Abstract
MicroRNAs are a group of small non-coding RNAs approximately 22 nucleotides in length. Recent work has shown differential expression of mature microRNAs in human cancers. We characterized the alteration in expression of a select group of microRNAs in primary peritoneal carcinoma relative to matched cases of ovarian serous carcinoma. MicroRNA expression was analysed using semi-quantitative stem-loop RT-PCR on a set of 34 formalin-fixed paraffin-embedded samples. Protein expression of p53 and bcl-2 was quantified in the corresponding tissue microarray. We provide definitive evidence that there is downregulation of a select group of microRNAs in tumours meeting Gynaecological Oncology Group criteria for primary peritoneal carcinoma relative to ovarian serous carcinoma. Specifically, we show decreased p53 expression and downregulation of miR-195 and miR-497 from the microRNA cluster site at chromosome 17p13.1 in primary peritoneal carcinoma relative to ovarian serous carcinoma. miR-195 and miR-497 may have potential roles as tumour-suppressor genes in primary peritoneal tumourigenesis.
Similar content being viewed by others
Main
Primary peritoneal carcinoma is an uncommon disease that occurs exclusively in women and diffusely involves both abdominal and pelvic peritoneum. Primary peritoneal carcinoma is similar to ovarian serous carcinoma histologically, however it is characterized by minimal surface ovarian involvement or minimal surface invasion of the ovarian cortex. The reported incidence of primary peritoneal carcinoma is 10–15% that of ovarian serous carcinoma.1, 2, 3 Although the recognition of primary peritoneal carcinoma is increasing its pathogenesis remains obscure. Chromosome 17 has been implicated as a potential location of genetic events important in the pathogenesis of both tumour types.4, 5, 6 High frequencies of loss of heterozygosity have been described at chromosomal band 17p13.1 (the genomic locus for p53 and microRNA cluster miR-195 and miR-497) in both tumour types.4 Primary peritoneal carcinoma exhibits distinct allelic loss patterns in comparison to ovarian serous carcinoma, implying that different sets of tumour-suppressor genes may be involved in the development of both tumour types.5
MicroRNAs are a class of small non-coding RNAs, approximately 22 nucleotides long that have been found to negatively regulate gene expression. They have been found to have roles in cell growth, differentiation, apoptosis and tumourigenesis.7, 8, 9, 10, 11, 12, 13, 14 Furthermore, unique microRNA expression profiles have been able to classify various cancers. Recently, microRNAs were implicated in the development of ovarian cancer: 39 microRNAs are differentially regulated between tumour and normal ovarian tissue.9
MicroRNA production involves cleavage of a long nascent transcript (primary microRNA) to form a 70–100 nucleotide hairpin precursor (precursor microRNA), which is in turn processed to form a microRNA duplex. One strand of this duplex is incorporated into the RISC complex where it will bind through partial sequence homology to the 3′ UTR of target mRNAs causing their translational repression.15, 16, 17 A variety of studies have linked aberrant microRNA expression to carcinogenesis where they can potentially act as both oncogenes and tumour-suppressor genes. Individual microRNAs such as miR-15 and miR-16 that target bcl-2 are frequently deleted and downregulated in CLL and thus can act as tumour-suppressor genes.7, 18
The purpose of this study was to examine the differential expression of a select group of microRNAs and their confirmed targets in primary peritoneal carcinoma relative to ovarian serous carcinoma. These microRNAs were chosen based on cluster site (17p13.1), their potential target pathways and on their relative fold changes as determined in a pilot study (unpublished observations, RJ Flavin). Furthermore, DNA copy number alterations at genomic loci for selected microRNAs have been described in ovarian serous carcinoma.19
Materials and methods
Ethical approval for the study was obtained from the St James's and Federation of Dublin Voluntary Hospitals ethics committee.
Case Selection and Tumour Sample Preparation
In total, 34 matched serous tumours (n=17 for primary peritoneal carcinoma and serous ovarian carcinoma) of advanced FIGO stage and grade were selected from archival formalin-fixed, paraffin-embedded tissue, between the years 1991 and 2006 from St James's Hospital, Dublin. All tumour samples were taken from the ovary. H&E slides of all tumours were reviewed by a histopathologist (RF) and original diagnoses confirmed. Primary peritoneal carcinoma was identified according to the Gynaecological Oncology Group criteria as previously described20 namely: (1) both ovaries were of normal size or enlarged by a benign process, (2)involvement of the extra-ovarian sites was greater than involvement on the surface of either ovary and (3) microscopically the ovarian component was either: non-existent, confined to the ovarian surface epithelium or involved the ovarian surface epithelium and underlying cortical stroma with tumour size not more than 5 × 5 mm.
Archival blocks were selected that contained over 90% tumour with contaminating stromal tissue estimated to be not more than 10%. Those tumours with >10% stromal contamination (n=10) were laser capture microdissected, using the PixCell 11™ System (Arcturus Engineering Inc., CA, USA), to yield homogenous populations of malignant epithelial cells lining the papillae, as previously described.21
RNA was extracted from archival material using RecoverAll™ Total Nucleic Acid Isolation Kit (Ambion Ltd, Cambridgeshire, UK) following the manufacturer's protocol. RNA quantity was assessed using a Nanodrop® ND-1000 Spectrophotometer (Wilmington, USA). Table 1 lists the clinicopathological characteristics of the cases selected. The paraffin tissue microarray composed of 68 serous tumours arrayed in quadruplicate (n=30 for primary peritoneal carcinoma; n=38 for ovarian serous carcinoma).
Stem-Loop RT-PCR
Applied Biosystems TaqMan® microRNA assays are designed to detect and quantify mature microRNAs using looped-primer real time PCR. All assays were taken from an early access panel, which contained 330 individual assays for identified human microRNAs (AB). Primer sequences are available on demand. Assays included: miR-16, miR-195 and miR-497. The protocol involves three steps: reverse transcription, pre-PCR amplification and real-time PCR, and has been previously described.23 Each RT reaction contained 10 ng of total RNA, and assays were gene dose corrected using let-7 as endogenous control.
Immunohistochemical Stains and Statistical Analysis
Sections (4 μm) of the ovarian serous carcinoma tissue array were cut and mounted on glass slides. For antigen unmasking, heat-mediated antigen retrieval was performed on deparaffinized sections using Triology (p53, bcl-2) in citrate buffer (10 mmol/L sodium citrate buffer (pH 6.0)) before incubation with primary antibodies. p53 (Vision Biosystems, Newcastle, UK) and bcl-2 (Dakocytomation, Copenhagen, Denmark) protein levels were examined using mouse monoclonal antibodies at dilutions of 1:50 and 1:30, respectively. Antibody staining was performed using the Ventana Nexus automated system (Tucson, Arizona, USA) using the avidin–biotin procedure.
Stained TMA slides were graded jointly by two pathologists (RF, CB). A modified visual semiquantification method was used as previously described,24 using a two-score system for immunointensity (II) and immunopositivity (IP). II and IP scores were summated. The semiquantification for II was scored on a scale of: 0, negative; 1, weak; 2, moderate and 3, strong. The semiquantification for percentage of IP cells was scored on a scale of 1 (1–10%), 2 (11–40%), 3 (41–70%) and 4 (>70%). This produced immunoscore values ranging from 0 to +7. Scores from all cores from one case were averaged. A cutoff value of ⩾5 was used to determine immunohistochemical positivity.
MicroRNA Target Prediction
The analysis of microRNA clusters and predicted targets were determined using the miRGen algorithm (http://www.diana.pcbi.upenn.edu/miRGen). Predicted targets of miR-195 and miR-497 were determined by union of target programs: DIANA-microT, miRanda, miRBase, PicTar and TargetScans.
Gene Ontology Analysis
Gene ontology analysis was performed using an online database known as the Panther Classification System (http://www.pantherdb.org). A binomial statistical tool was employed to compare the gene list to a reference list (ie the complete human genome) to determine over- or underrepresentation of PANTHER classification categories.
Statistical Analysis
Real-time stem-loop RT-PCR data analysis was performed using Real-Time StatMiner™ software from Integromics™ (www.integromics.com). Fold changes were calculated on filtered and ΔΔCT method normalized data using the CT method. P-values were calculated using a t-test.
The similarity in associations between tumour histotype and clinicopathological characteristics were examined by means of Fisher's exact test (Analyse-It™ Software Ltd) for discontinuous variables and a t-test for parametric continuous variables. The association between histotype and protein expression levels (p53 and bcl-2) were examined by means of a Mann–Whitney test.
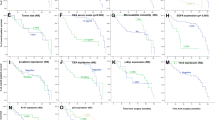
Kaplan–Meier estimates of survival time were compared using the logrank test (MedCalc™ Software Ltd). All tests were two-tailed, and the significance level was set at P<0.05.
Results
Real-Time Quantitative Stem-Loop RT-PCR Analysis of Differential MicroRNA in Primary Peritoneal Carcinoma Relative to Ovarian Serous Carcinoma
We used real-time stem-loop RT-PCR analysis to examine 17 matched cases for differential expression of miR-16, miR-195 and miR-497 in primary peritoneal carcinoma relative to ovarian serous carcinoma. All microRNAs were downregulated (Figure 1). miR-16 was downregulated by 4.42-fold (P=0.02) and miR-195 by 7.75-fold (P=0.006), whereas miR-497 was downregulated by 1149 fold (P=1.38E-11).
Immunohistochemical Analysis of p53 Expression in Primary Peritoneal Carcinoma Relative to Ovarian Serous Carcinoma
Our next investigation was to examine the protein expression levels of p53 and to see if there was any difference in relative expression between primary peritoneal carcinoma relative to ovarian serous carcinoma. This was undertaken as p53 is found at the same genomic locus on chromosomal band 17p13.1 as the microRNA cluster miR-195 and miR-497.
Background normal ovarian surface epithelium was predominantly non-reactive for p53. p53 overexpression was detected in 63% (19/30) of primary peritoneal carcinoma vs 84% (32/38) of ovarian serous carcinomas (Figure 2). In primary peritoneal carcinoma, the mean p53 immunoreactivity was 4.6 and in ovarian serous carcinoma, the mean p53 immunoreactivity was 6.0. The difference in p53 immunoreactivity between both tumour types was statistically significant (P=0.04). The results of the immunohistochemical analysis of p53 expression are summarized in Table 2.
Immunohistochemical Analysis of bcl-2 Expression in Primary Peritoneal Carcinoma Relative to Ovarian Serous Carcinoma
Our next investigation was to examine the protein expression levels of bcl-2 and to see if there was any difference in relative expression between primary peritoneal carcinoma and ovarian serous carcinoma. This was undertaken as bcl-2 is a validated target of miR-1625 and downregulation of this microRNA should be accompanied by an increased expression of its protein target.
Background normal ovarian surface epithelium was predominantly non-reactive for bcl-2. bcl-2 expression was detected in 20.69% (6/29) of primary peritoneal carcinoma vs 21.05% (8/38) of ovarian serous carcinomas (Figure 2). In primary peritoneal carcinoma, the mean bcl-2 immunoreactivity was 5.2 and in ovarian serous carcinoma, the mean bcl-2 immunoreactivity was 3.2. The difference in bcl-2 expression between both tumour types was statistically significant (P=0.02). The results of the immunohistochemical analysis of bcl-2 expression are summarized in Table 2.
Potential Targets of miR-195 and miR-497
Using the miRGen database, we looked for putative targets of miR-197 and miR-495 (no targets have been experimentally validated currently). Results were incorporated into PANTHER for gene ontology classification of the predicted mRNA targets. The cadherin, T-cell activation and WNT signalling pathways were significantly (P<0.01) overrepresented relative to a reference list. Potential targets include E-cadherin, α-catenin, cyclin D1 and cyclin D2. Table 3 lists the putative gene targets in significantly overrepresented pathways.
Discussion
In this study, we evaluated the expression of a select group of microRNAs and their target genes in primary peritoneal carcinoma relative to matched cases of ovarian serous carcinoma. Specifically, we show decreased p53 expression and downregulation of miR-195 and miR-497 from the microRNA cluster site at chromosome 17p13.1 in primary peritoneal carcinoma relative to ovarian serous carcinoma. Previous data have indicated that chromosome 17 may be a potential location of genetic events important in the pathogenesis of both tumour types.4, 5, 6 Throughout chromosome 17 frequent genetic deletions have been identified, suggesting that multiple tumour-suppressor genes may exist on this chromosome.26, 27, 28, 29 Our data therefore implicate miR-195 and miR-497 as potential tumour-suppressor genes.
Since its first description by Swerdlow in 1959, primary peritoneal carcinoma has been treated similarly to ovarian serous carcinoma by surgery and chemotherapy.30, 31 Despite histological and clinical similarities, some significant differences are seen in clinical, epidemiological and molecular characteristics of the two tumours, suggesting primary peritoneal carcinoma may be a distinct disease entity. Patients with primary peritoneal carcinoma tend to be older,32, 33 and when suboptimally debulked may have better survival rates than ovarian serous carcinoma patients.3, 34, 35 The prognosis of primary peritoneal carcinoma compared to ovarian serous carcinoma of the same stage is controversial with some studies reporting worse, same or better survival.32, 36, 37 A more recent matched case-comparison study demonstrated platinum resistance and impaired survival in patients with advanced primary peritoneal carcinomas.37 Of note, a recent epidemiological study found that parity and obesity are associated with an increased risk of peritoneal cancer, unlike ovarian serous cancer where there is an inverse correlation.38 The somewhat different patterns in terms of risk for primary peritoneal cancer suggests that primary peritoneal cancer may develop along different molecular pathways. Furthermore, differences in loss of heterozygosity,4, 5, 39 gene expression patterns40, 41 and cDNA copy number gains and loss have also been demonstrated between the two tumours possibly reflecting different carcinogeneic pathways.
Previous studies have suggested serous ovarian cancer may have a unifocal origin42, 43, 44, 45, 46 and that primary peritoneal cancer may have a multifocal origin.4, 40, 41, 47 Recent work proposes that epithelial ovarian cancer arises from three proposed origins, including the ovarian surface epithelium or mullerian inclusions, fallopian tube mucosa and mullerian epithelium elsewhere in the peritoneal cavity.2, 48, 49, 50, 51 Specifically, there is a growing body of evidence to suggest that both ovarian serous carcinoma and primary peritoneal carcinoma may arise from a precursor lesion in the distal tubal fimbria,49, 50, 51 and thus both tumour types may be genetically related. Importantly, our results do not preclude a common site of origin; they probably reflect temporal and spatial heterogenous microRNA expression in both tumour types, which maybe a consequence of biological properties such as preferential site of growth and spread.
Four recent studies reported aberrantly expressed microRNAs in human ovarian cancer.9, 14, 19, 52 Zhang et al19 described DNA copy number alterations at all genomic loci for our selected microRNAs in ovarian serous carcinoma. In the study by Iorio et al,9 miR-181a and miR-195 were downregulated in ovarian serous carcinoma, whereas Nam et al52 demonstrate differential expression of miR-16. Yang et al14 did not describe dysregulation of any of our selected microRNAs, however their group used microRNA microarrays (which are less sensitive and have a smaller dynamic range in comparison to PCR) to generate pilot data. Furthermore, a different RNA extraction protocol (Trizol Invitrogen) was used.
Interestingly, Iorio et al9 looked at differential expression of microRNAs in ovarian tumours with surface involvement and pelvic peritoneal involvement. miR-101, miR-182*, miR-22 and miR-133a were upregulated in those tumours with ovarian surface involvement, whereas miR-302c was upregulated in those tumours with pelvic peritoneal involvement. Though not stated in their study it would seem that ovarian surface and pelvic peritoneal involvement does not satisfy the Gynaecological Oncology Group diagnostic criteria for primary peritoneal carcinoma, rather reflecting pelvic and surface involvement by an ovarian serous carcinoma primary. Our data demonstrate decreased p53 expression and downregulation of a select group of microRNAs in tumours meeting Gynaecological Oncology Group criteria for primary peritoneal carcinoma relative to ovarian serous carcinoma. As the tissues studied were all taken from ovarian samples, it is possible that the differences observed are related to tumour progression (eg the peritoneal tumours began as p53 positive, miR-16, miR-195 and miR-497 retained tumours that then lost these features as they progressed and metastasized to the ovary. The ovarian tumours, as primary tumours, would have retained their p53 and microRNA profiles). Of note, dysregulation of miR-16, miR-195 and miR-497 have also been found in a range of other cancers, including chronic lymphocytic leukaemia, prostate, pancreas, cervical carcinomas together with sarcomas and benign uterine leiomyomata (Table 4).12, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63
Little is known about the mechanisms of microRNA deregulation in neoplastic tissue. MicroRNA genes are commonly located at minimal regions of amplification, loss of heterozygosity and breakpoint regions, suggesting that abnormal microRNA profiles can be caused by somatic gene mutation.19 Of note, previous studies have demonstrated high rates of loss of heterozygosity in both primary peritoneal carcinoma and ovarian serous carcinoma.4, 5, 39 A microRNA DNA copy number study of human ovarian cancer in combination with breast cancer and melanoma showed that a high proportion of genomic loci containing microRNA genes exhibit DNA copy number alterations.19 Furthermore, our group has demonstrated dysregulation of Dicer and Drosha proteins (both of which coordinate microRNA processing) in ovarian serous carcinoma.64
Recent research suggests that DNA methylation may also be involved in the regulation of microRNA expression in ovarian cancer, and the methylation can be reversed by DNA methyltransferase inhibitors.9, 65 Indeed, microRNA control and deregulation may be more complex as widespread microRNA repression by transcription factors such as c-myc (that targets miR-195/miR-497) have also been found to contribute to tumourigenesis.66
Our data demonstrate decreased p53 expression in primary peritoneal carcinoma relative to ovarian serous carcinoma. There was lower p53 overexpression in our cohort of primary peritoneal carcinomas vs ovarian serous carcinomas (63 vs 84%) with a lower mean p53 immunoreactivity score (4.6 vs 6.0; P=0.04). This difference may be due to distinct allelic loss patterns in both tumour types which has been demonstrated in previous studies.4, 5 The tumour-suppressor gene p53, which maps to chromosome 17p13.1, has a key role in cell-cycle regulation and apoptosis. Over 60% of ovarian carcinomas express mutations in the p53 gene.67, 68, 69 Previous studies70, 71, 72, 73 have reported both higher and lower p53 overexpression in ovarian serous carcinoma relative to primary peritoneal carcinomas, however these results were not statistically significant. True differences in p53 overexpression may however have been masked in these studies by both sample size and scoring methodology. The bcl-2 gene, an experimentally validated target of miR-16 in B-cell lymphomas,25 results in dysregulation of apoptosis. Expression of bcl-2 has been observed in 57% of ovarian cancer cases, and increased bcl-2 expression has been associated with improved patient survival.67, 69
Our study demonstrates both downregulation of miR-16 in primary peritoneal carcinoma relative to ovarian serous carcinoma and protein overexpression of bcl-2 in primary peritoneal carcinoma relative to ovarian serous carcinoma. In contrast to our study, previous groups have noted similarities in quantitative bcl-2 expression70 between primary peritoneal and ovarian serous carcinoma. However differences in qualitative expression were not examined specifically.
Potential overrepresented pathways targeted by mir-195 and miR-497 include the cadherin, WNT and T-cell activation pathways. Dysregulation of putative target genes within these lists have been described in ovarian cancer.74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88 E-cadherin and α-catenin, components of the cadherin pathway, have previously been found to be dysregulated in metatstatic ovarian carcinoma and serous effusions.79, 80, 81 Polymorphisms in cyclin D2 and protein kinase C isoform expression have been linked with prognosis in ovarian carcinoma patients.87, 88 AKT3, a component of the T-cell activation pathway, has been found to be highly expressed in a subset of ovarian serous carcinomas, and has been implicated as a key mediator of ovarian oncogenesis by the PI3K/AKT pathway.78 PIK3R1, part of the same family of lipid kinases has also been implicated as an oncogene in ovarian tumourigenesis.86 These putative targets indicate that miR-195 and miR-497 may have multiple biological roles in primary peritoneal carcinoma tumourigenesis.
In conclusion, we provide definitive evidence that there is downregulation of a select group of microRNAs in tumours meeting Gynaecological Oncology Group criteria for primary peritoneal carcinoma relative to ovarian serous carcinoma. Specifically, we show decreased p53 expression and downregulation of miR-195 and miR-497 from the microRNA cluster site at chromosome 17p13.1 in primary peritoneal carcinoma relative to ovarian serous carcinoma. Further investigation is needed to examine the functional role of miR-195 and miR-497 as potential tumour-suppressor genes in primary peritoneal tumour development.
References
Lele SB, Piver MS, Matharu J, et al. Peritoneal papillary carcinoma. Gynecol Oncol 1988;31:315–320.
Dalrymple JC, Bannatyne P, Russell P, et al. Extraovarian peritoneal serous papillary carcinoma. A clinicopathologic study of 31 cases. Cancer 1989;64:110–115.
Fromm GL, Gershenson DM, Silva EG . Papillary serous carcinoma of the peritoneum. Obstet Gynecol 1990;75:89–95.
Bandera CA, Muto MG, Welch WR, et al. Genetic imbalance on chromosome 17 in papillary serous carcinoma of the peritoneum. Oncogene 1998;16:3455–3459.
Huang LW, Garrett AP, Schorge JO, et al. Distinct allelic loss patterns in papillary serous carcinoma of the peritoneum. Am J Clin Pathol 2000;114:93–99.
Wojnarowicz PM, Breznan A, Arcand SL, et al. Construction of a chromosome 17 transcriptome in serous ovarian cancer identifies differentially expressed genes. Int J Gynecol Cancer 2007, Nov 19; e-pub ahead of print.
Calin GA, Dumitru CD, Shimizu M, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002;99:15524–15529.
Hayashita Y, Osada H, Tatematsu Y, et al. A polycistronic microRNA cluster, miR-17-92, is overexpressed in human lung cancers and enhances cell proliferation. Cancer Res 2005;65:9628–9632.
Iorio MV, Visone R, Di Leva G, et al. MicroRNA signatures in human ovarian cancer. Cancer Res 2007;67:8699–8707.
Karube Y, Tanaka H, Osada H, et al. Reduced expression of Dicer associated with poor prognosis in lung cancer patients. Cancer Sci 2005;96:111–115.
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature 2005;435:834–838.
Michael MZ, O’Connor SM, van Holst Pellekaan NG, et al. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res 2003;1:882–891.
Takamizawa J, Konishi H, Yanagisawa K, et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 2004;64:3753–3756.
Yang H, Kong W, He L, et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res 2008;68:425–433.
Ambros V . MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003;113:673–676.
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004;116:281–297.
Grosshans H, Slack FJ . Micro-RNAs: small is plentiful. J Cell Biol 2002;156:17–21.
Calin GA, Cimmino A, Fabbri M, et al. MiR-15a and miR-16-1 cluster functions in human leukemia. Proc Natl Acad Sci USA 2008;105:5166–5171.
Zhang L, Huang J, Yang N, et al. microRNAs exhibit high frequency genomic alterations in human cancer. Proc Natl Acad Sci USA 2006;103:9136–9141.
Bloss JD, Liao SY, Buller RE, et al. Extraovarian peritoneal serous papillary carcinoma: a case-control retrospective comparison to papillary adenocarcinoma of the ovary. Gynecol Oncol 1993;50:347–351.
Smyth P, Finn S, Cahill S, et al. ret/PTC and BRAF act as distinct molecular, time-dependant triggers in a sporadic Irish cohort of papillary thyroid carcinoma. Int J Surg Pathol 2005;13:1–8.
Silverberg SG . Histopathologic grading of ovarian carcinoma: a review and proposal. Int J Gynecol Pathol 2000;19:7–15.
Lao K, Xu NL, Yeung V, et al. Multiplexing RT-PCR for the detection of multiple miRNA species in small samples. Biochem Biophys Res Commun 2006;343:85–89.
Nisolle M, Gillerot S, Casanas-Roux F, et al. Immunohistochemical study of the proliferation index, oestrogen receptors and progesterone receptors A and B in leiomyomata and normal myometrium during the menstrual cycle and under gonadotrophin-releasing hormone agonist therapy. Hum Reprod 1999;14:2844–2850.
Cimmino A, Calin GA, Fabbri M, et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc Natl Acad Sci USA 2005;102:13944–13949.
Eccles DM, Brett L, Lessells A, et al. Overexpression of the p53 protein and allele loss at 17p13 in ovarian carcinoma. Br J Cancer 1992;65:40–44.
Eccles DM, Cranston G, Steel CM, et al. Allele losses on chromosome 17 in human epithelial ovarian carcinoma. Oncogene 1990;5:1599–1601.
Lee JH, Kavanagh JJ, Wildrick DM, et al. Frequent loss of heterozygosity on chromosomes 6q, 11, and 17 in human ovarian carcinomas. Cancer Res 1990;50:2724–2728.
Sato T, Saito H, Morita R, et al. Allelotype of human ovarian cancer. Cancer Res 1991;51:5118–5122.
Eltabbakh GH, Piver MS . Extraovarian primary peritoneal carcinoma. Oncology (Williston Park, NY) 1998;12:813–819; discussion 20, 25–26.
Swerdlow M . Mesothelioma of the pelvic peritoneum resembling papillary cystadenocarcinoma of the ovary; case report. Am J Obstet Gynecol 1959;77:197–200.
Bloss JD, Brady MF, Liao SY, et al. Extraovarian peritoneal serous papillary carcinoma: a phase II trial of cisplatin and cyclophosphamide with comparison to a cohort with papillary serous ovarian carcinoma—a Gynecologic Oncology Group Study. Gynecol Oncol 2003;89:148–154.
Eltabbakh GH, Piver MS, Natarajan N, et al. Epidemiologic differences between women with extraovarian primary peritoneal carcinoma and women with epithelial ovarian cancer. Obstet Gynecol 1998;91:254–259.
Barda G, Menczer J, Chetrit A, et al. Comparison between primary peritoneal and epithelial ovarian carcinoma: a population-based study. Am J Obstet Gynecol 2004;190:1039–1045.
Ben-Baruch G, Sivan E, Moran O, et al. Primary peritoneal serous papillary carcinoma: a study of 25 cases and comparison with stage III-IV ovarian papillary serous carcinoma. Gynecol Oncol 1996;60:393–396.
Dubernard G, Morice P, Rey A, et al. Prognosis of stage III or IV primary peritoneal serous papillary carcinoma. Eur J Surg Oncol 2004;30:976–981.
Eisenhauer EL, Sonoda Y, Levine DA, et al. Platinum resistance and impaired survival in patients with advanced primary peritoneal carcinoma: matched-case comparison with patients with epithelial ovarian carcinoma. Am J Obstet Gynecol 2008;198:213 e1-7.
Jordan SJ, Green AC, Whiteman DC, et al. Risk factors for benign serous and mucinous epithelial ovarian tumors. Obstet Gynecol 2007;109:647–654.
Cass I, Baldwin RL, Fasylova E, et al. Allelotype of papillary serous peritoneal carcinomas. Gynecol Oncol 2001;82:69–76.
Muto MG, Welch WR, Mok SC, et al. Evidence for a multifocal origin of papillary serous carcinoma of the peritoneum. Cancer Res 1995;55:490–492.
Schorge JO, Muto MG, Welch WR, et al. Molecular evidence for multifocal papillary serous carcinoma of the peritoneum in patients with germline BRCA1 mutations. J Natl Cancer Inst 1998;90:841–845.
Jacobs IJ, Kohler MF, Wiseman RW, et al. Clonal origin of epithelial ovarian carcinoma: analysis by loss of heterozygosity, p53 mutation, and X-chromosome inactivation. J Natl Cancer Inst 1992;84:1793–1798.
Li S, Han H, Resnik E, et al. Advanced ovarian carcinoma: molecular evidence of unifocal origin. Gynecol Oncol 1993;51:21–25.
Mok CH, Tsao SW, Knapp RC, et al. Unifocal origin of advanced human epithelial ovarian cancers. Cancer Res 1992;52:5119–5122.
Pejovic T, Heim S, Mandahl N, et al. Bilateral ovarian carcinoma: cytogenetic evidence of unicentric origin. Int J Cancer 1991;47:358–361.
Tsao SW, Mok CH, Knapp RC, et al. Molecular genetic evidence of a unifocal origin for human serous ovarian carcinomas. Gynecol Oncol 1993;48:5–10.
Karlan BY, Baldwin RL, Lopez-Luevanos E, et al. Peritoneal serous papillary carcinoma, a phenotypic variant of familial ovarian cancer: implications for ovarian cancer screening. Am J Obstet Gynecol 1999;180:917–928.
Feeley KM, Wells M . Precursor lesions of ovarian epithelial malignancy. Histopathology 2001;38:87–95.
Crum CP, Drapkin R, Kindelberger D, et al. Lessons from BRCA: the tubal fimbria emerges as an origin for pelvic serous cancer. Clin Med Res 2007;5:35–44.
Crum CP, Drapkin R, Miron A, et al. The distal fallopian tube: a new model for pelvic serous carcinogenesis. Curr Opin Obstet Gynecol 2007;19:3–9.
Kindelberger DW, Lee Y, Miron A, et al. Intraepithelial carcinoma of the fimbria and pelvic serous carcinoma: evidence for a causal relationship. Am J Surg Pathol 2007;31:161–169.
Nam EJ, Yoon H, Kim SW, et al. MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 2008;14:2690–2695.
Bloomston M, Frankel WL, Petrocca F, et al. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 2007;297:1901–1908.
Bruchova H, Yoon D, Agarwal AM, et al. Regulated expression of microRNAs in normal and polycythemia vera erythropoiesis. Exp Hematol 2007;35:1657–1667.
Calin GA, Ferracin M, Cimmino A, et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 2005;353:1793–1801.
Chen Y, Stallings RL . Differential patterns of microRNA expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 2007;67:976–983.
Iorio MV, Ferracin M, Liu CG, et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005;65:7065–7070.
Lui WO, Pourmand N, Patterson BK, et al. Patterns of known and novel small RNAs in human cervical cancer. Cancer Res 2007;67:6031–6043.
Murakami Y, Yasuda T, Saigo K, et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 2006;25:2537–2545.
Porkka KP, Pfeiffer MJ, Waltering KK, et al. MicroRNA expression profiling in prostate cancer. Cancer Res 2007;67:6130–6135.
Schetter AJ, Leung SY, Sohn JJ, et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 2008;299:425–436.
Subramanian S, Lui WO, Lee CH, et al. MicroRNA expression signature of human sarcomas. Oncogene 2008;27:2015–2026.
Wang T, Zhang X, Obijuru L, et al. A micro-RNA signature associated with race, tumor size, and target gene activity in human uterine leiomyomas. Genes Chromosomes Cancer 2007;46:336–347.
Flavin RJ, Smyth PC, Finn SP, et al. Altered eIF6 and Dicer expression is associated with clinicopathological features in ovarian serous carcinoma patients. Mod Pathol 2008;21:676–684.
Lu L, Katsaros D, de la Longrais IA, et al. Hypermethylation of let-7a-3 in epithelial ovarian cancer is associated with low insulin-like growth factor-II expression and favorable prognosis. Cancer Res 2007;67:10117–10122.
Chang TC, Yu D, Lee YS, et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 2008;40:43–50.
Casey G, Lopez ME, Ramos JC, et al. DNA sequence analysis of exons 2 through 11 and immunohistochemical staining are required to detect all known p53 alterations in human malignancies. Oncogene 1996;13:1971–1981.
Herod JJ, Eliopoulos AG, Warwick J, et al. The prognostic significance of Bcl-2 and p53 expression in ovarian carcinoma. Cancer Res 1996;56:2178–2184.
Teneriello MG, Ebina M, Linnoila RI, et al. p53 and Ki-ras gene mutations in epithelial ovarian neoplasms. Cancer Res 1993;53:3103–3108.
Chen LM, Yamada SD, Fu YS, et al. Molecular similarities between primary peritoneal and primary ovarian carcinomas. Int J Gynecol Cancer 2003;13:749–755.
Halperin R, Zehavi S, Hadas E, et al. Immunohistochemical comparison of primary peritoneal and primary ovarian serous papillary carcinoma. Int J Gynecol Pathol 2001;20:341–345.
Kowalski LD, Kanbour AI, Price FV, et al. A case-matched molecular comparison of extraovarian versus primary ovarian adenocarcinoma. Cancer 1997;79:1587–1594.
Soslow RA, Slomovitz BM, Saqi A, et al. Tumor suppressor gene, cell surface adhesion molecule, and multidrug resistance in mullerian serous carcinomas: clinical divergence without immunophenotypic differences. Gynecol Oncol 2000;79:430–437.
Aman P, Pejovic T, Wennborg A, et al. Mapping of the 19p13 breakpoint in an ovarian carcinoma between the INSR and TCF3 loci. Genes Chromosomes Cancer 1993;8:134–136.
Bourguignon LY, Gilad E, Peyrollier K . Heregulin-mediated ErbB2-ERK signaling activates hyaluronan synthases leading to CD44-dependent ovarian tumor cell growth and migration. J Biol Chem 2007;282:19426–19441.
Bourguignon LY, Peyrollier K, Gilad E, et al. Hyaluronan-CD44 interaction with neural Wiskott–Aldrich syndrome protein (N-WASP) promotes actin polymerization and ErbB2 activation leading to beta-catenin nuclear translocation, transcriptional up-regulation, and cell migration in ovarian tumor cells. J Biol Chem 2007;282:1265–1280.
Bourguignon LY, Zhu H, Zhou B, et al. Hyaluronan promotes CD44v3-Vav2 interaction with Grb2-p185(HER2) and induces Rac1 and Ras signaling during ovarian tumor cell migration and growth. J Biol Chem 2001;276:48679–48692.
Cristiano BE, Chan JC, Hannan KM, et al. A specific role for AKT3 in the genesis of ovarian cancer through modulation of G(2)-M phase transition. Cancer Res 2006;66:11718–11725.
Davidson B, Berner A, Nesland JM, et al. E-cadherin and alpha-, beta-, and gamma-catenin protein expression is up-regulated in ovarian carcinoma cells in serous effusions. J Pathol 2000;192:460–469.
Davies BR, Worsley SD, Ponder BA . Expression of E-cadherin, alpha-catenin and beta-catenin in normal ovarian surface epithelium and epithelial ovarian cancers. Histopathology 1998;32:69–80.
Fujimoto J, Ichigo S, Hirose R, et al. Expression of E-cadherin and alpha- and beta-catenin mRNAs in ovarian cancers. Cancer Lett 1997;115:207–212.
McPhillips F, Mullen P, MacLeod KG, et al. Raf-1 is the predominant Raf isoform that mediates growth factor-stimulated growth in ovarian cancer cells. Carcinogenesis 2006;27:729–739.
Melichar B, Nash MA, Lenzi R, et al. Expression of costimulatory molecules CD80 and CD86 and their receptors CD28, CTLA-4 on malignant ascites CD3+ tumour-infiltrating lymphocytes (TIL) from patients with ovarian and other types of peritoneal carcinomatosis. Clin Exp Immunol 2000;119:19–27.
Naora H, Yang YQ, Montz FJ, et al. A serologically identified tumor antigen encoded by a homeobox gene promotes growth of ovarian epithelial cells. Proc Natl Acad Sci USA 2001;98:4060–4065.
Pangas SA, Li X, Umans L, et al. Conditional deletion of Smad1 and Smad5 in somatic cells of male and female gonads leads to metastatic tumor development in mice. Mol Cell Biol 2008;28:248–257.
Philp AJ, Campbell IG, Leet C, et al. The phosphatidylinositol 3′-kinase p85alpha gene is an oncogene in human ovarian and colon tumors. Cancer Res 2001;61:7426–7429.
Song H, Hogdall E, Ramus SJ, et al. Effects of common germ-line genetic variation in cell cycle genes on ovarian cancer survival. Clin Cancer Res 2008;14:1090–1095.
Weichert W, Gekeler V, Denkert C, et al. Protein kinase C isoform expression in ovarian carcinoma correlates with indicators of poor prognosis. Int J Oncol 2003;23:633–639.
Acknowledgements
Dr Richard Flavin is funded by an HRB Ireland Clinical Research Fellowship under Grant no. CRT/2006/10. We acknowledge The Emer Casey Foundation, Ireland.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors have no conflict of interest to disclose.
Rights and permissions
About this article
Cite this article
Flavin, R., Smyth, P., Laios, A. et al. Potentially important microRNA cluster on chromosome 17p13.1 in primary peritoneal carcinoma. Mod Pathol 22, 197–205 (2009). https://doi.org/10.1038/modpathol.2008.135
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.135
Keywords
This article is cited by
-
Circulating microRNAs can predict chemotherapy-induced toxicities in patients being treated for primary breast cancer
Breast Cancer Research and Treatment (2023)
-
Low tissue levels of miR-125b predict malignancy in solitary fibrous tumors of the pleura
Respiratory Research (2017)
-
MicroRNA-195 acts as an anti-proliferative miRNA in human melanoma cells by targeting Prohibitin 1
BMC Cancer (2017)
-
Serum microRNA-195 is down-regulated in breast cancer: a potential marker for the diagnosis of breast cancer
Molecular Biology Reports (2014)
-
MicroRNA-497 increases apoptosis in MYCN amplified neuroblastoma cells by targeting the key cell cycle regulator WEE1
Molecular Cancer (2013)