Abstract
The biological behavior of chronic lymphocytic leukemia and small lymphocytic lymphoma is unpredictable. Nonetheless, non-mutated IgVH gene rearrangement, ATM (11q22–23) and p53 (17p13) deletion are recognized as unfavorable prognosticators in chronic lymphocytic leukemia. The mRNA expression of activation-induced cytidine deaminase (AID), an enzyme indispensable for somatic hypermutation processes, was claimed to be predictive of non-mutated chronic lymphocytic leukemia cells in blood. Here, we evaluated AID protein expression compared with known molecular and immunohistochemical prognostic indicators in 71 chronic lymphocytic leukemia/small lymphocytic lymphoma patients using a tissue microarray approach. We found AID heterogeneously expressed in tumor cells as shown by colocalization analysis for CD5 and CD23. Ki-67 positive paraimmunoblasts of the proliferation centers displayed the highest expression. This observation is reflected by a significant association of AID positivity with a high proliferation rate (P=0.012). ATM deletion was detected in 10% (6/63) of patients and p53 deletion in 19% (13/67) of patients. Moreover, both ATM (P=0.002) and p53 deletion (P=0.004) were significantly associated with AID. IgVH gene mutation was seen in 45% (27/60) of patients. Twenty-five percent (17/69) of patients with AID-positive chronic lymphocytic leukemia/small lymphocytic lymphoma displayed a shorter survival than AID-negative chronic lymphocytic leukemia/small lymphocytic lymphoma patients (61 vs 130 months, P=0.001). Although there was a trend, we could not show an association with the IgVH gene mutation status. Taken together, our study shows that AID expression is an indicator of an unfavorable prognosis in chronic lymphocytic leukemia/small lymphocytic lymphoma patients, although it is not a surrogate marker for the IgVH status. Furthermore, the microenvironment of proliferation centers seems to influence AID regulation and might be an initiating factor in its transformation.
Similar content being viewed by others
Main
B-cell chronic lymphocytic leukemia with its tissue counterpart, small lymphocytic lymphoma, is the most frequent type of chronic leukemia encountered in the Western Hemisphere.1 The disease is characterized by a highly variable clinical course and affects mainly elderly adults, as only 10–15% of these patients are diagnosed before the age of 50 years.2 In recent years, several studies on peripheral blood and bone marrow cells from patients with chronic lymphocytic leukemia have revealed a strong and independent prognostic value of the IgVH mutation status and further chromosomal aberrations. In the following, four subgroups of chronic lymphocytic leukemia with differing survival probabilities have been defined on the basis of IgVH mutation status, 11q deletion (likely to involve the ATM kinase), and 17p deletion (involving p53).3, 4, 5 Moreover, patients presenting with non-mutated chronic lymphocytic leukemia with a high germline homology (>98%) more frequently show additional unfavorable genomic alterations such as 11q deletion.6, 7 The association of 17p deletion with non-mutated chronic lymphocytic leukemia varies from strong (Krober et al6) to borderline (Oscier et al7) significance. Del (13q) and trisomy 12 represent two of the most frequently observed chromosomal aberrations in chronic lymphocytic leukemia. Provided that del(13q) is the only aberration, there is a beneficial effect, whereas trisomy 12 appears to be disadvantageous (reviewed by Mehes8).3 These latter chromosomal alterations have not been correlated with the IgVH status.
In addition to their prognostic role, these chromosomal aberrations allow insight into the pathogenesis of chronic lymphocytic leukemia signified by alterations of candidate genes (11q22–23: ATM and 17p13: p53) and to the time point of hypothetical transformational events during germinal center (GC) reaction, as reflected by the IgVH mutation status. The protein tyrosine kinase ZAP70 has emerged as the most reliable surrogate marker for the IgVH mutational status, with respect to disease progression and survival in chronic lymphocytic leukemia when assessed by flow cytometry.9, 10
Activation-induced cytidine deaminase (AID) can convert cytosine into uracil on either mRNA or DNA level and is involved in DNA cleavage.11, 12, 13, 14 AID expression is essential for somatic hypermutation and class switch recombination of the immunoglobulin (Ig) gene (reviewed by Okazaki et al14). An initial report found AID expression to be localized to GC B cells.11 Recent studies on mRNA on fresh material revealed conflicting results concerning the correlation of AID expression and the IgVH mutation status.15, 16, 17, 18, 19 Despite emerging knowledge on molecular alterations influencing the biological behavior of chronic lymphocytic leukemia, it is still not clear from which normal B-cell counterpart malignant chronic lymphocytic leukemia cells evolve. Moreover, there are no data available on the role of molecular alterations in small lymphocytic lymphoma. Furthermore, it is not known to which extent the known prognostic surrogate markers from blood chronic lymphocytic leukemia cells can be transcribed to small lymphocytic lymphoma, the tissue counterpart of chronic lymphocytic leukemia. Therefore, the aim of this study was to analyze AID protein in the small lymphocytic lymphoma and tissue presentation of chronic lymphocytic leukemia to (1) determine the immuno-architecture and assess the frequency of AID expression, (2) clarify its prognostic role, (3) determine the relationship with chromosomal aberrations and (4) to investigate whether AID is a surrogate marker for the IgVH mutation status.
Materials and methods
Patients
Archival biopsies of 71 patients with chronic lymphocytic leukemia/small lymphocytic lymphoma diagnosed between 1993 and 2003 were retrieved from the files of the Institute of Surgical Pathology, University Hospital Zurich, Switzerland. Sixty-one were of nodal and 10 of extranodal localization. Sixty-three patients were classified for a chronic lymphocytic leukemia, whereas eight patients had localized nodal disease compatible with small lymphocytic lymphoma. To underline the tissue nature of chronic lymphocytic leukemia, we will talk only of small lymphocytic lymphoma from now on. For technical reasons (acid decalcificiation), no bone marrow trephines were considered for this study. Forty-five of the patients (63%) were male and 26 (37%) were female. The median age was 72 years. Four patients were younger than 50 years41, 42 at diagnosis. Survival data were available for all 71 patients. Their follow-up time ranged from 1 month to 153 months (median 46 months) during which period 43 patients (61%) died. All biopsies were reviewed by two hematopathologists (CG and MT) using CD20, CD3, CD5, CD23 and CyclinD1 as basic immunohistochemical panel. A TMA was constructed by acquiring two 0.6 mm punches of each of the formalin-fixed, paraffin-embedded tumor biopsies and four reactive tonsils as recently described.20 Only punches containing at least 30% of tumor cells were selected for further investigations.
The study was approved by the local Ethics Committee of the Canton of Zurich (StV 2–2007).
Immunohistochemistry/Immunofluorescence
Immunohistochemistry (IHC) on TMA sections was performed by an automated immunostainer (Ventana Medical System, Tucson, AZ, USA). An exact identification and analysis of the tumor cell population was performed with the antibodies described in Table 1. AID protein was investigated in parallel with two different monoclonal antibodies (clone EK2 5G9 and ZA001) of rat and mouse species (Table 1). To prevent a selection bias by using TMA, whole sections of 17 lymphomas were analyzed for AID in parallel. Double staining for AID with either CD5 or CD23 was carried out using fluorescence labeling of CD5 or CD23 with a goat anti-mouse antibody coupled to Alexa 488 (Invitrogen). For this purpose, AID (clone ZA001) was marked with a polymeric alkaline phosphatase-linker antibody conjugate (Bond Polymer AP Detection by Vision BioSystems, UK), excited by TRITC and analyzed on an Olympus BX61 microscope (Schwerzenbach, Switzerland) for conventional fluorescence imaging. The image capture was controlled with the AnalySISPro software (Soft Imaging System, Münster, Germany).
The staining intensity for all markers investigated was assessed in a four-tiered system and graded as negative (−), faint (1+), moderate (2+) and strong (3+). Furthermore, the percentage of positive tumor cells was assessed. The cutoff for AID immunoreactivity was set at 5%. The remaining IHC markers were defined as positive if at least 10% of tumor cells were stained. On the basis of a median proliferation rate of 20% (range MIB1 nuclear staining 5–50%), Ki-67 labeling index (MIB1-LI) was grouped into a low (<20% MIB1-positive nuclei) and high (≥20% MIB1-positive nuclei) proliferation rate.
Interphase FISH
For analysis of 11q22.3 (ATM), 17p31 (p53), 13q34 deletions and trisomy 12 commercially available multicolor FISH probe sets (17p13.1LSIp53/11q22.3LSIATM) and LSID13S319/LSI13q34/CEP 12 (Abbott Molecular, Wiesbaden, Germany) were applied. The protocol was according to the manufacturer's instructions, accustomed to the need of our tissue. Briefly, deparaffinization, pretreatment, digestion and washing steps were all performed in the VP 2000 Processor (Abbott Molecular). The digestion step with 0.5% protease was 15 min for tissue and 10 min for smears. Denaturation and hybridization overnight were performed in a hybridizer (HyBrite, Abbott Molecular) at 37°C. FISH signals of at least 100 nuclei of each tumor area were evaluated using a monochrome fluorescence F-View II camera (Soft Imaging System) installed on an Olympus BX61 microscope with analySISB software (Soft Imaging System).
The cutoff for deletions and for trisomy of chromosome 12 was set at 20% on the basis of preliminary examinations on reactive tonsils. The number of chromosomal alterations was calculated for each tumor and categorized into groups with a high (≥2) or low number (≤1).
IgVh Mutation Analysis
DNA was extracted either from fresh-frozen or from paraffin-embedded whole tissue, sections according to standard procedures. Specific PCR amplification was carried out using BIOMED-2, VH1–VH6 FRI-III and JH consensus primers as described.21 Monoclonal DNA fragments without additional bands on 3% MetaPhor agarose gels were directly purified from amplification reactions using a MinElute PCR Purification Kit (Qiagen, Hilden, Germany). Fragments from monoclonal populations with biallelic rearrangements were excised and subsequently purified with a QIAquick Gel Extraction Kit (Qiagen). Direct sequencing of the PCR products was performed on an ABI 3130xl Sequencer using the Big Dye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, Rotkreuz, Switzerland). The sequences of both strands were edited with the Sequencher Software v3.1.1 (Gene Codes) and compared with the IMGT immunoglobulin databank using V-QUEST (imgt.cines.fr). A 98% DNA sequence identity with germline sequences was defined as non-mutated small lymphocytic lymphomas as recommended by the European Research Initiative on chronic lymphocytic leukemia.22
Statistical Analysis
A complete set of morphological, immunohistochemical, molecular and survival information for univariate statistical analysis was obtained from a variable number of subjects, mainly because of the loss of TMA tissue cylinders. Characteristics of all variables and the number of analyzable cases are shown in Table 2. A statistical association between AID expression and other parameters was tested using a two-sided Fisher's exact test. Overall survival (OS) was estimated by the Kaplan–Meier method. OS was measured from the date of diagnosis and compared between small lymphocytic lymphoma patients with or without any of the clinical, immunohistochemical or molecular risk factors by log-rank statistics. P-values <0.05 were considered significant. Statistical analysis was performed using the software package SPSS (Version 12.0.1 for Windows, ©SPSS Inc., Chicago, USA).
Results
AID Protein Expression
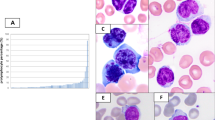
In reactive tonsils, AID was predominantly expressed in the dark zone of GC cells as previously reported by others.23, 24 A few scattered cells were seen in the interfollicular area (Figure 1a and 2c).25 Immunohistochemical AID expression in at least 5% of malignant cells was seen in 17 of 69 patients (25%). AID staining was found to be exclusively cytoplasmic (in small lymphocytes and paraimmunoblasts). Interestingly, AID expression pattern was rather heterogeneous regarding the percentage of positive cells as well as the staining intensity in an individual lymphoma. In general, only a small fraction of malignant cells (mean 7%, median 0%, range 0–60%) per specimen expressed AID (Figure 1b, c, 2a–b). Although the most intense staining was observed in proliferating (MIB1 positive) paraimmunoblasts and in cells of proliferation centers (PCs) (Figures 1d and 2b), a strong AID expression was not restricted to proliferating cells (Figure 1b–c). Staining patterns obtained on the TMA were confirmed using whole tissue sections in 11 patients with AID-positive and in 6 with AID-negative small lymphocytic lymphoma. In four patients, follow-up biopsies were available 3 months to 18 years after the initial investigation. In all these four patients with small lymphocytic lymphomas, AID immunoreactivity remained unchanged over time (negative in three cases, strongly positive in one case). The staining results were identical for both antibodies tested.
Activation-induced cytidine deaminase (AID) expression in the reactive germinal center and small lymphocytic lymphoma. (a) Predominant AID expression in the dark zone of a reactive germinal center, using the AID clone ZA001. (b) Small lymphocytic lymphoma on TMA with abundant AID-positive paraimmunoblasts. (c) Staining of the same case on whole tissue section showing a variable staining intensity in paraimmunoblasts, prolymphocytes and small lymphocytes. (d) Double staining for AID (cytoplasmic, brown) and Mib1 (nuclear, red) in a proliferation center showing proliferating and non-proliferating AID-positive cells.
Double staining for AID-CD23 and AID-CD5 in a small lymphocytic lymphoma and a reactive tonsil. (a) Double staining for AID-CD23 in a small lymphocytic lymphoma: (a) Overlay showing coexpression of AID (red) and CD23 (green); ai AID (red) and aii CD23 (green). (b) Double staining for AID-CD5 in a small lymphocytic lymphoma: Overlay showing variable AID (red) expression by CD5 (green) positive lymphoma cells, including para-immunoblasts of a proliferation center (upper right). (c) Double staining for AID-CD5 in a reactive tonsil as a control: AID (red) is expressed in germinal center cells and some activated parafollicular blasts (red), but not in CD5 (green)-positive T cells.
AID Expression and B-Cell Differentiation Markers
A high Ki-67 proliferation rate (≥20%) was significantly associated with positive AID expression in at least 5% of malignant cells (P=0.012). However, neither ZAP 70 nor the B-cell differentiation markers CD27 and Bcl-6 were associated with AID immunoreactivity (Table 2).
AID expression and chromosomal alterations
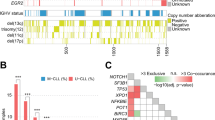
The number of chromosomal alterations differed in AID-positive and -negative small lymphocytic lymphomas, being higher in AID-positive small lymphocytic lymphomas (Figure 3a–b). AID-negative small lymphocytic lymphomas showed no or a low number (≤1) of chromosomal alterations in 40 of 46 (87%) cases. In this group, deletions of the 13q14 locus represent the prevailing aberration in 23 of 42 (55%) lymphomas, and were significantly associated with AID immunoreactivity (P=0.036). In contrast, AID-positive small lymphocytic lymphoma had a high number (≥2) of chromosomal alterations in 5 of 14 (36%) cases. ATM and p53 deletions were among the dominating aberrations, found in 12 of 14 (86%) cases in this group (Figure 3b). Deletion of the p53 gene was observed in 7 of 14 (50%) cases, and deletion of the ATM gene in 5 of 14 (36%) AID-positive small lymphocytic lymphomas, respectively; both alterations were statistically associated with AID expression as summarized in Table 2. Deletions of 13q34 and trisomy of chromosome 12 did not show any significant association with AID IHC.
Chromosomal aberration in AID-negative (a) and -positive (b) small lymphocytic lymphoma. (a) 13q deletion represents the most frequent aberration in AID-negative cases, whereas ATM and p53 genes are less frequently deleted. (b) The deletions of the tumor suppressor genes ATM and p53 show a mirror image, being predominantly present in these AID-positive lymphomas.
AID expression and mutation status
Activation-induced cytidine deaminase was expressed in a higher percentage of non-mutated small lymphocytic lymphoma with 10 of 33 (30%) compared with 3 of 27 (11%) mutated small lymphocytic lymphomas. However, owing to the low number of AID-positive cases, this trend did not reach statistical significance (P=0.115).
The mutation status could be assessed in 60 of 69 patients. In our cohort, 33 of 60 (55%) patients were non-mutated small lymphocytic lymphomas, and 27 of 60 (45%) were mutated small lymphocytic lymphomas. The VH3 family was the most frequently represented (n=28), followed by the VH4 and VH1 families with 13 cases each. VH1–69 was the most prevalent v-element detected (9/60, 15%), followed by VH3–23 (7/60, 12%). The IgVH families VH1–69 and VH3–23 were mainly found in non-mutated small lymphocytic lymphomas (8/9 and 6/7, respectively). In the mutated small lymphocytic lymphoma group one VH1–69 and two VH3–23 were observed. All rearrangements were in frame.
Survival analysis
The follow-up interval of the whole study population after diagnosis was 57 months (range 1–153 months). The estimated median survival time for patients with negative AID immunoreactivity was 130 months (95% confidence interval 96.6–163.4 months). For patients with AID-positive tumors (cutoff 5%), the estimated median survival time was 61 months (95% confidence interval 23.5–98.5 months) (Figure 4). The estimated 5- and 10-year survival rates were 76 and 34%, respectively. Only Bcl-6 protein expression in at least 10% of tumor cells (P=0.003) and deletion of the p53 gene using FISH (P=0.036) was associated with shorter OS. None of the other immunohistochemical or FISH markers correlated with survival.
The median survival time in patients with ATM deletion and p53 deletion was 74 and 50 months, contrasting with 153 months in patients with a 13q34 deletion only.
The median survival time of patients with non-mutated small lymphocytic lymphoma was shorter with 89 months (range 1–153 months) compared with mutated small lymphocytic lymphoma with 105 months (range 6–147 months). However, this trend was not statistically significant (P=0.25) (Table 2).
Discussion
By using a tissue-based approach studying the nodal manifestations of chronic lymphocytic leukemia or small lymphocytic lymphoma and by evaluating AID on a protein level by IHC, we extend the findings of previous groups investigating blood-derived leukemia cells. Hence, in this study, we show a significant association between AID protein expression and unfavorable clinical outcome in patients with nodal involvement of chronic lymphocytic leukemia or small lymphocytic lymphoma, respectively. On the basis of our findings, accelerated cell kinetics as well as p53 and ATM deletion might contribute to this adverse biological behavior as outlined below. For lack of clinical data, however, we are unable to indicate an influence on disease stage or treatment.
The prevalence of chromosomal aberrations of our study on formalin-fixed, paraffin-embedded tissue was comparable with data obtained on fresh blood cells.26, 27 In chronic lymphocytic leukemia, a significant association between early mortality and ATM deletion was reported. However, although we have seen a trend for such an association, we could not show significance.26, 28 This is very likely because of the low number of assessable cases. The strong association (P<0.05) of AID protein expression, deletion of both ATM and p53 and the adverse clinical outcome raises the question of an interaction of these genes. Indeed, AID-induced double-strand breaks can be resolved through the tumor suppressor pathways of ATM or p53 as shown in mouse models.29 Hence, one could speculate that AID overexpression promotes instability of the genome, which cannot be counter-regulated in the absence of these two tumor suppressor genes. Moreover, as the constitutive expression of AID triggers mutations of non-Ig genes, accumulation of further genetic alterations might be favored by prolonged AID expression.12, 14, 30 In this context, AID-induced p53 mutation using in vitro systems and the presence of hypermutation of the p53 gene associated with a high AID mRNA level in chronic lymphocytic leukemia patients is interesting.30, 31
Despite the association with these adverse biological markers and a high labeling index (MIB-LI >20%), a Cox regression model adjusted for OS did not reveal an independent and negative prognostic role of AID (data not shown).
We were able to define more precisely the immuno-architecture of AID in diseased tissue. AID protein was exclusively expressed in the cytoplasm, but not in the nucleus as would be expected from a protein with DNA mutator function, confirming previous results by western blot analysis on fractionated cells.32, 33 The staining heterogeneity concerns cell types (paraimmunoblasts and others) as well as staining intensity, the latter being strongest in paraimmunoblasts (Figure 5b–c). In particular, not all malignant cells stained positive for AID, not exceeding 60% of all tumor cells being the maximum. Using two different AID antibody clones (EK2 5G9 and ZA 001) showing an identical expression pattern, we could exclude technical reasons for this heterogeneity. Interestingly, our findings that only a subset of malignant B cells express AID corroborates results from mRNA analysis obtained from a dilution assay.15 Both these observations raise the question of whether these B cells are genetically different or inducible by their microenvironment. The fact that AID is particularly expressed in paraimmunoblasts, constituents of PCs, supports the hypothesis that PCs are sites of B-cell activation probably because of a particular microenvironment.15, 34 It favors the assumption that a specific cellular/chemical composition of the microenvironment is necessary to induce AID expression, leading to enhanced B-cell function and proliferation. Furthermore, the preferential localization of AID in PCs, particularly seen in lymph node manifestations of chronic lymphocytic leukemia, might explain why Pasqualucci et al33 did not detect the protein by western blotting in blood-derived chronic lymphocytic leukemia cells.
Although the process of somatic hypermutation and switch recombination requires AID, its unbalanced protein expression shown in mutated as well as non-mutated small lymphocytic lymphomas suggests that AID expression is, similar to normal GC B cells, very tightly regulated. It is tempting to speculate that this dissociation between AID expression and the absence/presence of somatic hypermutation in small lymphocytic lymphoma is because of one of the following mechanisms: (1) AID is expressed early in lymphomagenesis during the process of somatic hypermutation. Then, in the later stages of the disease, it might be shut off, requiring secondary genetic hits for lymphoma development, (2) AID is transiently and specifically induced in a particular microenvironment such as in B cells of PCs. Here, it might trigger disease progression and even transformation into aggressive lymphoma. Therefore, PC B cells might represent the initiator cells of prolymphocytic or Richter transformation as Reiniger et al35 indeed found an elevated AID mRNA level in such patients.
Here, we show that the IgVH gene mutation status stratifies small lymphocytic lymphoma patients into groups with different survival probability. Moreover, ZAP70 expression was correlated with non-mutated small lymphocytic lymphoma (P=0.028) (data not shown), further supporting our findings. Hence, the lack of statistical significance in survival difference between non-mutated and mutated small lymphocytic lymphoma might be explained either by the low number of cases investigated and/or the fact that patients with lymph node infiltration already have advanced disease at diagnosis. Our observation that AID is predominantly expressed in non-mutated small lymphocytic lymphoma is in line with previous studies on chronic lymphocytic leukemia cells.15, 16, 17, 18, 19 Identical to previous studies on peripheral blood chronic lymphocytic leukemia cells, the VH1 family member VH1–69 was also preferentially used in small lymphocytic lymphoma and associated with the non-mutated small lymphocytic lymphoma.36, 37 Similar to Mediterranean countries, we encountered the VH3–23 family much more frequently than VH3–21, the prevailing family in Scandinavian countries, which we observed in one single patient.38, 39
The constant presence of CD27 found in both mutated and non-mutated small lymphocytic lymphomas supports the view that small lymphocytic lymphoma cells derive from memory B cells.40, 41 Although the memory cell origin of mutated small lymphocytic lymphoma that has transited the GC is feasible, the same origin is difficult to understand for the non-mutated small lymphocytic lymphoma. There are two hypothetical explanations. Precursors of non-mutated small lymphocytic lymphoma may either be derived from B cells that have directly joined the pool of memory B cells circumventing the GC reaction, because of high antigen affinity, or by acquisition of the memory phenotype through a T-independent pathway.41, 42
Taken together, AID protein expression in small lymphocytic lymphoma correlates with an unfavorable clinical course and adverse biological parameters such as a higher proliferation rate and a complex genetic background. In contrast to normal B-cell differentiation, AID expression in small lymphocytic lymphoma is not directly linked to a hypermutated IgH status, as both mutated and non-mutated variable elements have been found in AID-expressing cases. Therefore, AID does not appear to be suitable as a surrogate marker for IgVH mutation status. As we were able to localize AID to prolymphocytes and cells of PCs, cells of this microenvironment are potential candidates for promoting AID expression.
References
Chiorazzi N, Rai KR, Ferrarini M . Chronic lymphocytic leukemia. N Engl J Med 2005;352:804–815.
Rozman C, Montserrat E . Chronic lymphocytic leukemia. N Engl J Med 1995;333:1052–1057.
Dohner H, Stilgenbauer S, Benner A, et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000;343:1910–1916.
Stilgenbauer S, Lichter P, Dohner H . Genetic features of B-cell chronic lymphocytic leukemia. Rev Clin Exp Hematol 2000;4:48–72.
Seiler T, Dohner H, Stilgenbauer S . Risk stratification in chronic lymphocytic leukemia. Semin Oncol 2006;33:186–194.
Krober A, Seiler T, Benner A, et al. V(H) mutation status, CD38 expression level, genomic aberrations, and survival in chronic lymphocytic leukemia. Blood 2002;100:1410–1416.
Oscier DG, Gardiner AC, Mould SJ, et al. Multivariate analysis of prognostic factors in CLL: clinical stage, IGVH gene mutational status, and loss or mutation of the p53 gene are independent prognostic factors. Blood 2002;100:1177–1184.
Mehes G . Chromosome abnormalities with prognostic impact in B-cell chronic lymphocytic leukemia. Pathol Oncol Res 2005;11:205–210.
Crespo M, Bosch F, Villamor N, et al. ZAP-70 expression as a surrogate for immunoglobulin-variable-region mutations in chronic lymphocytic leukemia. N Engl J Med 2003;348:1764–1775.
Rassenti LZ, Huynh L, Toy TL, et al. ZAP-70 compared with immunoglobulin heavy-chain gene mutation status as a predictor of disease progression in chronic lymphocytic leukemia. N Engl J Med 2004;351:893–901.
Muramatsu M, Sankaranand VS, Anant S, et al. Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. J Biol Chem 1999;274:18470–18476.
Okazaki IM, Hiai H, Kakazu N, et al. Constitutive expression of AID leads to tumorigenesis. J Exp Med 2003;197:1173–1181.
Ito S, Nagaoka H, Shinkura R, et al. Activation-induced cytidine deaminase shuttles between nucleus and cytoplasm like apolipoprotein B mRNA editing catalytic polypeptide 1. Proc Natl Acad Sci USA 2004;101:1975–1980.
Okazaki IM, Kotani A, Honjo T . Role of AID in tumorigenesis. Adv Immunol 2007;94:245–273.
Albesiano E, Messmer BT, Damle RN, et al. Activation-induced cytidine deaminase in chronic lymphocytic leukemia B cells: expression as multiple forms in a dynamic, variably sized fraction of the clone. Blood 2003;102:3333–3339.
Heintel D, Kroemer E, Kienle D, et al. High expression of activation-induced cytidine deaminase (AID) mRNA is associated with unmutated IGVH gene status and unfavourable cytogenetic aberrations in patients with chronic lymphocytic leukaemia. Leukemia 2004;18:756–762.
Kolar GR, Mehta D, Pelayo R, Capra JD . A novel human B cell subpopulation representing the initial population to express AID. Blood 2007;109:2545–2552.
McCarthy H, Wierda WG, Barron LL, et al. High expression of activation-induced cytidine deaminase (AID) and splice variants is a distinctive feature of poor-prognosis chronic lymphocytic leukemia. Blood 2003;101:4903–4908.
Oppezzo P, Vuillier F, Vasconcelos Y, et al. Chronic lymphocytic leukemia B cells expressing AID display dissociation between class switch recombination and somatic hypermutation. Blood 2003;101:4029–4032.
Ponzoni M, Berger F, Chassagne-Clement C, et al. Reactive perivascular T-cell infiltrate predicts survival in primary central nervous system B-cell lymphomas. Br J Haematol 2007;138:316–323.
van Dongen JJ, Langerak AW, Bruggemann M, et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 2003;17:2257–2317.
Ghia P, Stamatopoulos K, Belessi C, et al. ERIC recommendations on IGHV gene mutational status analysis in chronic lymphocytic leukemia. Leukemia 2007;21:1–3.
Greiner A, Tobollik S, Buettner M, et al. Differential expression of activation-induced cytidine deaminase (AID) in nodular lymphocyte-predominant and classical Hodgkin lymphoma. J Pathol 2005;205:541–547.
Smit LA, van Maldegem F, Langerak AW, et al. Antigen receptors and somatic hypermutation in B-cell chronic lymphocytic leukemia with Richter's transformation. Haematologica 2006;91:903–911.
Moldenhauer G, Popov SW, Wotschke B, et al. AID expression identifies interfollicular large B cells as putative precursors of mature B-cell malignancies. Blood 2006;107:2470–2473.
Sindelarova L, Michalova K, Zemanova Z, et al. Incidence of chromosomal anomalies detected with FISH and their clinical correlations in B-chronic lymphocytic leukemia. Cancer Genet Cytogenet 2005;160:27–34.
Glassman AB, Hayes KJ . The value of fluorescence in situ hybridization in the diagnosis and prognosis of chronic lymphocytic leukemia. Cancer Genet Cytogenet 2005;158:88–91.
Amiel A, Leopold L, Gronich N, et al. The influence of different chromosomal aberrations on molecular cytogenetic parameters in chronic lymphocytic leukemia. Cancer Genet Cytogenet 2006;167:145–149.
Ramiro AR, Jankovic M, Callen E, et al. Role of genomic instability and p53 in AID-induced c-myc-Igh translocations. Nature 2006;440:105–109.
Matsumoto Y, Marusawa H, Kinoshita K, et al. Helicobacter pylori infection triggers aberrant expression of activation-induced cytidine deaminase in gastric epithelium. Nat Med 2007;13:470–476.
Malcikova J, Smardova J, Pekova S, et al. Identification of somatic hypermutations in the TP53 gene in B-cell chronic lymphocytic leukemia. Mol Immunol 2008;45:1525–1529.
Begum NA, Kinoshita K, Muramatsu M, et al. De novo protein synthesis is required for activation-induced cytidine deaminase-dependent DNA cleavage in immunoglobulin class switch recombination. Proc Natl Acad Sci USA 2004;101:13003–13007.
Pasqualucci L, Guglielmino R, Houldsworth J, et al. Expression of the AID protein in normal and neoplastic B cells. Blood 2004;104:3318–3325.
Soma LA, Craig FE, Swerdlow SH . The proliferation center microenvironment and prognostic markers in chronic lymphocytic leukemia/small lymphocytic lymphoma. Hum Pathol 2006;37:152–159.
Reiniger L, Bodor C, Bognar A, et al. Richter's and prolymphocytic transformation of chronic lymphocytic leukemia are associated with high mRNA expression of activation-induced cytidine deaminase and aberrant somatic hypermutation. Leukemia 2006;20:1089–1095.
Schroeder Jr HW, Dighiero G . The pathogenesis of chronic lymphocytic leukemia: analysis of the antibody repertoire. Immunol Today 1994;15:288–294.
Johnson TA, Rassenti LZ, Kipps TJ . Ig VH1 genes expressed in B cell chronic lymphocytic leukemia exhibit distinctive molecular features. J Immunol 1997;158:235–246.
Donisi PM, Di Lorenzo N, Riccardi M, et al. Pattern and distribution of immunoglobulin VH gene usage in a cohort of B-CLL patients from a Northeastern region of Italy. Diagn Mol Pathol 2006;15:206–215.
Bomben R, Dal Bo M, Capello D, et al. Comprehensive characterization of IGHV3-21-expressing B-cell chronic lymphocytic leukemia: an Italian multicenter study. Blood 2007;109:2989–2998.
Klein U, Rajewsky K, Kuppers R . Human immunoglobulin (Ig)M+IgD+ peripheral blood B cells expressing the CD27 cell surface antigen carry somatically mutated variable region genes: CD27 as a general marker for somatically mutated (memory) B cells. J Exp Med 1998;188:1679–1689.
Klein U, Tu Y, Stolovitzky GA, et al. Gene expression profiling of B cell chronic lymphocytic leukemia reveals a homogeneous phenotype related to memory B cells. J Exp Med 2001;194:1625–1638.
Fais F, Ghiotto F, Hashimoto S, et al. Chronic lymphocytic leukemia B cells express restricted sets of mutated and unmutated antigen receptors. J Clin Invest 1998;102:1515–1525.
Acknowledgements
We are grateful to Silvia Behnke, Martina Storz and Helene Steinmann for excellent technical support, to Andreas Kofler for informatics support, and to Silvia Morf, Brigitte Steinauer and Danielle Schüler for registration. The work was supported in part by the Cancer League of the Canton of Zurich.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare no competing financial interest.
Rights and permissions
About this article
Cite this article
Leuenberger, M., Frigerio, S., Wild, P. et al. AID protein expression in chronic lymphocytic leukemia/small lymphocytic lymphoma is associated with poor prognosis and complex genetic alterations. Mod Pathol 23, 177–186 (2010). https://doi.org/10.1038/modpathol.2009.156
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2009.156
Keywords
This article is cited by
-
High expression of OX-40, ICOS, and low expression PD-L1 of follicular helper and follicular cytotoxic T cells in chronic lymphocytic leukemia
Journal of Hematopathology (2022)
-
Activation-induced deaminase and its splice variants associate with trisomy 12 in chronic lymphocytic leukemia
Annals of Hematology (2019)
-
Expression of activation-induced cytidine deaminase is associated with a poor prognosis of diffuse large B cell lymphoma patients treated with CHOP-based chemotherapy
Journal of Cancer Research and Clinical Oncology (2016)
-
Duodenal follicular lymphoma lacks AID but expresses BACH2 and has memory B-cell characteristics
Modern Pathology (2013)
-
Using Cytogenetic Rearrangements for Cancer Prognosis and Treatment (Pharmacogenetics)
Current Genetic Medicine Reports (2013)