Abstract
Tissue-resident memory (TRM) CD8+ T-cells are non-recirculating, long-lived cells housed in tissues that can confer protection against mucosal pathogens. Human immunodeficiency virus-1 (HIV-1) is a mucosal pathogen and the gastrointestinal tract is an important site of viral pathogenesis and transmission. Thus, CD8+ TRM cells may be an important effector subset for controlling HIV-1 in mucosal tissues. This study sought to determine the abundance, phenotype, and functionality of CD8+ TRM cells in the context of chronic HIV-1 infection. We found that the majority of rectosigmoid CD8+ T-cells were CD69+CD103+S1PR1− and T-betLowEomesoderminNeg, indicative of a tissue-residency phenotype similar to that described in murine models. HIV-1-specific CD8+ TRM responses appeared strongest in individuals naturally controlling HIV-1 infection. Two CD8+ TRM subsets, distinguished by CD103 expression intensity, were identified. CD103Low CD8+ TRM primarily displayed a transitional memory phenotype and contained HIV-1-specific cells and cells expressing high levels of Eomesodermin, whereas CD103High CD8+ TRM primarily displayed an effector memory phenotype and were EomesoderminNeg. These findings suggest a large fraction of CD8+ T-cells housed in the human rectosigmoid mucosa are tissue-resident and that TRM contribute to the anti-HIV-1 immune response. Further exploration of CD8+ TRM will inform development of anti-HIV-1 immune-based therapies and vaccines targeted to the mucosa.
Similar content being viewed by others
Introduction
The gastrointestinal mucosa is an important site of human immunodeficiency virus-1 (HIV-1) pathogenesis, as it serves both as a portal of entry and site of HIV-1 persistence throughout chronic infection.1 Accordingly, immune-based strategies to prevent and/or eradicate HIV-1 infection will likely require durable and robust HIV-1-specific immune responses in the rectosigmoid mucosa and other vulnerable tissues.2, 3, 4, 5 Characterized as long-lived, non-recirculating effector memory T-cells (TEFF) localized to tissues including the gastrointestinal tract, tissue-resident memory T-cells (TRM) represent a potential immunotherapeutic target for combating mucosal pathogens such as HIV-1 (refs 6, 7, 8, 9).
First described in the murine model, TRM are thought to develop from killer cell lectin-like receptor G1 (KLRG1)-negative precursor effector T-cells following migration into peripheral tissues.10 Here, exposure to tissue-specific cytokine mixtures involving transforming growth factor (TGF)-β drives expression of early activation marker CD69 and integrin αE(CD103)β7, which promote tissue accumulation and retention and are considered hallmarks of the TRM phenotype.6, 11, 12, 13, 14 Interestingly, although T-box transcription factors Eomesodermin (Eomes) and T-bet regulate CD8+ T-cell development and effector function, a feature of CD103+ CD8+ TRM conserved across models of infection is strong down-regulation of Eomes.12, 15 Recently, using a murine Herpes simplex virus-1 model, skin CD8+ TRM were shown to display low T-bet and negligible Eomes expression.16 Unlike circulating effector memory CD8+ T-cells, TRM in the gastrointestinal tract appear to be maintained independently of cognate antigen for long periods.7, 11, 17, 18 Positioned at sites of pathogen exposure, TRM initiate rapid and robust defenses upon reinfection, notably cytokine production, mobilizing both innate and adaptive arms of the immune system.10 Studies of the lung, skin, genital mucosa, and small intestine have all demonstrated the protective capacity of TRM against a range of pathogens after secondary infection and during reactivation of latent viral infection.17, 19, 20, 21, 22, 23, 24 Together, these data suggest TRM may be beneficial in controlling HIV-1 infection in peripheral, non-lymphoid tissues such as the gastrointestinal mucosa.
Although the protective attributes of TRM have been characterized extensively in murine models, a knowledge gap exists regarding their role in HIV-1 infection. Previous characterization of mucosal CD8+ T-cells in chronic HIV-1 infection revealed them to be phenotypically and functionally different from CD8+ T-cells circulating in blood. Rectosigmoid CD8+ T-cells displayed weak perforin-mediated cytotoxicity and diminished expression of T-bet and Eomes compared with their blood counterparts.25, 26, 27 Rather, rectosigmoid CD8+ T-cells were primarily T-betLowEomesoderminNeg and displayed robust cytokine/chemokine polyfunctionality characteristic of TRM. Strong polyfunctional HIV-1-specific CD8+ T-cell responses in rectosigmoid mucosa have been identified as a correlate of immune control, as they are particularly robust in individuals who naturally control HIV-1 (i.e., “controllers”).26 Whether these observations reflect an abundance of canonical ‘tissue-resident’ CD8+ T-cells in human gastrointestinal mucosa and participation of TRM in controlling HIV-1-infection is unknown.
The goal of this study was to determine whether gastrointestinal HIV-1-specific CD8+ T-cells in chronically HIV-1-infected and healthy participants share the characteristics of tissue resident T-cells as described in murine models of infectious disease, and to better understand the implications of this understudied population for HIV-1 pathogenesis, immune control, and vaccine design.
Results
Most CD8+ T-cells in human rectosigmoid mucosa display a tissue-resident phenotype
TRM can be distinguished from recirculating populations via expression of tissue-specific markers and the absence of recirculation markers.8, 19 Co-expression of CD69 and αE integrin (CD103) in the absence of sphingosine-1-phosphate receptor (S1PR1) and C–C motif chemokine receptor 7 (CCR7) are commonly used to identify tissue-resident CD8+ T-cells in a range of tissues.6, 13, 14, 28 To determine whether this strategy also identifies tissue-resident CD8+ T-cells in human rectosigmoid mucosa, we used flow cytometry to analyze cells for expression of CD103, CD69, and S1PR1 in chronically HIV-1-infected and seronegative (SN) participants.
Irrespective of HIV-1 disease status, large fractions of rectosigmoid CD8+ T-cells were CD103+CD69+S1PR1−; in contrast, this subset was negligible in blood (Figure 1a,b). In tissues, effector memory CD8+ T-cells often display a tissue-residency phenotype;6, 28, 29, 30, 31 however, the extent to which other memory subsets display such a phenotype is not known. We next analyzed expression of CD103, CD69, and S1PR1 within rectosigmoid CD8+ T-cell memory subsets in HIV-1+ and SN participants. Memory subsets were defined as follows: naïve (CD45RO−CD27+CCR7+), central memory (CD45RO+CD27+CCR7+), transitional memory (CD45RO+CD27+CCR7−), effector memory (CD45RO+CD27−CCR7−), and effector (CD45RO−CD27−CCR7−) (Supplementary Figure 1a,b online).32, 33 Although the abundance of naive (Tnaive) and central memory (TCM) CD8+ T-cells in mucosa was low compared with effector (TEFF) and TEM T-cells, all memory subsets contained some CD103+CD69+S1PR1− CD8+ T-cells, the proportion of which increased with memory differentiation. Only a small fraction of Tnaive and TCM CD8+ T-cells were CD103+CD69+S1PR1− (medians 2.16% and 19.5%, respectively); in contrast, large proportions of TEFF and TEM were CD103+CD69+S1PR1− (medians 64.7% and 74.9%, respectively) (Figure 1c,d). We also observed CD103 expressed at high and low median fluorescence intensities on rectosigmoid CD8+ T-cells. Although the majority of rectosigmoid tissue-resident CD8+ T-cells were CD103High, a small percentage was CD103Low; this percentage varied with memory differentiation. The proportion of CD103Low CD69+S1PR1− putative tissue-resident CD8+ T-cells was highest among TTM and TCM, and lowest among Tnaive (Figure 1c–e).
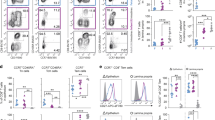
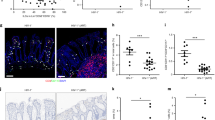
Characterization and distribution of tissue-resident CD8+ T-cells in the rectosigmoid mucosa of chronically human immunodeficiency virus-1 (HIV-1)-infected and seronegative (SN) participants. (a) Representative flow cytometry plot displaying surface staining of tissue-resident markers and gating strategy used to identify live, CD3+, tissue resident CD8+ T-cells as CD103+CD69+S1PR1− in blood (red) and rectosigmoid mucosa (blue). (b) Differences in the frequency of CD103+CD69+S1PR1− CD8+ T-cells between blood and rectosigmoid mucosa across HIV-1 disease status. (c) Representative flow cytometry plot demonstrating the difference in CD103 fluorescence intensity and CD69 expression in rectosigmoid naive (Tnaive, aqua), central memory (TCM, orange), transitional memory (TTM, green), effector memory (TEM, purple), and effector (TEFF, red) CD8+ T-cells. Numbers in parentheses indicate the percentage of the memory subset in the total rectosigmoid CD8+ T-cell population; colored numbers indicate frequency of the memory subset in each quadrant. Memory/effector subsets were first identified by differential expression of CD45RO, CD27, and CCR7 as described in the text. (d) Comparison of the frequencies of rectosigmoid CD103High and CD103Low CD69+S1PR1− CD8+ T-cells within each memory subset, or (e) between memory subsets in a combined group of chronically HIV-1-infected and SN participants. Horizontal bars represent median, whiskers indicate interquartile ranges with the level of significance shown with asterisks as follows: *P<0.05, **P<0.01, ****P<0.0001.
CD103 binds β7 to form the heterodimeric integrin αEβ7, which interacts with E-cadherin on epithelial cells, facilitating accumulation of tissue-resident T-cells in close proximity to the epithelium.34, 35, 36 We used immunohistochemistry and quantitative image analysis to confirm the distribution of CD103Neg, CD103Low, and CD103High CD8+ T cells. Cells were quantified in three anatomical compartments within rectosigmoid mucosa: intraepithelial (IE) lymphocytes, lamina propria (LP), and areas bordering the IE and LP (Supplementary Figure 2a). CD103Neg CD8+ T cells were located primarily in the LP, in contrast, greater than 60% of CD103High CD8+ T cells and approximately 50% of CD103Low CD8+ T cells were IE (Supplementary Figure 2a–c online). These findings corroborate previous observations concerning the localization of CD103-expressing CD8+ T-cells in human intestine34, 35 and reveal that a large proportion of rectosigmoid CD8+ T-cells resemble tissue-resident cells described previously. In human rectosigmoid mucosa, this population is composed mainly of CD103High effector and effector memory cells.
CD103High and CD103Low tissue-resident CD8+ T-cells differ in abundance across HIV-1 disease status, association with rectosigmoid CD4+ T-cells, and expression of memory markers
Disruption of the gut epithelial barrier and dramatic depletion of gut CD4+ T-cells are hallmarks of HIV-1 infection.5, 37 CD4+ T-cells have been shown to aid development of lung CD103High CD8+ TRM during influenza infection.10, 38 Accordingly, we hypothesized that the abundance of CD103High tissue-resident CD8+ T-cells might be lower in chronically HIV-1-infected compared with SN participants. To test this, we evaluated expression of tissue-resident markers on unstimulated rectosigmoid CD8+ T-cells in the following groups: HIV-1+ controllers; HIV-1+ viremic individuals not on antiretroviral therapy(ART) (viremic); HIV-1+ individuals on ART (Tx); and SN controls (Table 1). As CD8+ T-cells displaying a tissue-resident phenotype were primarily observed in effector memory and effector subsets, we investigated both CD45RO+ and CD45RONeg populations, referring to CD45RO+ CD103+CD69+S1PR1− cells as tissue-resident memory cells (TRM) and CD45RONeg CD103+CD69+S1PR1− cells as tissue-resident effector cells, abbreviated rTEFF.
As expected, HIV-1+ participants not on ART displayed lower proportions of CD103High CD8+ TRM and rTEFF compared with HIV-1+ participants on ART and SNs, reaching statistical significance for ART-treated compared with controllers and viremic untreated individuals for TRM, and between viremic untreated and SNs for rTEFF ( Figure 2 a,b ). No differences between subject groups were observed in the proportions of CD103Low TRM and rTEFF (data not shown). When all participant groups were combined, a strong positive correlation was observed between the percentage of rectosigmoid CD4+ T-cells and that of CD103High but not CD103Low TRM (Figure 2c). A similar correlation was seen when all HIV-1+ participants were considered together (P=0.0048, R2=0.44); however, when each subgroup (including SNs) was examined individually, no significant differences or trends were observed (not shown). No correlations were observed between rectosigmoid CD4+ T-cells and the percentages of CD103High or CD103Low rTEFF cells (Supplementary Figure 3a).
Distribution, association with rectosigmoid CD4+ T-cells, and maturation status of CD103High and CD103Low tissue-resident CD8+ T-cells. (a) Representative flow cytometry plot showing differences in the proportion of CD103High (blue) and CD103Low (orange) CD45RO+ and CD45RONeg CD8+ T-cells between a chronically human immunodeficiency virus-1 (HIV-1)-infected individual on antiretroviral therapy (ART) (top panel) and a chronically HIV-1-infected individual not on ART (bottom panel). (b) Differences in the frequency of rectosigmoid CD103High CD8+ tissue-resident memory T-cells (TRM) and rTEFF cells across HIV-1-disease status. (c) Association between the frequency of rectosigmoid CD4+ T-cells and the frequency of rectosigmoid CD103High and CD103Low TRM CD8+ T-cells in all study participants. (d) Representative flow cytometry plot showing differences in the expression of memory markers CD27 and CCR7 between CD103High and CD103Low rectosigmoid CD8+ TRM cells. (e) Distribution of central memory (TCM), transitional memory (TTM), and effector memory (TEM) subsets within CD103High and CD103Low rectosigmoid CD8+ TRM in a combined group of all study participants. Associations calculated using non-parametric Spearman’s correlation and linear regression analyses. Horizontal bars represent median, whiskers indicate interquartile range with the level of significance indicated with asterisks as follows: *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001.
CD103High and CD103Low tissue-resident CD8+ T-cells also differed in memory phenotype. The majority of CD103High TRM displayed an effector memory phenotype (TEM median 67.7%, TTM median 32.2%, and TCM median 0.225%); in contrast, CD103Low TRM cells mainly displayed a transitional memory phenotype characterized by expression of CD27 (TTM median 77.6%, TEM median 20.6%, and TCM median 1.05%) (Figure 2d,e). Among rTEFF, CD103High cells primarily displayed the canonical effector phenotype (CD45RO−CD27−CCR7−, median 73.5%), whereas a substantial fraction of CD103Low rTEFF expressed CD27 (CD45RO−CD27+CCR7− median 55.4%) (Supplementary Figure 3b,c). Collectively, these data reveal lower proportions of highly differentiated TEM/TEFF CD103High tissue-resident CD8+ T-cells in HIV-1+ individuals not on ART compared with ART-treated and SN groups, and a strong association between CD103High CD8+ TRM cells and rectosigmoid CD4+ T-cells.
HIV-1 gag-specific tissue-resident CD8+ T-cell responses originate from CD103Low cells and appear strongest in controllers
Although tissue-resident T-cells represent a potential immunotherapeutic target for HIV-1 eradication, little is known about the contribution these cells to the HIV-1-specific immune response during chronic infection. This and previous work have revealed the presence of HIV-1 Gag-specific, major histocompatibility complex (MHC) class I tetramer-binding CD8+ T-cells expressing αEβ7 in human rectosigmoid mucosa30 (Supplementary Figure 2d), and studies in rhesus macaques demonstrated vaccine-inducible long-lasting mucosal T-cell responses, suggesting TRM contribute to the HIV-1-specific immune response.18, 39 We investigated Gag-specific tissue-resident CD8+ T-cell responses and their contribution to the total rectosigmoid Gag-specific CD8+ T-cell response across HIV-1-disease status using an ex vivo 6 h stimulation assay followed by intracellular staining for effector molecules.
Degranulation (CD107a) and cytokine production in response to Gag-peptide stimulation were detected in rectosigmoid CD8+ TRM and rTEFF, with TRM displaying significantly higher percentages of Gag and Staphylococcal enterotoxin B (SEB)-responding cells compared with rTEFF (Figure 3a,b). Both controllers and viremic participants not on ART displayed significantly greater total and polyfunctional Gag-specific tissue-resident CD8+ T-cell responses compared with ART-treated participants (Figure 3c–e). This parallels earlier cross-sectional studies, in which we found low percentages of total and polyfunctional Gag-specific T-cells in mucosal tissues of participants on ART, particularly those with viral suppression.25, 26, 27 Although the total HIV-1 Gag-specific response did not differ significantly in magnitude between controllers and viremic untreated participants, the highest median polyfunctional responses (three- and four-function) were detected in controllers (Figure 3d,e). This was true for both TRM (Figure 3d) and rTEFF (Figure 3e). The strongest polyfunctional responses elicited by both TRM and rTEFF involved CD107a, MIP-1β, and interferon (IFN)-γ, with all notable polyfunctional responses involving CD107a (Figure3d,e). With the exception of interleukin-2-producing cells, Gag-responding CD8+ TRM were primarily CD103Low, reaching statistical significance for CD107a- and IFN-γ-producing TRM; in contrast, SEB-responding TRM were primarily CD103High (Supplementary Figure 4a,b). Similar trends were observed for Gag-responding CD8+ rTEFF (Supplementary Figure 4c).
Responses of rectosigmoid CD8+ tissue-resident memory T-cells (TRM) cells to antigenic stimulation. (a) Representative flow cytometry plot demonstrating degranulation (CD107a) and interferon (IFN)-γ-production following stimulation with an human immunodeficiency virus-1 (HIV)-1 Gag-peptide pool or medium containing dimethyl sulfoxide (DMSO) (negative control) by rectosigmoid CD8+ TRM (top) and rTEFF (bottom) from an HIV-1+ controller. (b) Difference in the frequency of CD8+ TRM and rTEFF cells responding to Gag-peptide and Staphylococcal enterotoxin B (SEB) stimulation. (c) Total percentage of rectosigmoid CD8+ TRM and rTEFF responding in any way (tumor necrosis factor (TNF)-α, IFN-γ, interleukin (IL)-2, CD107a, and/or macrophage inflammatory protein (MIP)-1β) to HIV-1 Gag stimulation. For each subject, the total HIV-1 Gag-specific response shown in (c) was calculated by combining all non-overlapping Boolean categories that are displayed individually in d for TRM and in e for rTEFF. (d) Difference in percentages of poly- and monofunctional Gag-specific rectosigmoid CD8+ TRM cells across HIV-1 disease status. (e) Difference in percentages of poly-and monofunctional Gag-specific rectosigmoid CD8+ rTEFF cells across HIV disease status. Response data were analyzed using Boolean gating of responding populations within live, CD3+, CD8+, CD45RO+ or CD45RONeg, CD103+CD69+S1PR1− gates. Co-expression analyses were generated using SPICE software. Poly- and monofunctional combinations with negligible responses were omitted. Horizontal bars represent medians, whiskers indicate interquartile ranges with the level of significance shown with asterisks as follows: *P<0.05, **P<0.01.
Previous work has demonstrated robust polyfunctional HIV-1-specific CD8+ T-cell responses in rectosigmoid mucosa of controllers;26 however, it is unknown whether these responses originate from resident and/or circulating populations. To understand the extent to which tissue-resident cells contribute to the total HIV-1 Gag-specific CD8+ T-cell response in rectosigmoid mucosa, we analyzed expression of CD69, CD103, and S1PR1 on Gag-specific polyfunctional CD8+ T-cells, focusing on the CD107a+MIP-1β+IFN-γ+ (7+M+F+) response as it had the highest median frequency (Supplementary Figure 5). Rectosigmoid Gag-specific 7+M+F+ CD8+ T-cells displayed three main patterns: the canonical tissue-resident phenotype (CD103+CD69+S1PR1−), CD69 single positive (CD103−CD69+S1PR1−), and the absence of all three markers (CD103−CD69−S1PR1−); the proportions differed between viremic individuals not on ART and controllers. In viremic individuals, rectosigmoid Gag-specific 7+M+F+ CD8+ T-cells were primarily tissue-resident (median 35.9%) or CD69 single positive (median 38.7%) with a smaller fraction lacking all three markers (median 11.6%). In controllers, the majority of 7+M+F+ CD8+ T-cells were CD69 single positive (median 52.2%) with much smaller proportions exhibiting tissue-residency (median 16.0%) or absence of all three markers (median 16.7%) (Supplementary Figure 5a–c). These trends were conserved across all polyfunctional responses (data not shown). The results presented here, in conjunction with previous work, demonstrate that HIV-1-specific tissue-resident CD8+ T-cells are present in rectosigmoid mucosa and capable of polyfunctional responses. In the context of chronic HIV-1 infection, Gag-specific tissue-resident CD8+ T-cells appear to be mainly CD103Low. Notably, although tissue-resident Gag-specific CD8+ T-cell responses appeared strongest in controllers, the tissue-resident population constituted a smaller proportion of the total rectosigmoid Gag-specific CD8+ T-cell response in controllers compared with viremic participants not on ART.
Eomes expression in tissue-resident CD8+ T-cells is associated with low CD103 fluorescence intensity and is greatest in HIV-1+ participants not on ART
Previous work demonstrated that although human rectosigmoid CD8+ T-cells typically had a T-betLowEomesoderminNeg phenotype characteristic of TRM, HIV-1+ individuals not on ART displayed greater Eomes expression compared with ART-treated and healthy participants.27 Whether this reflects upregulation of Eomes by tissue-resident CD8+ T-cells or an influx of EomesHigh CD8+ T-cells from circulation is unknown. To elucidate the origin of EomesHigh CD8+ T-cells in rectosigmoid mucosa and understand modulations in T-bet and Eomes expression during chronic HIV-1 infection, we used flow cytometry to detect intracellular T-bet and Eomes, in conjunction with tissue-resident markers, in unstimulated rectosigmoid CD8+ T-cells from HIV-1+ individuals and healthy controls (Figure 4 and Supplementary Figure 6).
Expression of T-bet and Eomesodermin (Eomes) in tissue-resident CD8+ T-cells varies with human immunodeficiency virus-1 (HIV-1)-disease status and CD103 fluorescence intensity. (a) Representative flow cytometry plot displaying differences in expression of T-bet and Eomes in rectosigmoid CD8+ tissue-resident memory T-cells (TRM) between a chronically HIV-1-infected participant not on antiretroviral therapy (ART) (far left), a chronically HIV-1-infected participant on ART (second from left), and a seronegative (SN) participant (second from right). T-bet fluorescence minus one (FMO) control is displayed for comparison (far right). (b) Differences in the frequency of EomesHigh and EomesNeg rectosigmoid CD8+ TRM across HIV-1-disease status. Horizontal bars represent medians, whiskers represent interquartile ranges. (c) Representative flow cytometry plot displaying differences in CD103 fluorescence intensity between EomesHigh and EomesNeg rectosigmoid CD8+ TRM. (d) Difference in Eomes expression between CD103High and CD103Low rectosigmoid CD8+ TRM cells in a combined group of all study participants. Vertical bars represent interquartile ranges, horizontal bars represent medians, whiskers represent s.e.m. Co-expression analysis was generated using SPICE software. Level of significance is indicated with asterisks as follows: *P<0.05, **P<0.01, ***P<0.001.
Rectosigmoid EomesHigh CD8+ T-cells were primarily single positive for CD69 (CD103−CD69+S1PR1−, median 47.4%); however, a notable fraction displayed the canonical tissue-resident phenotype (CD103+CD69+S1PR1−, median 26.6%) (Supplementary Figure 6b,c). To determine whether high Eomes expression observed in the CD103+CD69+S1PR1− subset was associated with chronic HIV-1 infection, we compared Eomes expression in CD8+ TRM and rTEFF cells across HIV-1 disease status (Figure 4a,b). Similar to the total rectosigmoid CD8+ T-cell population, tissue-resident CD8+ T-cells were primarily T-betLow/NegEomesNeg; however, HIV-1+ participants not on ART had higher percentages of EomesHigh tissue-resident CD8+ T-cells compared with ART-treated and SN participants. Unlike Eomes, no differences in T-bet expression were observed between groups (Figure 4). Within the TRM subset, viremic individuals not on ART displayed the highest median frequency of EomesHigh cells; however, within rTEFF, only HIV-1 controllers displayed significantly greater Eomes expression compared with ART-treated and SN groups (Figures 4a,b and Supplementary Figure 7a). The majority of EomesHigh tissue-resident CD8+ T-cells were CD103Low; in contrast, EomesNeg cells were primarily CD103High (Figure 4c,d and Supplementary Figure 7b). Together, these data demonstrate that rectosigmoid tissue-resident CD8+ T-cells are primarily CD103High T-betLow/NegEomesNeg; however, in untreated HIV-1 infection, a substantial population of CD103Low EomesHigh tissue-resident CD8+ T-cells emerges.
Discussion
Several studies have demonstrated the disparity in phenotype and function between CD8+ T-cells isolated from gastrointestinal mucosa and peripheral blood.25, 26, 27 As the gut is an important site of HIV-1 pathogenesis, these findings highlight the importance of understanding the distinct properties of mucosal immune responses to HIV-1.5 With the discovery of tissue-resident CD8+ T-cells, numerous studies have eloquently described the tissue-specific mechanisms that shape the tissue-resident population as well as the capacity of tissue-resident CD8+ T-cells to protect the host against mucosal pathogens.9, 10 Missing from this body of knowledge, however, is an understanding of the role of tissue-resident CD8+ T-cells in fighting HIV-1 infection in the gastrointestinal mucosa, a site of viral transmission and pathogenesis.5, 37 Thus, the aim of this study was to investigate the abundance of tissue-resident CD8+ T-cells in human rectosigmoid mucosa and the contribution of tissue-resident CD8+ T-cells to the HIV-1-specific immune response during chronic infection.
The majority of CD8+ T-cells in human rectosigmoid mucosa were CD103+CD69+S1PR1−, with a T-betLowEomesNeg transcriptional profile reminiscent of tissue-resident CD8+ T-cells described in the literature.10 This subset was negligible in blood, further evidence of the tissue-resident nature of these cells. As effector memory cells predominate in mucosal tissues and have an important role in secondary immune responses, most tissue-resident work has focused on memory T-cells.31 Indeed, the majority of rectosigmoid effector memory CD8+ T-cells were CD103+CD69+S1PR1−, expanding on previous work demonstrating an abundance of effector memory CD8+ T-cells expressing CD69 and CD103 in jejunum, ileum, and colon of human cadavers.28, 31 In addition, the majority of effector CD8+ T-cells in rectosigmoid mucosa also displayed tissue residency. Although the origin of this putative resident effector population, termed rTEFF, could not be definitively determined, one interpretation is that rTEFF are memory precursor effector cells transitioning into TRM. In murine skin and intestine, TRM develop from KRLG1Neg memory precursor effector cells.12, 36 In an oral Listeria monocytogenes murine model, intestinal IE KLRG1Neg memory precursor effector cells were CD103+CD69+ and formed rapidly compared with spleen or lung memory precursor effector cells in a TGF-β-dependent manner.36 Interestingly, we observed that although expression of KLRG1 was relatively low on rectosigmoid rTEFF and TRM compared with blood CD8+ T-cells, rTEFF displayed significantly lower expression of KLRG1 compared with TRM cells (these authors, unpublished data). Alternatively, rTEFF may represent a mature tissue-resident population distinct from TRM. Longitudinal studies designed to track the appearance and longevity of pathogen-specific rTEFF may help elucidate the origin and role of rTEFF in mucosal immunity.
Previous studies reported robust polyfunctional Gag-specific CD8+ T-cells in rectosigmoid mucosa of controllers, identifying these cells as a possible correlate of immune control.25, 26, 40 However, it remains unclear whether recent emigrants from blood or resident populations mediate these responses. We detected Gag-specific responses in both TRM and rTEFF populations, demonstrating that tissue-resident T-cells contribute to the HIV-1-specific immune response. Although this study did not have the statistical power to discern differences between controllers and viremic untreated individuals, the strongest median polyfunctional TRM and rTEFF responses were detected in controllers, suggesting tissue-resident CD8+ T-cells might have a role in controlling HIV-1 in rectosigmoid mucosa. Paradoxically, although controllers displayed the highest median percentages of polyfunctional Gag-specific tissue-resident CD8+ T-cell responses, these responses constituted a smaller proportion of the total rectosigmoid Gag-specific CD8+ T-cell response in controllers compared with viremic participants not on ART. Instead, in controllers, the majority of rectosigmoid Gag-specific polyfunctional CD8+ T-cells were single positive for CD69. CD69 is an early activation marker upregulated on tissue-resident precursor cells after entering the tissue. Expression of CD69 occurs early in tissue-resident development, preceding CD103 expression, and aids in tissue-retention by repressing expression of the tissue egress receptor S1PR1 (refs12, 13, 14, 41). Tissue-resident CD8+ T-cells have been shown to initiate immune responses, mediated through cytokine and chemokine secretion, leading to recruitment of circulating immune cells to the site of infection.22, 23 Accordingly, one interpretation of these data is that individuals with more robust Gag-specific tissue-resident CD8+ T-cell responses (that is, HIV-1 controllers) might in turn experience greater recruitment of HIV-1-specific CD8+ T-cells from circulation compared with individuals with weaker CD8+ tissue-resident responses. This enhanced recruitment of cells lacking the tissue-resident phenotype would dilute the proportion of Gag-specific cells in rectosigmoid mucosa with a tissue-resident phenotype, consistent with our observations. Clearly, further studies, likely involving non-human primate models, will be required to fully elucidate the role of tissue-resident CD8+ T-cells in the total mucosal anti-HIV-1 immune response.
Within both TRM and rTEFF populations, we observed two subsets of tissue-resident CD8+ T-cells that could be differentiated by CD103 fluorescence intensity. The CD103High subset resembled canonical tissue-resident CD8+ T-cells, consisting mainly of TEM/TEFF cells with a T-betLowEomesNeg profile. CD103High cells were enriched in epithelia and revealed a strong positive correlation with the percentage of rectosigmoid CD4+ T-cells. In contrast, the CD103Low subset was mainly composed of less-differentiated CD8+ T-cells expressing CD27 and was more abundant in LP compared with CD103High cells. HIV-1+ individuals not on ART had lower percentages of CD103High but higher percentages of EomesHigh tissue-resident CD8+ T-cells compared with ART-treated participants and SN controls. Interestingly, both Gag-specific and EomesHigh tissue-resident CD8+ T-cells were primarily CD103Low. Collectively, these observations reveal heterogeneity in the tissue-resident CD8+ T-cell population that may be exacerbated during chronic untreated HIV-1 infection.
Three main interpretations, not mutually exclusive, emerged to explain the observed HIV-1-associated alterations in the tissue-resident CD8+ T-cell population. First, gut epithelial cells express E-cadherin and are a constitutive source of TGF-β, a prominent driver of tissue-resident development.6, 7, 12, 16, 36, 42, 43 Certain subsets of CD4+ T-cells also produce TGF-β and CD4+ T-cells have been shown to aid development of TRM in the lung.38, 44 Accordingly, loss of epithelial barrier function and/or CD4+ T-cells during HIV-1 infection may result in a reduction of gut TGF-β and E-cadherin, ultimately leading to a reduction in an accumulation and/or development of CD103High tissue-resident CD8+ T-cells. Second, chronic antigen exposure can affect CD103 expression. In murine lymphocytic choriomeningitis virus infection, chronic in vivo exposure to antigen in combination with TGF-β reduced CD103 expression on antigen-specific CD8+ T-cells, leading the authors to propose CD103 as a marker for resting versus recently stimulated TRM.7 Consistent with this finding, we observed Gag-specific tissue-resident CD8+ T-cells to be primarily CD103Low, whereas tissue-resident CD8+ T-cells responding to SEB were primarily CD103High. It seems unlikely that the significant reduction in the percentage of CD103High tissue-resident CD8+ T-cells observed in HIV-1+ individuals not on ART would be due solely to CD103 downregulation on HIV-1-specific cells. However, the pool of rectosigmoid tissue-resident CD8+ T-cells undergoing activation-induced CD103 down-regulation might be broadened in individuals not on ART due to ‘bystander’ activation as a result of HIV-1-associated inflammation.37 Third, the HIV-1-associated inflammatory state in individuals not on ART might lead to greater recruitment of CD8+ T-cells from circulation. An influx of blood CD8+ T-cells into rectosigmoid mucosa would reduce the proportion of CD103High cells in the total rectosigmoid CD8+ T-cell population and increase proportions of CD103Low/Neg cells as recent immigrants mature into CD103High tissue-residents. In support of this interpretation, a large fraction of CD103Low tissue-resident CD8+ T-cells displayed a less differentiated memory phenotype characterized by CD27 expression.
Again, these interpretations are likely not mutually exclusive. The CD103Low tissue-resident CD8+ T-cell subset may be heterogeneous, composed both of recent immigrants developing into tissue-resident CD8+ T-cells and activated, mature tissue-resident CD8+ T-cells. Interestingly, within the CD103Low subset, we observed heterogeneity in Eomes expression. Unlike CD103High cells, which displayed a homogenous T-betLowEomesNeg phenotype, the CD103Low subset contained cells expressing high, low, or no Eomes. Notably, although CD103Low cells were primarily EomesNeg, EomesHigh tissue-resident CD8+ T-cells were almost exclusively CD103Low. Murine skin and brain tissue-resident CD8+ T-cells severely down-regulate Eomes; thus, detecting high Eomes expression within this population was surprising.12, 15, 16 However, high Eomes expression has been associated with a ‘terminally exhausted’ phenotype when co-expressed with the inhibitory receptor programmed cell death-1 (PD-1).45 As the frequency of EomesHigh tissue-resident CD8+ T-cells was greatest in HIV-1+ individuals not on ART, we surmised that the high Eomes expression in rectosigmoid tissue-resident CD8+ T-cells likely represents activated and/or exhausted cells. The data presented here do not contradict previous findings, but rather suggest that Eomes expression may vary during chronic infection and that tissue-resident CD8+ T-cells may be susceptible to exhaustion.
To our knowledge, this is the first report examining the characteristics and functionality of tissue-resident CD8+ T-cells in rectosigmoid mucosa during HIV-1 infection. As the gastrointestinal mucosa is an important site of HIV-1 transmission and pathogenesis,1 the long-lived tissue resident T-cell population can potentially be exploited in the development of vaccines and immunotherapies designed to prevent and/or eradicate HIV-1 infection. Recent successes in HIV-1 vaccine development have lent support to the concept that antigen-specific CD8+ T-cells, positioned near the sites of mucosal viral entry, can have critical roles in preventing and/or clearing infection.4, 18, 39 However, more work will be required to fully elucidate the pathways driving induction and maintenance of these populations and to understand the relative contributions of both tissue-resident and recirculating memory T-cell subsets to host immunity.
Methods
Participants and Sample Collection
HIV-1 positive and SN participants were enrolled through the SCOPE study at San Francisco General Hospital. Participants were categorized by plasma viral load and ART: Controllers with viral load consistently <2,000 copies per ml without ART; Viremic subjects with viral load ≥2,000 copies per ml without ART; ART-treated (Tx) participants with viral load<40 copies per ml; and HIV-1 SNs. SN participants were healthy volunteers from whom samples were collected for research purposes only. Written informed consent for phlebotomy and rectosigmoid biopsy was obtained through protocols approved by the Committee on Human Subjects Research, University of California, San Francisco (Protocols 10-01218, 10-00263, and 10-01330). Exclusion criteria included active sexually transmitted infections other than HIV-1, inflammatory bowel disease or other known inflammatory conditions affecting the gastrointestinal tract, colorectal cancer or other malignancies, and severe anemia or blood clotting disorders.
Twenty to 40 ml of blood was collected by sterile venipuncture into tubes containing EDTA. Twenty-four to 30 rectosigmoid biopsies were obtained by flexible sigmoidoscopy at 10–30 cm from the anal verge.2 Biopsies were immediately stored in R-15 (RPMI-1640 supplemented with fetal calf serum (15%), penicillin (100 U ml−1), streptomycin (100 mg ml−1), and glutamine (2 mM)). Samples were transported at room temperature to Davis, CA, and processed immediately upon arrival.
Blood and Tissue Processing
Peripheral blood mononuclear cells were isolated from blood using Ficoll-Paque (Pfizer-Pharmacia, New York, NY) and rested overnight in R-15 at 37 °C, 5% CO2. Rectosigmoid mononuclear cells were isolated from biopsies as described.46, 47 Briefly, biopsies were washed in R-15 and subjected to shaking incubation in 25 mg ml−1 Liberase DL (Roche, Indianapolis, IN) at 37 °C for 30 min, then passed through a 16-gauge blunt end needle to mechanically disrupt tissue. Disrupted tissue was passed through a sterile 70 μm cell strainer to collect cells. Undigested tissue was again incubated in Liberase and the process repeated until all tissue was digested. Isolated cells were washed three times in R-15 and rested overnight at 37 °C, 5% CO2 in R-15 with 200 × Zozyn (Pfizer-Pharmacia).
Antibodies and peptide pools
The following monoclonal antibodies (Abs) were used in flow cytometry: CD8 (SK1: APC-H7), CCR7 (3D12: PE-Cy7), CD69 (L78: PE), IFN-γ (B27: PE-Cy7), and MIP-1β (D21-1351: PerCP-Cy5.5) from BD Biosciences (San Jose, CA); CD103 (Ber-ACT8: Ax488), TNF-α (MAb11: BV605), IL-2 (MQ1-17H12: BV650), CD107a (H4A3: BV711), CD4 (RPA-T4: BV570, BV605), CD45RO (UCHL1: BV785), CD27 (O323: BV650), CD3 (UCHT-1: Ax700), and T-bet (4B10: BV711) from Biolegend (San Diego, CA). Unlabeled CD28 (L293) and CD49d (L25) were from BD Pharmingen (San Diego, CA). Eomes (WD1928: PE-eFluor610) and S1PR1 (SW4GYPP: eFluor660) were from ThermoFisher (Waltham, MA). The HIV-1 Gag peptide pool (p55, HXB2 sequence) consisted of 15–mers with an 11 amino acid overlap (BD Biosciences). SEB was from Sigma Aldrich (St. Louis, MO).
Antigen stimulation and intracellular cytokine staining
Surface and intracellular staining was performed as described25 on freshly isolated blood and rectosigmoid mononuclear cells rested overnight at 37 °C, 5% CO2. For stimulation assays, cells were incubated at 2 × 106 cells per 200 μl R-15 for 6 h in the presence of CD28 (1 μg ml−1) and CD49d (1 μg ml−1), anti-CD107a, 1 μM GolgiStop (BD Biosciences), brefeldin A (5 μg ml−1, Sigma Aldrich), and the appropriate antigens as follows: Gag-peptide pool (3.5 μg ml−1), SEB (5 μg ml−1), or media containing dimethyl sulfoxide (peptide solvent) as negative control. Cells were incubated for 10 min in phosphate-buffered saline with 0.5 mM EDTA before staining for surface markers and viability (Aqua Dead cell stain kit, Invitrogen, Waltham, MA) for 20 min at room temperature. Cells were fixed in 4% paraformaldehyde and permeabilized using FACS Perm 2 (BD Biosciences) before intracellular staining for CD3, IFN-γ, MIP-1β, TNF-α, and IL-2. The Transcription Factor Buffer Set (BD Biosciences) was substituted for 4% paraformaldehyde and FACS Perm 2 when staining for transcription factors. In such cases, cells were fixed with Transcription Factor Fix/Perm Buffer for 40 min at 4 °C, washed twice in Transcription Factor Perm/Wash Buffer, and stained with Abs for 40 min at 4 °C in Transcription Factor Perm/Wash. For all protocols, after staining, cells were re-suspended in 1% paraformaldehyde and stored at 4 °C in the dark until analysis within 24 h.
Data acquisition and analysis
Flow cytometry data were acquired with an LSR II cytometer and FACSDiva software (Becton Dickinson Immuno-cytometry Systems, San Jose, CA). Analysis was performed using Flowjo software (Flowjo LLC, Ashland, OR). Fluorescence minus one samples and known negative populations were used to aid in gating (Supplementary Figures 1a, 6b and 8). The distinction between CD103 negative, low, and high populations was made visually as shown in Supplementary Figure 8. Tissue-resident responses were identified using Boolean gating within the live, CD3+, CD8+, CD45RO+, and CD103+CD69+S1PR1− subset for TRM or live, CD3+, CD8+, CD45RO−, and CD103+CD69+S1PR1− subset for rTEFF. An in-house statistical algorithm was used to determine whether antigen-specific responses differed significantly from the negative control.25 For responses deemed statistically significant, net responses were then calculated by subtracting negative control values from antigen-specific responses. SPICE software was used to visualize multifunctional T-cell populations.48 Statistical analyses were performed with GraphPad Prism V.6 (Graphpad Software, San Diego, CA) or SPICE. Non-parametric statistical tests included Wilcoxon matched-pairs signed rank test for paired samples, Mann–Whitney test for unpaired samples, Spearman’s correlation, and linear regression analysis.
In situ tetramer staining and immunohistochemistry
Fresh biopsy specimens were shipped in complete media on ice overnight to the University of Minnesota. Biopsies were cut into two to four sections using a scalpel and stained with MHC-tetramers, CD103, and CD8 Abs as previously described.49, 50 Red background from MHC tetramer staining was used for morphological identification of intestinal crypts. Biotinylated MHC-class I monomers were loaded with peptides (National Institute of Health Tetramer Core Facility, Emory University, Atlanta, GA) and converted to MHC-class I tetramers. HLA-B*57 molecules were loaded with HIV-1 Gag IW9 (ISPRTLNAW) or QW9 (QASQEVKNW) peptides. HLA-B*07 molecules were loaded with Nef TL10 (TPNGSIFTTL) peptides. An irrelevant peptide MART (ELAGIGILTV) from melanoma protein was used as negative control. Fresh tissue sections were incubated with MHC-class I tetramers (0.5 μg ml−1), mouse-anti-human CD103 Abs (2.5 μg ml−1, Biolegend), and rat-anti-human CD8 Abs (2 μg ml−1, Acris, San Diego, CA). For secondary incubations, sections were incubated with rabbit-anti-fluorescein isothiocyanate Abs (0.5 μg ml−1, BioDesign, Saco, ME). For tertiary incubations, sections were incubated with Cy3-conjugated goat-anti-rabbit Abs (0.3 μg ml−1, Jackson ImmunoResearch Laboratories, West Grove, PA), Alexa 488-conjugated goat-anti-mouse Abs (0.75 μg ml−1, Molecular Probes, Eugene, OR), and Cy5-conjugated goat-anti-rat Abs (0.3 μg ml−1, Jackson ImmunoResearch Laboratories). Sections stained with secondary Abs alone were used as negative controls for CD103 and CD8 staining. Sections were imaged using an Olympus FluoView 1000 microscope (Olympus America, Center Valley, PA). Confocal z-series were collected from ∼3 μm from the surface of the section to 30–45 μm into the tissue. Montage images of multiple 800 × 800 pixels were created and used for analysis.
Quantitative Image Analysis
Anatomical compartments of rectosigmoid mucosa, crypts and IE lymphocytes were identified morphologically by increasing the red background from MHC tetramers to allow identification of goblet cells. The ∼10 μm area directly adjacent was defined as the border. Surrounding areas with abundant CD8+ T cells and diffuse red background were identified as LP. Areas that were ambiguous in structure were not included. To determine levels of CD103 expression, CD8+ T cells with no detectable CD103 staining above background were scored as CD103 negative, cells with CD103 staining 2–5 × greater than background were scored as CD103low, cells with CD103 staining 6 × or greater than background were scored as CD103high.
References
Hatano, H. et al. Comparison of HIV DNA and RNA in gut-associated lymphoid tissue of HIV-infected controllers and noncontrollers. AIDS 27, 2255–2260 (2013).
Shacklett, B.L. & Ferre, A.L. Mucosal immunity in HIV controllers: the right place at the right time. Curr. Opin. HIV AIDS 6, 202–207 (2011).
Barouch, D.H. & Deeks, S.G. Immunologic strategies for HIV-1 remission and eradication. Science 345, 169–174 (2014).
Masopust, D. & Picker, L.J. Hidden memories: frontline memory T cells and early pathogen interception. J. Immunol. 188, 5811–5817 (2012).
Shacklett, B.L. & Anton, P.A. HIV infection and gut mucosal immune function: updates on pathogenesis with implications for management and intervention. Curr. Infect. Dis. Rep. 12, 19–27 (2010).
Masopust, D., Vezys, V., Wherry, E.J., Barber, D.L. & Ahmed, R. Cutting edge: gut microenvironment promotes differentiation of a unique memory CD8 T cell population. J. Immunol. 176, 2079–2083 (2006).
Casey, K.A. et al. Antigen-independent differentiation and maintenance of effector-like resident memory T cells in tissues. J. Immunol. 188, 4866–4875 (2012).
Schenkel, J.M. & Masopust, D. Tissue-resident memory T cells. Immunity 41, 886–897 (2014).
Thome, J.J. & Farber, D.L. Emerging concepts in tissue-resident T cells: lessons from humans. Trends Immunol. 36, 428–435 (2015).
Mueller, S.N. & Mackay, L.K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2016).
Hofmann, M. & Pircher, H. E-cadherin promotes accumulation of a unique memory CD8 T-cell population in murine salivary glands. Proc. Natl Acad. Sci. USA 108, 16741–16746 (2011).
Mackay, L.K. et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Skon, C.N. et al. Transcriptional downregulation of S1pr1 is required for the establishment of resident memory CD8+ T cells. Nat. Immunol. 14, 1285–1293 (2013).
Mackay, L.K. et al. Cutting edge: CD69 interference with sphingosine-1-phosphate receptor function regulates peripheral T cell retention. J. Immunol. 194, 2059–2063 (2015).
Wakim, L.M. et al. The molecular signature of tissue resident memory CD8 T cells isolated from the brain. J. Immunol. 189, 3462–3471 (2012).
Mackay, L.K. et al. T-box transcription factors combine with the cytokines TGF-beta and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Mackay, L.K. et al. Long-lived epithelial immunity by tissue-resident memory T (TRM) cells in the absence of persisting local antigen presentation. Proc. Natl Acad. Sci. USA 109, 7037–7042 (2012).
Li, H. et al. Durable mucosal simian immunodeficiency virus-specific effector memory T lymphocyte responses elicited by recombinant adenovirus vectors in rhesus monkeys. J. Virol. 85, 11007–11015 (2011).
Carbone, FR Tissue-resident memory T cells and fixed immune surveillance in nonlymphoid organs. J. Immunol. 195, 17–22 (2015).
Zhu, J. et al. Immune surveillance by CD8alphaalpha+ skin-resident T cells in human herpes virus infection. Nature 497, 494–497 (2013).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Schenkel, J.M. et al. T cell memory. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014).
Schenkel, J.M., Fraser, K.A., Vezys, V. & Masopust, D. Sensing and alarm function of resident memory CD8(+) T cells. Nat. Immunol. 14, 509–513 (2013).
Bergsbaken, T. & Bevan, M.J. Proinflammatory microenvironments within the intestine regulate the differentiation of tissue-resident CD8(+) T cells responding to infection. Nat. Immunol. 16, 406–414 (2015).
Critchfield, J.W. et al. Magnitude and complexity of rectal mucosa HIV-1-specific CD8+ T-cell responses during chronic infection reflect clinical status. PLoS ONE 3, e3577 (2008).
Ferre, A.L. et al. Mucosal immune responses to HIV-1 in elite controllers: a potential correlate of immune control. Blood 113, 3978–3989 (2009).
Kiniry, B.E. et al. Predominance of weakly cytotoxic, T-betLowEomesNeg CD8+ T-cells in human gastrointestinal mucosa: implications for HIV infection. Mucosal Immunol. 10, 1008–1020 (2016).
Sathaliyawala, T. et al. Distribution and compartmentalization of human circulating and tissue-resident memory T cell subsets. Immunity 38, 187–197 (2013).
Booth, J.S. et al. Characterization and functional properties of gastric tissue-resident memory T cells from children, adults, and the elderly. Front. Immunol. 5, 294 (2014).
Shacklett, B.L. et al. Trafficking of human immunodeficiency virus type 1-specific CD8+ T cells to gut-associated lymphoid tissue during chronic infection. J. Virol. 77, 5621–5631 (2003).
Thome, J.J. et al. Spatial map of human T cell compartmentalization and maintenance over decades of life. Cell 159, 814–828 (2014).
Hamann, D. et al. Phenotypic and functional separation of memory and effector human CD8+ T cells. J. Exp. Med. 186, 1407–1418 (1997).
Mackay, C.R. Dual personality of memory T cells. Nature 401, 659–660 (1999).
Cepek, K.L. et al. Adhesion between epithelial cells and T lymphocytes mediated by E-cadherin and the alpha E beta 7 integrin. Nature 372, 190–193 (1994).
Agace, W.W., Higgins, J.M., Sadasivan, B., Brenner, M.B. & Parker, C.M. T-lymphocyte-epithelial-cell interactions: integrin alpha(E)(CD103)beta(7), LEEP-CAM and chemokines. Curr. Opin. Cell Biol. 12, 563–568 (2000).
Sheridan, B.S. et al. Oral infection drives a distinct population of intestinal resident memory CD8(+) T cells with enhanced protective function. Immunity 40, 747–757 (2014).
Burgener, A., McGowan, I. & Klatt, N.R. HIV and mucosal barrier interactions: consequences for transmission and pathogenesis. Curr. Opin. Immunol. 36, 22–30 (2015).
Laidlaw, B.J. et al. CD4+ T cell help guides formation of CD103+ lung-resident memory CD8+ T cells during influenza viral infection. Immunity 41, 633–645 (2014).
Hansen, S.G. et al. Immune clearance of highly pathogenic SIV infection. Nature 502, 100–104 (2013).
Ferre, A.L. et al. Immunodominant HIV-specific CD8+ T-cell responses are common to blood and gastrointestinal mucosa, and Gag-specific responses dominate in rectal mucosa of HIV controllers. J. Virol. 84, 10354–10365 (2010).
Park, S.L., Mackay, L.K. & Gebhardt, T. Distinct recirculation potential of CD69+CD103- and CD103+ thymic memory CD8+ T cells. Immunol. Cell Biol. 94, 975–980 (2016).
Mowat, A.M. Anatomical basis of tolerance and immunity to intestinal antigens. Nat. Rev. Immunol. 3, 331–341 (2003).
Zhang, N. & Bevan, M.J. Transforming growth factor-beta signaling controls the formation and maintenance of gut-resident memory T cells by regulating migration and retention. Immunity 39, 687–696 (2013).
Oh, S.A. & Li, M.O. TGF-beta: guardian of T cell function. J. Immunol. 191, 3973–3979 (2013).
Pauken, K.E. & Wherry, E.J. Overcoming T cell exhaustion in infection and cancer. Trends Immunol. 36, 265–276 (2015).
Shacklett, B.L., Critchfield, J.W. & Lemongello, D. Isolating mucosal lymphocytes from biopsy tissue for cellular immunology assays. Methods Mol. Biol. 485, 347–356 (2009).
Shacklett, B.L. et al. Optimization of methods to assess human mucosal T-cell responses to HIV infection. J. Immunol. Methods 279, 17–31 (2003).
Roederer, M., Nozzi, J.L. & Nason, M.C. SPICE: exploration and analysis of post-cytometric complex multivariate datasets. Cytometry Part A 79, 167–174 (2011).
Skinner, P.J., Daniels, M.A., Schmidt, C.S., Jameson, S.C. & Haase, A.T. Cutting edge: in situ tetramer staining of antigen-specific T cells in tissues. J. Immunol. 165, 613–617 (2000).
Skinner, P.J. & Haase, A.T. In situ staining using MHC class I tetramers. Curr. Protoc. Immunol. Chapter 17, Unit 17, 4 (2005).
Acknowledgements
We thank Rebecca Hoh, Montha Pao, Monika Deswal, and the clinical staff at San Francisco General Hospital for their assistance with participant recruitment and sample collection. We thank the participants for their willingness to contribute to this study. The SCOPE cohort was supported by the Delaney AIDS Research Enterprise (DARE; AI096109 and AI127966), NIAID (K24 AI069994), and the UCSF/Gladstone Institute of Virology & Immunology CFAR (P30 AI027763). We also thank Dr Alex Khoruts, University of Minnesota, for assistance with identification of anatomical compartments in rectosigmoid tissue samples. This work was supported by NIH/NIAID R01-AI057020 to B.L.S. The UC Davis Flow Cytometry Shared Resource Laboratory (FCSR) was supported by grants from the National Institutes of Health: NCI P30 CA0933730, and NIH NCRR C06-RR12088, S10-RR12964, S10-RR026825, and S10-OD018223-01A1; and the James B. Pendleton Charitable Trust. We thank the FCSR staff members Ms. Bridget McLaughlin and Mr Jonathan Van Dyke, for assistance with flow cytometry.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
Author contributions
Project concept and direction: B.L.S. with consultation from P.S., P.W.H., and S.G.D. Characterization of participant cohort and procurement of clinical samples: M.S., P.W.H., and S.G.D. Laboratory assays: B.E.K. (flow cytometry) with assistance from A.G.; S.L. (in situ tetramer staining). Data analysis: B.E.K., S.L., P.S., and B.L.S. Manuscript writing and editing: B.E.K. and B.L.S. All authors read and approved the manuscript.
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kiniry, B., Li, S., Ganesh, A. et al. Detection of HIV-1-specific gastrointestinal tissue resident CD8+ T-cells in chronic infection. Mucosal Immunol 11, 909–920 (2018). https://doi.org/10.1038/mi.2017.96
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2017.96
This article is cited by
-
Mucosal T-cell responses to chronic viral infections: Implications for vaccine design
Cellular & Molecular Immunology (2024)
-
Human mucosal tissue-resident memory T cells in health and disease
Mucosal Immunology (2022)
-
Local heroes or villains: tissue-resident memory T cells in human health and disease
Cellular & Molecular Immunology (2020)
-
Defining T Cell Tissue Residency in Humans: Implications for HIV Pathogenesis and Vaccine Design
Current HIV/AIDS Reports (2020)