Abstract
Copy number gains involving the long arm of chromosome 8, including high-level amplifications at 8q21 and 8q24, have been frequently reported in breast cancer. Although the role of the MYC gene as the driver of the 8q24 amplicon is well established, the significance of the 8q21 amplicon is less clear. The breast cancer cell line SK-BR-3 contains three separate 8q21 amplicons, the distal two of which correspond to putative target genes TPD52 and WWP1. To understand the effect of proximal 8q21 amplification on breast cancer phenotype and patient prognosis, we analyzed 8q21 copy number changes using fluorescence in situ hybridization (FISH) in a tissue microarray containing more than 2000 breast cancers. Amplification at 8q21 was found in 3% of tumors, and was associated with medullary type (P<0.03), high tumor grade (P<0.0001), high Ki67 labeling index (P<0.05), amplification of MYC (P<0.0001), HER2, MDM2, and CCND1 (P<0.05 each), as well as the total number of gene amplifications (P<0.0001). 8q21 copy number gains were significantly related to unfavorable patient outcome in univariate analysis. However, multivariate Cox regression analysis did not reveal an independent prognostic value of 8q21 amplification. The position of our FISH probe and data of a previously performed high-resolution CGH study in the breast cancer cell line SK-BR-3 involve TCEB1 and TMEM70 as new possible candidate oncogenes at 8q21 in breast cancer.
Similar content being viewed by others
Main
Structural and numerical alterations of chromosome 8 have been reported in up to 60% of breast cancers.1, 2 In the majority of cases, these alterations occur as low-level copy number changes, including partial or complete deletions of 8p and gains of 8q.3 Recurrent high-level amplifications have been found at 8p12, 8q21, and 8q24.4, 5 Gene amplification is an important mechanism for protein overexpression and oncogene activation in tumor cells.6 At 8q24, the transcription factor v-myc myelocytomatosis viral oncogene homolog (avian) (MYC) is generally accepted as the biologically relevant amplification target.7 Several studies have shown that MYC amplification occurs in approximately 5% of breast cancers and it has been linked with high grade, advanced tumor stage, and poor patient survival.8, 9, 10, 11 At 8p12, the fibroblast growth factor receptor (FGFR1) has been suggested as the candidate amplification target in breast and bladder cancer.12, 13 Kallioniemi et al14 first reported amplification of 8q21–q23 in breast carcinomas that occurred independently of MYC amplification. Subsequent studies showed that the amplicon was not restricted to breast cancer but also occurred in carcinomas of lung, bladder, and prostate.15, 16, 17 High-resolution array CGH in combination with interphase fluorescence in situ hybridization (FISH) showed a high variability of amplicons at 8q21–24 with several discontinuous target regions.18 Rodriguez et al19 identified three separate amplicons within 8q21 in the SK-BR-3 breast cancer cell line by using high-resolution BAC arrayCGH. Putative target genes of two distal regions include tumor protein D52 (TPD52) and the ubiquitin-protein ligase WWP1, the amplification of which has been confirmed in clinical breast cancer specimens.20, 21 In contrast, data on the prevalence and clinical relevance of the first (proximal) amplicon involving a 70–80 Mb stretch at 8q21 are lacking in breast cancer. To explore the potential significance of 8q21 amplification in breast cancer, we analyzed a tissue microarray containing more than 2000 breast cancer specimens using a FISH probe that maps to the center of the proximal 8q21 amplicon.
Materials and methods
Breast Cancer Tissue Microarray
The breast cancer tissue microarray used for this study has been described in detail.22 In brief, a total of 2197 formalin-fixed (buffered neutral aqueous 4% solution), paraffin-embedded tumors with a median patient age of 62 (range 26–101) years and a median follow-up time of 68 months (range 1–176) were assembled in a tissue microarray format (Table 1). We punched one tissue cylinder per case with a diameter of 0.6 mm from representative tumor areas of a ‘donor’ tissue block using a home-made semiautomatic robotic precision instrument. The histological grade was determined according to a modified scoring system by Elston and Ellis (BRE score).23 Several molecular data used in this study were available from previously published studies. These included amplification data obtained by FISH for HER2, MYC, CCND1, MDM2, and EGFR, as well as expression data obtained by immunohistochemistry for estrogen receptor (ER), progesteron receptor (PR), and Ki67.8, 22
The use of these human tissues for protein expression and FISH studies was approved by the local ethics committee of the University of Hamburg.
FISH Analysis
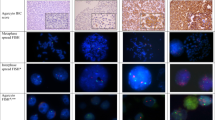
A FISH probe was generated from genomic clone RZPDB737E022003D for 8q21 containing the entire TMEM70 gene and part of the adjacent LY96 gene. The probe was labeled with digoxigenin-dUTP by nick translation (Invitrogen). A commercially available pericentromeric probe for chromosome 8 was used as reference (CEP 8Z2 SpectrumOrange, Vysis, Downers Grove, IL, USA). For dual-color FISH analysis, 4-μm sections of the breast cancer tissue microarray blocks were transferred to an adhesive-coated slide system (Instrumedics, Hackensack, NJ, USA). For proteolytic slide pretreatment, a commercial kit was used (Paraffin pretreatment reagent kit; Vysis). Before hybridization, tissue microarray sections were deparaffinized, air dried, and dehydrated in 70, 85, and 100% ethanol followed by denaturation for 5 min at 74 °C in 70% formamide-2 × SSC solution. After overnight hybridization at 37 °C in a humidified chamber, slides were washed and counterstained with 0.2 μmol/l 4′,6-diamidino-2-phenylindole in an antifade solution. Detection of the digoxigenin-labeled probe was conducted using fluorescent antibody enhancer set (Roche) containing an FITC-conjugated antibody. For each tissue spot, the predominant gene and centromere copy numbers in the tumor cell nuclei were estimated.
A tumor was considered amplified if the ratio of 8q21/centromere 8 was ≥2.0. Ratios of >1.0 and <2.0 were considered as gains and a ratio of ≤1.0 as normal.
Statistics
Pearson's chi-squared test and Student's t-test were used to study the relationship between 8q21 copy number and clinicopathological or molecular parameters. Survival effect of 8q21 and MYC amplification was assessed using Kaplan–Meier curves and log-rank tests. A Cox proportional-hazards model was used to identify independent factors associated with overall survival. Analysis was performed using R statistical software package for Windows (version 2.7.2, R Foundation for statistical computing).
Results
8q21 Amplification Frequency
A total of 1458 (66%) arrayed cancer samples were assessable using FISH (Table 1 and Figure 1). Copy number alterations of the 8q21 locus were found in 157 interpretable breast cancers, including amplification in 50 (3%) tumors and gains in 107 (7%) tumors according to our predefined criteria (Figure 2). Almost all amplified tumors showed clusters of <10 gene copies, but two cases with large clusters of >20 FISH signals were also found.
Association with Clinicopathological and Molecular Features
Amplifications of the 8q21 locus showed significant correlations with various histopathological and molecular features of breast carcinomas. 8q21 copy number changes were related to medullary phenotype (P=0.03) and high-grade tumors (P<0.0001, Table 1). In addition, 8q21 alterations were related to high Ki67 labeling index (P<0.05, Figure 3). Tumors with 8q21 gains or amplifications were characterized by an increased overall frequency of amplifications of other known oncogenes (P<0.0001, Figure 4), including HER2, CCND1, MDM2 (P<0.05 each), and MYC (P<0.0001, Table 2). The same trend was also found for EGFR amplification; however, the low prevalence of EGFR amplifications (1%) in our patient set did not allow for a statistically sound analysis. A comparison between the expected and the observed frequency of co-amplifications with at least one of the other genes revealed that tumors with 8q21 amplification had a twofold increased likelihood to develop other amplifications (expected probability 1.5%, observed probability 2.7%; P=0.0053). There was no association between aberrations of the 8q21 locus and tumor stage, presence of lymph node metastases, or hormone receptor status (P>0.05 each).
Association with MYC Amplification
Data on MYC amplification were available for the breast cancer tissue microarray from a previous study with a total number of 121 (5%) MYC-amplified tumors.8 A subset of 1132 tumors with data available for both 8q21 and 8q24 (MYC) were included in the current study. A combined analysis of MYC and 8q21 identified 90 tumors with amplifications of MYC and/or 8q21. Of these, 28 tumors were amplified for 8q21 only and 54 for MYC only. Co-amplification of both genes was found in the remaining eight tumors.
Prognostic Significance of 8q21Amplification
8q21 amplification was strongly associated with adverse prognosis in univariate survival analysis (Figure 5a). There was no effect of 8q21 gains on patient survival (P=0.48). Furthermore, we analyzed the overall patient survival in the subset of 1132 tumors with complete copy number data for 8q21 and MYC (Figure 5b). In this subgroup, no statistically relevant survival differences could be found between tumors with MYC amplification and tumors with 8q21 amplification, with or without included co-amplifications (P=0.255 and P=0.15, respectively). However, the adverse effect of 8q21 amplification was retained and a tendency to worse outcome in 8q21-amplified tumors was observed. 8q21/q24 co-amplification (n=8) was too rare for further statistically analysis. A multivariate analysis including the established prognostic markers of breast cancer (pT, pN, and BRE grade) and the 8q21 or 8q24 (MYC) amplification status did not reveal an independent prognostic value of either locus (Table 3).
Discussion
A recently published study using high-resolution array CGH on SK-BR-3 breast cancer cells reported three separate amplicons within 8q21.19 Studies on clinical breast cancer specimens suggested TPD52 and WWP1 as amplification target genes in this chromosomal region.20, 21 In concordance with the SK-BR-3 mapping study, TPD52 and WWP1 are located within the two distal 8q21 amplicons. The specific aim of this study was to gain more insight into the potential significance of the third (proximal) amplicon located between 70 and 80 Mb (8q21.11).
Amplification driver genes often map to central portions of an amplified region.24 A FISH probe mapping directly to the center of the proximal 8q21 amplicon identified in SK-BR-3 breast cancer cell line was therefore used for this study. The high number of analyzed breast cancer samples in combination with the extensive database collected during previous studies enabled a comprehensive comparison of the presence or absence of the proximal 8q21 copy number gains with multiple clinicopathological and molecular features, including survival data.8, 22 The associations with high grade, tumor cell proliferation, medullary phenotype, and poor clinical outcome argue for relevant biological role of at least one gene in the proximal 8q21 region. The comparatively weaker link between 8q21 gains and these parameters as well as the lack of prognostic significance underscores the biological effect of 8q21 amplification. Medullary and medullary-like cancers are well known for their high proliferative activity and expression of other relevant tumor proteins, such as EGFR and CD117.25, 26 This tumor entity comprises a heterogeneous subgroup of basal-like carcinomas, consisting of pure medullary carcinomas, atypical medullary carcinomas, and poorly differentiated ductal carcinomas with strong stromal inflammatory response.27 Accumulation of 8q21 amplifications in medullary-like breast carcinomas constitutes another argument in favor of a biological uniqueness of this rare subtype of breast cancer.
The relatively high number of 8q21 amplifications in our examination (3%) may be viewed as an indirect argument for our FISH probe mapping not so far from the target gene of the proximal 8q21 amplicon. The BAC for the used FISH probe contains entire transmembrane protein 70 (TMEM70), which is one interesting candidate target gene in the region. The gene encodes a small 30 kD protein located at the inner mitochondrial membrane. TMEM70 wild-type protein is necessary for regular biogenesis and assembly of the ATP synthase, as shown in some mitochondrial disorders with decreased activity of this protein.28, 29 Enhanced activity of ATP synthase results in elevated levels of reactive oxygen species (ROS) in the cell. High intracellular ROS was described in many cancer types, including breast carcinomas.30, 31, 32 It has been suggested that high ROS levels cause elevated expression of the transcription factor HIF-1α, which is also implicated in breast tumor development.33, 34 Stabilization of HIF-1α with increased aerobic glycolysis (Warburg effect) has a central role in many common human cancer types.35 Therefore, it might be that TMEM70 amplification with elevation of ROS led to a growth advantage of breast tumor cells.
Transcription elongation factor B, polypeptide 1 (TCEB1) is another interesting candidate oncogene. TCEB1 locates in the close proximity of the hybridized region, only 4-kb upstream from TMEM70. The gene encodes the protein elongin C, which serves as a cofactor for activation of transcriptional elongation by RNA polymerase II.36 TCEB1 was suggested to have oncogenic potential in prostate cancer.37 One study has described TCEB1 overexpression using quantitative RT-PCR in TCEB1-amplified SK-BR-3 cells.38
Although the co-amplification rate of 8q21 and MYC (13%) in our study was somewhat higher than the co-amplification rate of 8q21 and other analyzed amplicons (4–10%), amplifications of 8q21 and 8q24 occurred independently in most cases. MYC was not amplified in 28 of 36 8q21-amplified cancers, confirming the independent nature of the 8q21 amplicon. Several previous studies, including reports using the same tissue microarray as in this study, have shown a nonrandom accumulation of amplifications of different genomic regions in certain breast cancers that are considered to show an ‘amplifier’ phenotype.2, 8, 39, 40, 41 Amplification of the 8q21 locus is probably also part of a spectrum of breast carcinomas with high genomic instability and frequent amplifications.
TPD52 and WWP1, the most promising candidate target genes in the two other 8q21 amplicons in breast cancer, map 6 Mb and 12.5 Mb distal from our FISH probe. Amplification and overexpression of these genes were recently also found to be associated with short patient survival in breast cancer.42, 43 Although it has been hypothesized that each of these genes can cause a malignant phenotype in breast cancer by its own, it is also possible that a cumulative effect of multiple 8q21 genes contributes to the adverse prognosis in tumors with high genomic instability and many gene amplifications.
In summary, amplification of 8q21 occurs in a small but significant subgroup of genomic-instable breast carcinomas with poor prognosis. Copy number changes in 8q21 are independent of MYC and represent a separate amplicon in this chromosomal segment. Possible candidate oncogenes within this region include TCEB1 and TMEM70.
References
Tirkkonen M, Tanner M, Karhu R, et al. Molecular cytogenetics of primary breast cancer by CGH. Genes Chromosomes Cancer 1998;21:177–184.
Courjal F, Theillet C . Comparative genomic hybridization analysis of breast tumors with predetermined profiles of DNA amplification. Cancer Res 1997;57:4368–4377.
Rummukainen JK, Salminen T, Lundin J, et al., Amplification of c-myc oncogene by chromogenic and fluorescence in situ hybridization in archival breast cancer tissue array samples. Lab Invest 2001;81:1545–1551.
Buerger H, Otterbach F, Simon R, et al. Comparative genomic hybridization of ductal carcinoma in situ of the breast-evidence of multiple genetic pathways. J Pathol 1999;187:396–402.
Cingoz S, Altungoz O, Canda T, et al. DNA copy number changes detected by comparative genomic hybridization and their association with clinicopathologic parameters in breast tumors. Cancer Genet Cytogenet 2003;145:108–114.
Myllykangas S, Bohling T, Knuutila S . Specificity, selection and significance of gene amplifications in cancer. Semin Cancer Biol 2007;17:42–55.
Escot C, Theillet C, Lidereau R, et al. Genetic alteration of the c-myc protooncogene (MYC) in human primary breast carcinomas. Proc Natl Acad Sci USA 1986;83:4834–4838.
Al-Kuraya K, Schraml P, Torhorst J, et al. Prognostic relevance of gene amplifications and coamplifications in breast cancer. Cancer Res 2004;64:8534–8540.
Chen Y, Olopade OI . MYC in breast tumor progression. Expert Rev Anticancer Ther 2008;8:1689–1698.
Aulmann S, Adler N, Rom J, et al. c-myc amplifications in primary breast carcinomas and their local recurrences. J Clin Pathol 2006;59:424–428.
Rodriguez-Pinilla SM, Jones RL, Lambros MB, et al. MYC amplification in breast cancer: a chromogenic in situ hybridisation study. J Clin Pathol 2007;60:1017–1023.
Elbauomy Elsheikh S, Green AR, Lambros MB, et al. FGFR1 amplification in breast carcinomas: a chromogenic in situ hybridisation analysis. Breast Cancer Res 2007;9:R23.
Simon R, Richter J, Wagner U, et al. High-throughput tissue microarray analysis of 3p25 (RAF1) and 8p12 (FGFR1) copy number alterations in urinary bladder cancer. Cancer Res 2001;61:4514–4519.
Kallioniemi A, Kallioniemi OP, Piper J, et al. Detection and mapping of amplified DNA sequences in breast cancer by comparative genomic hybridization. Proc Natl Acad Sci USA 1994;91:2156–2160.
Kallioniemi A, Kallioniemi OP, Citro G, et al. Identification of gains and losses of DNA sequences in primary bladder cancer by comparative genomic hybridization. Genes Chromosomes Cancer 1995;12:213–219.
Cher ML, MacGrogan D, Bookstein R, et al. Comparative genomic hybridization, allelic imbalance, and fluorescence in situ hybridization on chromosome 8 in prostate cancer. Genes Chromosomes Cancer 1994;11:153–162.
Zhu H, Lam DC, Han KC, et al. High resolution analysis of genomic aberrations by metaphase and array comparative genomic hybridization identifies candidate tumour genes in lung cancer cell lines. Cancer Lett 2007;245:303–314.
Hicks J, Krasnitz A, Lakshmi B, et al. Novel patterns of genome rearrangement and their association with survival in breast cancer. Genome Res 2006;16:1465–1479.
Rodriguez V, Chen Y, Elkahloun A, et al. Chromosome 8 BAC array comparative genomic hybridization and expression analysis identify amplification and overexpression of TRMT12 in breast cancer. Genes Chromosomes Cancer 2007;46:694–707.
Balleine RL, Fejzo MS, Sathasivam P, et al., The hD52 (TPD52) gene is a candidate target gene for events resulting in increased 8q21 copy number in human breast carcinoma. Genes Chromosomes Cancer 2000;29:48–57.
Chen C, Zhou Z, Ross JS, et al., The amplified WWP1 gene is a potential molecular target in breast cancer. Int J Cancer 2007;121:80–87.
Ruiz C, Seibt S, Al Kuraya K, et al. Tissue microarrays for comparing molecular features with proliferation activity in breast cancer. Int J Cancer 2006;118:2190–2194.
Elston CW, Ellis IO . Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 1991;19:403–410.
Albertson DG, Ylstra B, Segraves R, et al. Quantitative mapping of amplicon structure by array CGH identifies CYP24 as a candidate oncogene. Nat Genet 2000;25:144–146.
Matkovic B, Juretic A, Separovic V, et al. Immunohistochemical analysis of ER, PR, HER-2, CK 5/6, p63 and EGFR antigen expression in medullary breast cancer. Tumori 2008;94:838–844.
Simon R, Panussis S, Maurer R, et al. KIT (CD117)-positive breast cancers are infrequent and lack KIT gene mutations. Clin Cancer Res 2004;10:178–183.
Rakha EA, Aleskandarany M, El-Sayed ME, et al. The prognostic significance of inflammation and medullary histological type in invasive carcinoma of the breast. Eur J Cancer 2009;45:1780–1787.
Cizkova A, Stranecky V, Mayr JA, et al. TMEM70 mutations cause isolated ATP synthase deficiency and neonatal mitochondrial encephalocardiomyopathy. Nat Genet 2008;40:1288–1290.
Houstek J, Kmoch S, Zeman J . TMEM70 protein - A novel ancillary factor of mammalian ATP synthase. Biochim Biophys Acta 2009;1787:529–532.
Szatrowski TP, Nathan CF . Production of large amounts of hydrogen peroxide by human tumor cells. Cancer Res 1991;51:794–798.
Santamaria G, Martinez-Diez M, Fabregat I, et al. Efficient execution of cell death in non-glycolytic cells requires the generation of ROS controlled by the activity of mitochondrial H+-ATP synthase. Carcinogenesis 2006;27:925–935.
Panayiotidis M . Reactive oxygen species (ROS) in multistage carcinogenesis. Cancer Lett 2008;266:3–5.
Chandel NS, Maltepe E, Goldwasser E, et al., Mitochondrial reactive oxygen species trigger hypoxia-induced transcription. Proc Natl Acad Sci USA 1998;95:11715–11720.
Chandel NS, McClintock DS, Feliciano CE, et al. Reactive oxygen species generated at mitochondrial complex III stabilize hypoxia-inducible factor-1alpha during hypoxia: a mechanism of O2 sensing. J Biol Chem 2000;275:25130–25138.
Denko NC . Hypoxia, HIF1 and glucose metabolism in the solid tumour. Nat Rev Cancer 2008;8:705–713.
Aso T, Lane WS, Conaway JW, et al., Elongin (SIII): a multisubunit regulator of elongation by RNA polymerase II. Science 1995;269:1439–1443.
Jalava SE, Porkka KP, Rauhala HE, et al., TCEB1 promotes invasion of prostate cancer cells. Int J Cancer 2009;124:95–102.
Porkka K, Saramaki O, Tanner M, et al. Amplification and overexpression of Elongin C gene discovered in prostate cancer by cDNA microarrays. Lab Invest 2002;82:629–637.
Courjal F, Cuny M, Simony-Lafontaine J, et al. Mapping of DNA amplifications at 15 chromosomal localizations in 1875 breast tumors: definition of phenotypic groups. Cancer Res 1997;57:4360–4367.
Melchor L, Alvarez S, Honrado E, et al. The accumulation of specific amplifications characterizes two different genomic pathways of evolution of familial breast tumors. Clin Cancer Res 2005;11:8577–8584.
Albertson DG . Gene amplification in cancer. Trends Genet 2006;22:447–455.
Shehata M, Bieche I, Boutros R, et al. Nonredundant functions for tumor protein D52-like proteins support specific targeting of TPD52. Clin Cancer Res 2008;14:5050–5060.
Nguyen Huu NS, Ryder WD, Zeps N, et al. Tumour-promoting activity of altered WWP1 expression in breast cancer and its utility as a prognostic indicator. J Pathol 2008;216:93–102.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Choschzick, M., Lassen, P., Lebeau, A. et al. Amplification of 8q21 in breast cancer is independent of MYC and associated with poor patient outcome. Mod Pathol 23, 603–610 (2010). https://doi.org/10.1038/modpathol.2010.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2010.5
Keywords
This article is cited by
-
6q deletion is frequent but unrelated to patient prognosis in breast cancer
Breast Cancer (2022)
-
Basal-like breast cancer: molecular profiles, clinical features and survival outcomes
BMC Medical Genomics (2017)
-
Development and validation of a novel clinical fluorescence in situ hybridization assay to detect JAK2 and PD-L1 amplification: a fluorescence in situ hybridization assay for JAK2 and PD-L1 amplification
Modern Pathology (2017)
-
New cytogenetically visible copy number variant in region 8q21.2
Molecular Cytogenetics (2011)
-
Clinical significance of high focal adhesion kinase gene copy number and overexpression in invasive breast cancer
Breast Cancer Research and Treatment (2011)