=0.050 for PFS and 0.026 for OS), but its significance disappeared after multivariate analysis (P=0.144 for PFS and 0.360 for OS). Other clinicopathological factors including age, ARID1A expression, PIK3CA mutations, PI3K-Akt pathway and ZNF217 amplification were not significant prognostic factors.
Discussion
Compared with other epithelial ovarian cancers, clear cell carcinoma patients are more likely to be younger, of Asian descent, present at an early stage and suffer a poorer survival after adjusting other prognostic factors such as age and grade of disease.21 Owing to less favorable responses to platinum-based first-line chemotherapy in the patients of ovarian clear cell carcinoma,2 effort for discovering molecular genetic changes are ongoing. The major differential diagnosis of ovarian clear cell carcinoma in the adult is high-grade serous adenocarcinoma. Combined with morphological features, a panel of immunostains including HNF-1beta, ER, PR, WT-1 and p53 is helpful for the differential diagnosis.16, 17 To date, the mutations of chromatin-remodeling gene ARID1A, activating mutations in PIK3CA11 and ZNF217 amplification12 are the most common molecular genetic alterations in ovarian clear cell carcinoma. Loss of PTEN protein expression is also found in one-third of the ovarian clear cell carcinoma patients.22 To our knowledge, no study has reported the combination of all these common molecular genetic alterations in the same patient cohort.
The SWI/SNF complex in mammals is an ATP-dependent chromatin-remodeling complex crucial for cell proliferation, development and differentiation.23 ARID1A is one of the ARID family members with non-sequence-specific DNA-binding activity, and is the large subunit of SWI/SNF complex.3 Among the related transcription factors and transcriptional complexes, SMARCA4, SMARCA2 and SMARCB1 are important.4 ARID1A is a tumor-suppressor gene located at 1p36, which is frequently mutated in ovarian, endometrial, breast, urinary bladder and gastric cancers.5, 6, 7, 8, 9 In gynecologic cancers, ARID1A promotes formation of SWI/SNF-mediated chromatin remodeling and is present in 46% of ovarian clear cell carcinomas and 30% of endometrioid carcinomas.5, 24 In our study, we confirmed that loss of ARID1A expression was present in 52% of ovarian clear cell carcinomas. The result was close to those of previous studies by Maeda et al (59.1%) and Xiao et al (57.7%).10, 25 Compared with ARID1A, loss of expression of other components related to SWI/SNF complex such as SMARCA2, SMARCA4 and SMARCB1 were rare events in ovarian clear cell carcinoma. Similar findings were reported by Bosse et al in endometrial cancer.26 PI3K-Akt pathway activation is a novel molecular genetic change in ovarian clear cell carcinoma, and can be caused by either PIK3CA mutations, PTEN loss or a combination of these alterations.13 In this study, the ARID1A expression did not significantly correlate with PIK3CA mutations or PTEN expression alone. Although the frequency of PIK3CA mutations in the ARID1A-deficient carcinomas was higher than that in the ARID1A-intact carcinomas, it was not significantly different. Similar results have been reported in several studies, with or without statistical significance.27, 28 We also found that loss of ARID1A expression was related to younger patient age and activated PI3K-Akt pathway (activating PIK3CA mutations or loss of PTEN expression). A similar correlation between the ARID1A expression and PI3K-Akt pathway was noted in endometrial cancer but was not statistically significant.26
ZNF217 encodes a transcriptional repressor protein and forms the ZNF217 complex with CoREST, histone deacetylases HDAC1 and 2, and LSD-1.14 ZNF217 overexpression can cause cellular immortalization, telomerase repression, antiapoptosis and increased metastatic potential.29, 30 Recently, Krig et al31 reported that ZNF217 could activate the PI3K-Akt pathway by regulating ErbB3 expression in breast cancer cells. ZNF217 amplification and PIK3CA mutations were almost mutually exclusive in the study by Rahman et al.32 In our study, the correlation between PIK3CA mutations and ZNF217 amplification was not statistically significant (data not shown, P=0.541). PIK3CA mutations and ZNF217 amplification affected 38 cases, and 6 cases (16%) harbored both PIK3CA mutations and ZNF217 amplification. The results were not consistent with Rahman’s study.32 Compared with Rahman’s study, we used different FISH probes and different criteria for amplification.
If we combined activating PIK3CA mutations, ZNF217 amplification and loss of PTEN expression as indicators of activating Akt pathway, the relationship between ARID1A expression and the activating Akt pathway was more statistically significant (P=0.007) than the PI3K-Akt pathway (P=0.048). The results clearly pointed out that cases with activating Akt pathway (PIK3CA mutations, ZNF217 amplification or loss of PTEN expression) had a higher frequency of loss of ARID1A expression (64%) than those without PI3K-Akt pathway alteration (31%). Our study also revealed that ZNF217 amplification occurred significantly more frequently in the group with loss of ARID1A expression. This was the first report on the correlation between ARID1A expression and ZNF217 amplification. Further investigation is needed to find the mechanism.
Except for the FIGO staging, all of the other clinicopathological factors were not associated with PFS and OS after multivariate analysis. Loss of ARID1A expression was found to be an adverse prognostic factor in gastric cancer,9 cervical cancer33 and endometrial clear cell carcinoma.6 In ovarian clear cell carcinoma, the prognostic effects of ARID1A expression were inconsistent in different studies by Shih et al10 and Katagiri et al.34 Both our study and Shih’s study revealed that ARID1A expression was not a prognostic factor for ovarian clear cell carcinoma. Absence of prognostic significance of PIK3CA mutations and PI3K-Akt pathway was confirmed in our study as in the previous studies.10, 27, 35 In this study, we demonstrated that loss of PTEN expression was not an independent prognostic factor after multivariate analysis; this finding is similar to that of Ho et al.36 ZNF217 amplification was reported to be a poor prognostic factor in the study by Rahman et al,32 while in this study, we did not find the prognostic effect of ZNF217 amplification. In brief, FIGO staging was the most important and the only prognostic factor in our study.
Considering the complexities of the PI3K-Akt pathway, a combination of different therapeutic agents related to this pathway may be considered in order to get the greatest effect. One recent example involved combining two inhibitors targeting PI3K/AKT/mTOR and RAF/MEK/ERK pathways in ovarian cancer patients.37 Our study suggested that additional factors such as ZNF217 may also activate the PI3K-Akt pathway, providing a potential target for this dual-targeting strategy.
In conclusion, this is the first report demonstrating the correlation between ARID1A, PI3K-Akt pathway and ZNF217 in ovarian clear cell carcinoma, by integration of expression of ARID1A, SMARCA2, SMARCA4, SMARCB1 and PTEN, PIK3CA mutations, as well as ZNF217 amplification. More studies have to be conducted to clarify the truncated relationship between these common molecular genetic changes for a better understanding of the disease pathogenesis and subsequent therapeutic strategy.
References
Anglesio MS, Carey MS, Kobel M, et al. Vancouver Ovarian Clear Cell Symposium Speakers. Clear cell carcinoma of the ovary: a report from the first Ovarian Clear Cell Symposium, 24 June 2010. Gynecol Oncol 2011;121:407–415.
Pectasides D, Fountzilas G, Aravantinos G et al. Advanced stage clear-cell epithelial ovarian cancer: the Hellenic Cooperative Oncology Group experience. Gynecol Oncol 2006;102:285–291.
Dallas PB, Pacchione S, Wilsker D et al. The human SWI-SNF complex protein p270 is an ARID family member with non-sequence-specific DNA binding activity. Mol Cell Biol 2000;20:3137–3146.
Phelan ML, Sif S, Narlikar GJ et al. Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell 1999;3:247–253.
Wiegand KC, Shah SP, Al-Agha OM et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N Engl J Med 2010;363:1532–1543.
Fadare O, Renshaw IL, Liang SX . Does the loss of ARID1A (BAF-250a) expression in endometrial clear cell carcinomas have any clinicopathologic significance? A pilot assessment. J Cancer 2012;3:129–136.
Zhang X, Zhang Y, Yang Y et al. Frequent low expression of chromatin remodeling gene ARID1A in breast cancer and its clinical significance. Cancer Epidemiol 2012;36:288–293.
Gui Y, Guo G, Huang Y et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat Genet 2011;43:875–878.
Wang DD, Chen YB, Pan K et al. Decreased expression of the ARID1A gene is associated with poor prognosis in primary gastric cancer. PLoS One 2012;7:e40364.
Maeda D, Mao TL, Fukayama M et al. Clinicopathological significance of loss of ARID1A immunoreactivity in ovarian clear cell carcinoma. Int J Mol Sci 2010;11:5120–5128.
Kuo KT, Mao TL, Jones S et al. Frequent activating mutations of PIK3CA in ovarian clear cell carcinoma. Am J Pathol 2009;174:1597–1601.
Kuo KT, Mao TL, Chen X et al. DNA copy numbers profiles in affinity-purified ovarian clear cell carcinoma. Clin Cancer Res 2010;16:1997–2008.
Dobbin ZC, Landen CN . The importance of the PI3K/AKT/MTOR pathway in the progression of ovarian cancer. Int J Mol Sci 2013;14:8213–8227.
Quinlan KG, Verger A, Yaswen P et al. Amplification of zinc finger gene 217 (ZNF217) and cancer: when good fingers go bad. Biochim Biophys Acta 2007;1775:333–340.
Lee KR, Tavassoli FA, Part J et al. Surface epithelial-stromal tumours In: Tavassoli FA, Devilee P (eds). World Health Organization Classification of Tumours. Pathology and Genetics of Tumours of the Breast and Female Genital Organs. IARC Press: Lyon, 2003, pp 117–145.
DeLair D, Oliva E, Kobel M et al. Morphologic spectrum of immunohistochemically characterized clear cell carcinoma of the ovary: a study of 155 cases. Am J Surg Pathol 2011;35:36–44.
Kobel M, Kalloger SE, Carrick J et al. A limited panel of immunomarkers can reliably distinguish between clear cell and high-grade serous carcinoma of the ovary. Am J Surg Pathol 2009;33:14–21.
Maeda D, Shih I . Pathogenesis and the role of ARID1A mutation in endometriosis-related ovarian neoplasms. Adv Anat Pathol 2013;20:45–52.
Kurose K, Zhou XP, Araki T et al. Frequent loss of PTEN expression is linked to elevated phosphorylated Akt levels, but not associated with p27 and cyclin D1 expression, in primary epithelial ovarian carcinomas. Am J Pathol 2001;158:2097–2106.
Nakayama K, Nakayama N, Kurman RJ et al. Sequence mutations and amplification of PIK3CA and AKT2 genes in purified ovarian serous neoplasms. Cancer Biol Ther 2006;5:779–785.
Chan JK, Teoh D, Hu JM et al. Do clear cell ovarian carcinomas have poorer prognosis compared to other epithelial cell types? A study of 1411 clear cell ovarian cancers. Gynecol Oncol 2008;109:370–376.
Hashiguchi Y, Tsuda H, Inoue T et al. PTEN expression in clear cell adenocarcinoma of the ovary. Gynecol Oncol 2006;101:71–75.
Clapier CR, Cairns BR . The biology of chromatin remodeling complexes. Annu Rev Biochem 2009;78:273–304.
Guan B, Wang TL, Shih I . ARID1A, a factor that promotes formation of SWI/SNF-mediated chromatin remodeling, is a tumor suppressor in gynecologic cancers. Cancer Res 2011;71:6718–6727.
Xiao W, Awadallah A, Xin W . Loss of ARID1A/BAF250a expression in ovarian endometriosis and clear cell carcinoma. Int J Clin Exp Pathol 2012;5:642–650.
Bosse T, Ter Haar NT, Seeber LM et al. Loss of ARID1A expression and its relationship with PI3K-Akt pathway alterations, TP53 and microsatellite instability in endometrial cancer. Mod Pathol. 2013;26:1525–1535.
Yamamoto S, Tsuda H, Takano M et al. PIK3CA mutations and loss of ARID1A protein expression are early events in the development of cystic ovarian clear cell adenocarcinoma. Virchows Arch 2012;460:77–87.
Yamamoto S, Tsuda H, Takano M et al. Loss of ARID1A protein expression occurs as an early event in ovarian clear-cell carcinoma development and frequently coexists with PIK3CA mutations. Mod Pathol 2012;25:615–624.
Nonet GH, Stampfer MR, Chin K et al. The ZNF217 gene amplified in breast cancers promotes immortalization of human mammary epithelial cells. Cancer Res 2001;61:1250–1254.
Huang G, Krig S, Kowbel D et al. ZNF217 suppresses cell death associated with chemotherapy and telomere dysfunction. Hum Mol Genet 2005;14:3219–3225.
Krig SR, Miller JK, Frietze S et al. ZNF217, a candidate breast cancer oncogene amplified at 20q13, regulates expression of the ErbB3 receptor tyrosine kinase in breast cancer cells. Oncogene 2010;29:5500–5510.
Rahman MT, Nakayama K, Rahman M et al. Prognostic and therapeutic impact of the chromosome 20q13.2 ZNF217 locus amplification in ovarian clear cell carcinoma. Cancer 2012;118:2846–2857.
Cho H, Kim JS, Chung H et al. Loss of ARID1A/BAF250a expression is linked to tumor progression and adverse prognosis in cervical cancer. Hum Pathol 2013;44:1365–1374.
Katagiri A, Nakayama K, Rahman MT et al. Loss of ARID1A expression is related to shorter progression-free survival and chemoresistance in ovarian clear cell carcinoma. Mod Pathol 2012;25:282–288.
Rahman M, Nakayama K, Rahman MT et al. Clinicopathologic and biological analysis of PIK3CA mutation in ovarian clear cell carcinoma. Hum Pathol 2012;43:2197–2206.
Ho CM, Lin MC, Huang SH et al. PTEN promoter methylation and LOH of 10q22-23 locus in PTEN expression of ovarian clear cell adenocarcinomas. Gynecol Oncol 2009;112:307–313.
Shimizu T, Tolcher AW, Papadopoulos KP et al. The clinical effect of the dual-targeting strategy involving PI3K/AKT/mTOR and RAS/MEK/ERK pathways in patients with advanced cancer. Clin Cancer Res 2012;18:2316–2325.
Acknowledgements
This work was supported by National Taiwan University Hospital (NTUH100-S1520 to K-T Kuo) and Far Eastern Memorial Hospital (FEMH-99-D040 to W-C Huang).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Huang, HN., Lin, MC., Huang, WC. et al. Loss of ARID1A expression and its relationship with PI3K-Akt pathway alterations and ZNF217 amplification in ovarian clear cell carcinoma. Mod Pathol 27, 983–990 (2014). https://doi.org/10.1038/modpathol.2013.216
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.216
Keywords
This article is cited by
-
Treatment for ovarian clear cell carcinoma with combined inhibition of WEE1 and ATR
Journal of Ovarian Research (2023)
-
Clinical relevance of molecular characteristics in Burkitt lymphoma differs according to age
Nature Communications (2022)
-
Mismatch repair deficiency is associated with specific morphologic features and frequent loss of ARID1A expression in ovarian clear cell carcinoma
Diagnostic Pathology (2021)
-
ARID1A mutations and expression loss in non-small cell lung carcinomas: clinicopathologic and molecular analysis
Modern Pathology (2020)
-
Establishment and molecular characterization of a human ovarian clear cell carcinoma cell line (FDOV1)
Journal of Ovarian Research (2018)