Abstract
In inflammatory bowel diseases, a breakdown in host microbial interactions accompanies sustained activation of immune cells in the gut. Functional studies suggest a key role for interleukin-23 (IL-23) in orchestrating intestinal inflammation. IL-23 can be produced by various mononuclear phagocytes (MNPs) following acute microbial stimulation, but little is known about the key cellular sources of IL-23 that drive chronic intestinal inflammation. Here we have addressed this question using a physiological model of bacteria-driven colitis. By combining conditional gene ablation and gene expression profiling, we found that IL-23 production by CD11c+ MNPs was essential to trigger intestinal immunopathology and identified MHCII+ monocytes and macrophages as the major source of IL-23. Expression of IL-23 by monocytes was acquired during their differentiation in the intestine and correlated with the expression of major histocompatibility complex class II (MHCII) and CD64. In contrast, Batf3-dependent CD103+ CD11b- dendritic cells were dispensable for bacteria-induced colitis in this model. These studies reinforce the pathogenic role of monocytes in dysregulated responses to intestinal bacteria and identify production of IL-23 as a key component of this response. Further understanding of the functional sources of IL-23 in diverse forms of intestinal inflammation may lead to novel therapeutic strategies aimed at interrupting IL-23-driven immune pathology.
Similar content being viewed by others
Introduction
Inflammatory bowel diseases, including Crohn’s disease and ulcerative colitis, are chronic inflammatory disorders of the gastrointestinal tract commonly associated with high morbidity. The precise etiology of inflammatory bowel diseases is unknown, but involves a complex combination of host genetic, microbial, and environmental factors that alter mucosal immune responses.1
In healthy individuals, intestinal homeostasis is ensured through the tight control of aberrant immune responses by a complex regulatory network. Distinct subsets of mononuclear phagocytes (MNPs) are strategically located at the mucosal interface, residing either in gut-associated lymphoid tissue or dispersed throughout the lamina propria. Within these sites, two major populations of MNPs can be identified:2 classical dendritic cells (DCs), expressing the integrins CD11c and CD103 (αEβ7) with or without coexpression of CD11b, and monocyte-derived MNPs, expressing the fractalkine receptor (CX3CR1), CD11b, and various levels of CD11c. In addition, two minor populations of DCs can be found: CD103- CD11b+ CX3CR1int and CD103- CD11b- CX3CR1- DCs.3 Lamina propria CD103+ DCs arise from common DC progenitors and classical DC-committed precursor under the control of Fms-like tyrosine kinase 3 ligand. CD103+ CD11b+ DCs are predominantly located in the small intestine, where they constitutively sample antigen and migrate to the mesenteric lymph nodes to prime and polarize naive T cells.4, 5 In contrast, the colon mostly harbors CD103+ CD11b- DCs6 that are developmentally related to lymphoid-tissue resident CD8α+ DCs.7
In addition to classical DCs, a network of macrophages expressing high levels of CD11c and CX3CR1 is spread throughout the intestinal lamina propria. These cells contrast with macrophage populations in other tissues by their relatively short life span and their constitutive replenishment from the blood monocyte pool.8, 9, 10 Under homeostatic conditions, Ly6Chi monocytes entering the gut transit through a continuum of developmental intermediates before giving rise to fully mature CX3CR1hi macrophages. This process involves the stepwise acquisition of anti-inflammatory genes along with the expression of the major histocompatibility complex class II (MHCII) and downregulation of typical proinflammatory genes.11, 12 Ly6Chi monocytes also give rise to a heterogeneous Ly6Clo CX3CR1int cellular subset distinct from macrophages that share characteristics of macrophages and DCs11, 13 and can be distinguished by their differential expression of the tissue macrophage marker Fc gamma receptor 1 (CD64)14, 15 and F4/80.
Lately, many studies have highlighted the critical roles of these complementary subsets in shaping both homeostatic and inflammatory responses.5, 16, 17 Their precise roles in inflammation are only beginning to be unraveled, and definitive reports on their functional contribution to chronic intestinal inflammation are currently lacking.
Interleukin-23 (IL-23) is a heterodimeric cytokine comprising the IL-12p40 and IL-23p19 subunits.18 It has been well documented as a critical driver of multiple inflammatory conditions in mice and humans, and plays a key role in the pathogenesis of inflammatory bowel diseases.19, 20 Although the mechanism through which IL-23 drives intestinal inflammation has been studied extensively, little is known about the cellular sources implicated in the generation of IL-23-dependent intestinal inflammation in vivo. In this study, we used a bacterially driven model of intestinal inflammation with many cardinal features of human inflammatory bowel diseases to address the functional relevance of IL-23 production by CD11c+ antigen-presenting cells and to characterize the cellular subsets involved in this key pathophysiological process.
Results
H. hepaticus-driven inflammation alters the intestinal mononuclear phagocyte compartment
The Gram-negative bacterium Helicobacter hepaticus (Hh) is a noninvasive organism commonly found in the mucosal layer of the murine lower intestinal tract. Wild-type animals infected with the bacterium do not develop intestinal immunopathology and serve as carriers of the infection.21 However, mice with genetic deficiencies in the IL-10/IL-10R pathway are susceptible to experimental Hh infection and develop severe inflammation of the colon and cecum (typhlocolitis) associated with epithelial hyperplasia.22, 23 Similarly, Hh infection of wild-type mice with concomitant administration of anti-IL-10R monoclonal antibodies (mAbs) triggers IL-23-dependent intestinal inflammation along with a robust T helper type 1/type 17 (Th1/Th17)-polarized effector T cell response.23 Mice infected with Hh and treated with anti-IL-10R antibodies, but not uninfected or single-treated controls, develop histological features of colitis within 2–3 weeks after infection (Figure 1a). Hh and anti-IL-10R-treated mice further show a marked infiltration of leukocytes within the lamina propria (Figure 1b), including predominant populations of neutrophils and MHCII+ monocytes (Figure 1c and Supplementary Figure S1a online). In the noninflamed colon, monocytes entering the lamina propria from the blood preferentially give rise to tissue macrophages11 (defined as CD11b+ CX3CR1hi Ly6Clo cells, green subset), expressing high levels of CD64, MHCII, F4/80 and CD11c (Figure 1d and Supplementary Figure S1b). However, this process is dramatically altered during inflammation and we observed the marked accumulation of MHCII+ monocytes (defined as CD11b+ CX3CR1int Ly6Chi cells, red subset) within the colonic lamina propria. These cells are characterized by the expression of CD64 and MHCII and intermediate levels of F4/80, CD24, and CD11c (Figure 1c–e and Supplementary Figure S1a). At steady state, MHCII+ monocytes and macrophages express the mannose receptor (CD206), a marker associated with anti-inflammatory macrophage function.24 However, expression of CD206 was lower in MHCII+ monocytes and macrophages during colitis (Figure 1f), suggesting that not only monocyte but also macrophage functions were modified during inflammation. Indeed, we found that during inflammation, MHCII+ monocytes expressed higher amounts of tumor necrosis factor-α and IL-6 protein, and Nos2 (inducible nitric oxide synthase) mRNA, and that IL-6 was also increased in macrophages (Supplementary Figure S1c). IL-10 production by both MHCII+ monocytes and macrophages was also increased during inflammation (Supplementary Figure S1c), most likely reflecting an inflammation-driven negative feedback pathway.
Helicobacter hepaticus (Hh)-driven inflammation alters the intestinal mononuclear phagocyte compartment. CX3CR1GFP/+ mice were infected with Hh and treated with anti-IL-10R monoclonal antibody (mAb), or with the respective single controls as indicated, and were compared with uninfected controls (Ctrl). Mice were analyzed after 2–3 weeks. (a) Colon histopathology scores. (b) Total number of colonic lamina propria cells per mouse. (c) Frequency of myeloid cell subsets among total colonic CD11b+ leukocytes. (d) Representative fluorescence-activated cell sorting (FACS) plots and (e) histograms of the indicated myeloid subsets: MHCII+ monocytes (CD11b+ CX3CR1int Ly6Chi, red subset), macrophages (CD11b+ CX3CR1hi Ly6Clo cells, green subset), CD11b+ DCs (CD11c+ CD103- CD11b+ CX3CR1int cells, gray subset), and CD103+ DCs (CD11c+ CD103+ CX3CR1- cells, orange subset). All cells were pregated on live CD45+ leukocytes, excluding neutrophils and eosinophils. Fluorescence minus one (FMO) controls for CD11b+ MHCII+ cells are shown. (f) Frequency of CD206+ cells among the indicated subsets determined by flow cytometry. Data points represent individual mice, bars indicate medians. All data are representative of at least two independent experiments. **P<0.01, ***P<0.001 as determined by one-way analysis of variance (ANOVA) with Bonferroni’s post-test. DC, dendritic cell; IL-10, interleukin-10; LPL, lamina propria leukocyte.
Besides their differentiation into macrophages, MHCII+ monocytes have also been shown to differentiate into a Ly6Clo CX3CR1int heterogeneous subset related to both macrophages and classical DCs.11 Among this heterogeneous compartment, CD11b+ DCs (defined as CD11c+ CD103- CD11b+ CX3CR1int cells, gray subset) can easily be distinguished from macrophages under homeostatic conditions. CD11b+ DCs express MHCII, CD24, and CD11c, but lack expression of CD64 and F4/80, markers commonly associated with macrophages (Figure 1d and Supplementary Figure S1b). However, the distinction between macrophages and CD11b+ DCs was blurred during inflammation by the appearance of a CD64-expressing population (Figure 1d,e) and it is currently unclear how these cells ontogenically and functionally relate to steady-state macrophages or CD11b+ DCs.
During colitis, the marked accumulation of MHCII+ monocytes correlated with a reduced frequency of macrophages, CD11b+ DCs, and CD103+ DCs (defined as CD11c+ CD103+ CX3CR1- cells, orange subset) among CD45+ lamina propria cells (Figure 1c,d). However, absolute numbers indicated a small increase in CD11b+ DCs and CD103+ CD11b+ DCs. In contrast, the numbers of macrophages remained unchanged and CD103+ CD11b- DCs were slightly decreased (Supplementary Figure S1a).
Altogether, Hh infection in the presence of IL-10R blockade triggers intestinal inflammation associated with dramatic changes in the myeloid compartment. We observed the predominant accumulation of granulocytes and MHCII+ monocytes within the colonic lamina propria. In addition, MHCII+ monocytes and macrophages acquired a more proinflammatory profile.
CD11c+ mononuclear phagocytes drive colitis through the production of IL-23
Because IL-23 has been identified as the major driver of colitis in this model,23 we next investigated the functional relevance of IL-23 production by mononuclear phagocytes in the development of colitis in the Hh+anti-IL-10R model. Therefore, we crossed mice expressing green fluorescent protein (GFP) and Cre recombinase under the control of the CD11c promoter (encoded by Itgax)25, 26 with mice possessing two loxP sites flanking four exons of the Il23a gene27 to generate Cd11c-cre.Il23afl/fl (CD11cIL-23-) mice.
As CD11cIL-23- mice express GFP under the control of the CD11c promoter and therefore cannot be crossed to CX3CR1-GFP reporter mice, we relied on an alternative gating strategy to assess the myeloid compartment in these mice (Supplementary Figure S2). At steady state, CD11cIL-23- mice had similar numbers of leukocytes in the colon as their IL-23-proficient littermates (CD11cIL-23+) (Supplementary Figure S3a) and no significant changes in the frequencies of myeloid or T cells were observed (Supplementary Figure S3b). However, as IL-23 is required for maintaining differentiated Th17 cells,28 CD11cIL-23- mice displayed strongly reduced levels of IL-17+ and IL-17+ IFNγ+ CD4+ T cells in the colon (Supplementary Figure S3c).
Upon infection with Hh and anti-IL-10R treatment, CD11cIL-23- mice exhibited a striking reduction in colitis compared with CD11cIL-23+ mice, with reduced epithelial crypt hyperplasia and lower leukocyte infiltration throughout the colon and cecum (Figure 2a,b). CD11cIL-23- mice also showed no development of splenomegaly (Figure 2c), indicating that they were also protected from systemic signs of disease. Reduced pathology observed in CD11cIL-23- mice was not a consequence of a differential bacterial load, as Hh colonization levels were similar in CD11cIL-23- and CD11cIL-23+ groups (Figure 2d). Consistent with reduced pathological changes, both innate and adaptive effector cytokines were strongly reduced in the colon of CD11cIL-23- mice following Hh and IL-10R blockade (Figure 2e), as was the amount of IL-23 itself (Figure 2f).
Interleukin-23 (IL-23) from CD11c+ cells drives Helicobacter hepaticus (Hh)-induced colitis. CD11cIL-23+ and CD11cIL-23- mice were infected with Hh combined with anti-IL-10R monoclonal antibody (mAb) treatment and analyzed after 3 weeks. Uninfected mice served as control. (a) Representative micrographs of hematoxylin and eosin-stained sections (bar-200 μm) and histopathology scores of the colon and cecum. (b) Total number of lamina propria cells per mouse. (c) Spleen weight. (d) Hh colonization level quantified by quantitative PCR (qPCR). (e,f) Expression of cytokines assessed by (e) multiplex flow cytometric assay or (f) enzyme-linked immunosorbent assay (ELISA) on the supernatants of unstimulated lamina propria cells cultured for 48 h. Data points represent individual mice, bars indicate medians. Data are pooled from two independent experiments and are representative of four experiments. **P<0.01, ***P<0.001 as determined by (a–c) one-way analysis of variance (ANOVA) with Bonferroni’s post-test or (d–f) Mann–Whitney U-test. IFNγ, interferon-γ; TNFα, tumor necrosis factor-α.
In accordance with their reduced leukocyte infiltration, CD11cIL-23- mice showed a reduced frequency and number of both MHCII- and MHCII+ monocytes (Figure 3a) and neutrophils (Supplementary Figure S3d) within the lamina propria following Hh infection and IL-10R blockade compared with similarly treated CD11cIL-23+ littermates. The lower infiltration of monocytes in CD11cIL-23- mice correlated with a higher frequency of lamina propria macrophages, whereas the frequencies of CD11b+ DCs and CD103+ DCs were unchanged and absolute numbers were reduced (Supplementary Figure S3d).
MHCII+ monocytes and pathogenic T cells are reduced in Helicobacter hepaticus (Hh)-infected and anti-IL-10R-treated CD11cIL-23- mice. CD11cIL-23+ and CD11cIL-23- mice were infected with Hh combined with anti-IL-10R monoclonal antibody (mAb) treatment and analyzed after 3 weeks. (a) Representative fluorescence-activated cell sorting (FACS) plots, frequencies among CD45+ leukocytes, and absolute numbers of the indicated myeloid subsets in the colonic lamina propria. (b) Frequencies of CD4+ T cells among CD45+ leukocytes. (c) Representative FACS plots and frequencies of IFNγ+, IFNγ+ IL-17A+, and IL-17A+ cells among CD4+ T cells upon restimulation with phorbol 12-myristate 13-acetate (PMA) and ionomycin. Data points represent individual mice, bars indicate medians. Data are representative of two independent experiments. *P<0.05, **P<0.01 as determined by Mann–Whitney U-test. IFNγ, interferon-γ; IL, interleukin; MHCII, major histocompatibility complex class II; TCR, T cell receptor.
Because IL-23 drives intestinal inflammation through direct effects on T cells, promoting their accumulation and proliferation in the colon,29 we examined the colonic T cell compartment of CD11cIL-23- and CD11cIL-23+ mice during colitis. CD11cIL-23- mice displayed lower frequencies of colonic CD4+ T cells compared with their IL-23-proficient littermates (Figure 3b). Whereas CD11cIL-23+ mice mounted a robust Th1/Th17 effector response characterized by the emergence of single- and double-producing IL-17A+/IFNγ+ T cells, CD11cIL-23- mice showed only mild induction of the corresponding cytokines (Figure 3c).
Overall, the absence of colitis in CD11cIL-23- mice indicates that the production of IL-23 by CD11c-expressing MNPs plays a nonredundant role in the intestinal inflammatory response.
MHCII+ monocytes and macrophages are the major source of IL-23 upon H. hepaticus infection
To better understand the sequence of events associated with the development of colitis in IL-23-proficient mice, we first monitored IL-23 expression in leukocytes purified from the colonic lamina propria. Il23a mRNA expression was consistently detectable 4–5 days following initiation of colitis (Figure 4a), suggesting that a tissue-resident and/or a rapidly mobilized cellular subset is involved in the early production of IL-23 in the colon. We therefore chose this time point for subsequent analyses. At 4 days after infection with Hh and anti-IL-10R treatment, CD11cIL-23+ mice showed an increased infiltration of leukocytes in the lamina propria relative to uninfected control mice (Figure 4b). This influx correlated with increased numbers of all MNP subsets in the colon (Supplementary Figure S4a,b), as well as increased frequencies of both MHCII- and MHCII+ monocytes (Figure 4c,d), neutrophils (Figure 4d), CD103+ CD11b- DCs, and CD103+ CD11b+ DCs (Figure 4e). However, frequencies of macrophages and CD11b+ DCs among total leukocytes were unchanged at this time point (Figure 4d).
MHCII+ monocytes and macrophages are the major producers of interleukin-23 (IL-23) during Helicobacter hepaticus (Hh)-induced colitis. CD11cIL-23+ mice (Cre-GFP+) were infected with Hh combined with anti-IL-10R monoclonal antibody (mAb) treatment and analyzed 4 days after infection alongside uninfected controls (Ctrl). (a) Quantitative PCR (qPCR) analysis of Il23a mRNA expression in colonic tissue collected from five individual mice at the indicated time points. (b) Total number of lamina propria cells per mouse at day 4. (c) Representative fluorescence-activated cell sorting (FACS) plots and (d,e) frequencies of the indicated myeloid subsets (as defined in Supplementary Figure S2) among colonic CD45+ leukocytes. (f,g) Representative FACS plots of CD11c-Cre-associated green fluorescent protein (GFP) expression vs. (f) CD11c and (g) MHCII. (h) qPCR analysis of Il23a mRNA expression by Cre-GFP+ and Cre-GFP- cells FACS-sorted from colonic leukocytes. (i) Frequency of CD11c-Cre-associated GFP expression among the indicated colonic myeloid subsets. Rectangle indicates Cre-positive subsets that were selected for further analysis in j. (j) qPCR analysis of Il23a mRNA expression by myeloid subsets FACS-sorted from CD11cIL-23+ and CD11cIL-23- mice. Statistics comparing cells from CD11cIL-23+ and CD11cIL-23- mice are shown. (k) qPCR analysis of Il23a mRNA expression by FACS-sorted MHCII- and MHCII+ monocytes. Data points represent individual mice, bars indicate medians. All bar charts of FACS-sorted cells represent the mean±s.e.m. of 10–15 pooled mice, sorted in two biological replicates. Data are representative of two independent experiments. *P<0.05, **P<0.01, ***P<0.001, as determined by (a, j) one-way analysis of variance (ANOVA) with Bonferroni’s post-test or (b–h, k) Mann–Whitney U-test. DC, dendritic cell; MHCII, major histocompatibility complex class II.
To further characterize the myeloid subsets involved in IL-23-dependent intestinal inflammation, we examined CD11c-Cre-associated GFP expression by lamina propria cells from CD11cIL-23+ mice. Analysis of CD45+ cells confirmed that Cre-GFP expression was restricted to leukocytes expressing CD11c on their cell surface (Figure 4f), but was predominantly observed in CD11chi cells. Furthermore, Cre-associated GFP fluorescence was only observed in MHCII+ cells (Figure 4g) and, interestingly, Il23a mRNA expression was restricted to Cre-GFP+ cells (Figure 4h). The majority of lamina propria MNPs express high levels of CD11c (Figure 1e and Supplementary Figure S1b). Analysis of distinct myeloid subsets in CD11cIL-23+ mice revealed that CD11c-Cre-associated GFP expression was predominantly detected among macrophages, CD103+ CD11b- DCs, CD103+ CD11b+ DCs, (Figure 4i), and CD11b+ DCs (not shown). In addition, more than half of MHCII+ monocytes were positive for Cre-GFP, indicating that all those subsets might be a potential source of IL-23. In contrast, Cre expression was absent from eosinophils, neutrophils, and MHCII- monocytes (Figure 4i).
To characterize which of these subsets represent a functional source of IL-23 in vivo, we assessed Il23a mRNA expression by the corresponding populations sorted by flow cytometry from CD11cIL-23- and CD11cIL-23+ mice 4 days after Hh infection in the presence of IL-10R blockade. Strikingly, Il23a mRNA expression in CD11cIL-23+ cells was mostly restricted to MHCII+ monocytes and macrophages, with low expression detectable among CD103+ CD11b- DCs and CD103+ CD11b+ DCs (Figure 4j). Furthermore, Il23a expression by both MHCII+ monocytes and macrophages was markedly reduced in CD11cIL-23- mice, indicating that CD11c-Cre-expression was active within these cellular subsets (Figure 4j). The low Il23a expression observed in MHCII- monocytes (Figure 4k) suggests that induction of IL-23 occurs in the lamina propria during their local maturation and acquisition of MHCII.
As the level of CX3CR1 expression identifies discrete subsets of MNPs, we repeated our analysis in CX3CR1GFP/+ mice. In uninfected mice, low levels of constitutive Il23a were detected among MHCII+ monocytes, CD103+ CD11b+ DCs, (Supplementary Figure S4c,d), and CD11b+ DCs (Supplementary Figure S4d). However, increased expression of Il23a at 4 days following infection with Hh and IL-10R blockade was only consistently detected in MHCII+ monocytes (Supplementary Figure S4d) and macrophages (not shown). During the course of inflammation, high Il23a expression was maintained in MHCII+ monocytes and their developmental progeny, but not CD103+ DCs (Figure 5a). Among CX3CR1int subsets, expression of Il23a was further restricted to CD64+ cells (Figure 5b).
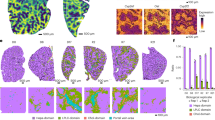
Interleukin-23 (IL-23) production during colitis is restricted to CX3CR1-expressing CD64+ subsets. CX3CR1GFP/+ mice were infected with Helicobacter hepaticus (Hh) combined with anti-IL-10R monoclonal antibody (mAb) treatment and analyzed 2 weeks after infection alongside uninfected controls (Ctrl). Colonic lamina propria cells were sorted by flow cytometry and Il23a mRNA expression was analyzed by quantitative PCR (qPCR). (a) Il23a mRNA expression and representative fluorescence-activated cell sorting (FACS) plots of the following sorted populations: (1) MHCII+ monocytes (Mono), (2) Ly6Clo CX3CR1int macrophages/DCs, (3) CX3CR1hi macrophages, and (4) CD103+ DCs. (b) Il23a mRNA expression and representative FACS plots of the following sorted populations: (1) CX3CR1int MHCII+ CD64- and (2) CX3CR1int MHCII+ CD64+ cells. Bar charts represent the mean+s.e.m. of 10–15 pooled mice, sorted in two biological replicates. Data are representative of three independent experiments. **P<0.01, ***P<0.001, as determined by one-way analysis of variance (ANOVA) with Bonferroni’s post-test. DC, dendritic cell; Macs, macrophages; MHCII, major histocompatibility complex class II; Mono, monocyte; NS, not significant.
Taken together, our data indicate that induction of IL-23 in colitis is contained within CX3CR1-expressing CD11c+ cells that also express MHCII and CD64.
Batf3-dependent CD103+ DCs are dispensable for chronic colitis
CD103+ CD11b+ DCs are predominantly found in the mouse small intestine and are nearly absent from the colon.6 Instead, most colonic CD103+ DCs are CD11b- and require the transcription factor Batf3 for their development.30 Batf3−/− mice therefore lack tissue-resident CD8α+ and CD103+ CD11b- nonlymphoid DC subsets. As expected, CD103+ CD11b- cells were strongly reduced in the colonic lamina propria of Batf3−/− mice, whereas CD103+ CD11b+ DCs were unaffected (Figure 6a,b). To assess whether CD103+ CD11b- DCs might be involved in the development of colitis, we infected Batf3−/− mice with Hh combined with anti-IL-10R treatment. At 2 weeks after initiation of colitis, numbers of CD103+ CD11b- DCs were still significantly reduced in the lamina propria (Figure 6b), but no differences in leukocyte infiltration (Figure 6c) or severity of colitis (Figure 6d) were observed in Batf3−/− mice compared with their wild-type counterparts, indicating that Batf3-dependent DCs are dispensable for the induction of IL-23-driven intestinal inflammation in this model.
Batf3-dependent dendritic cells (DCs) are dispensable for the production of interleukin-23 (IL-23) in Helicobacter hepaticus (Hh)-driven colitis. Wild-type (WT) and Batf3−/− mice were infected with Hh combined with anti-IL-10R monoclonal antibody (mAb) treatment and analyzed after 2 weeks alongside uninfected controls (Ctrl). (a) Representative fluorescence-activated cell sorting (FACS) plots and (b) frequencies and numbers of CD103+ CD11b- and CD103+ CD11b+ cells among CD45+ colonic leukocytes. (c) Total number of lamina propria cells per mouse. (d) Histopathology scores of the colon. Data points represent individual mice, bars indicate medians. *P<0.05 as determined by Mann–Whitney U-test, NS, not significant.
Mature macrophages produce IL-23 in response to H. hepaticus stimulation in vitro, but share functional redundancy with MHCII+ monocytes in vivo
To decipher the contribution of macrophages to IL-23 production, we generated bone marrow-derived macrophages (BMDMs) from CD11cIL-23- and CD11cIL-23+ mice. After differentiation with L929 cell-conditioned medium, around half of the BMDMs were positive for Cre-GFP and expressed CD64, F4/80, and CD11c (Supplementary Figure S5a). BMDMs were stimulated in vitro with live Hh bacteria in the presence of anti-IL-10R antibody and their response was compared with the respective controls. Strikingly, stimulation of CD11cIL-23+ BMDMs with Hh resulted in Il23a mRNA expression and this response was enhanced in the presence of IL-10R signaling blockade, whereas Il23a was not expressed by unstimulated BMDMs or those treated with anti-IL-10R antibody alone (Supplementary Figure S5b). Similarly, infection of mice with Hh only induced a low burst of IL-23 in monocytes and macrophages in vivo and this response was strongly increased in the absence of IL-10R signaling (Supplementary Figure S5c).
Intestinal tissue macrophages exclusively derive from blood monocytes and do not self-renew.8, 10 Colony-stimulating factor 1 receptor (CSF-1R) signaling controls the differentiation of Ly6Chi into Ly6Clo monocytes31 and is required for the maturation and replacement of resident-type Ly6Clo monocytes and tissue macrophages, but is dispensable for Ly6Chi monocyte production or inflammatory function.32 To assess the relative contributions of MHCII+ monocytes and tissue macrophages to the development of IL-23-dependent intestinal inflammation, we pretreated CX3CR1GFP/+ mice with an anti-CSF-1R-blocking mAb or isotype control before induction of colitis. A group of mice was analyzed at day 0 (Supplementary Figure S6a,b), whereas the remaining mice were treated with Hh and anti-IL-10R and continued to receive anti-CSF-1R treatment. At 4 days after Hh infection and anti-IL-10R treatment, macrophages were almost absent from the colon of anti-CSF-1R-treated mice. The heterogeneous Ly6Clo CX3CR1int macrophage/DC compartment was also significantly reduced in these mice (Figure 7a,b and Supplementary Figure S6c). Although the total number of colonic leukocytes was unchanged (Figure 7c), there was an increase in the proportion of granulocytes in these conditions (Figure 7d). Interestingly, the depletion of macrophages further correlated with increased Il23a expression within the lamina propria at day 4 (Figure 7e). However, anti-CSF-1R-mediated depletion of macrophages throughout the whole course of colitis did not affect the number of inflammatory cells accumulating in the colon (Figure 7f) or the severity of intestinal pathology (Figure 7g) 2 weeks following Hh infection and IL-10R signaling blockade, suggesting that MHCII+ monocytes and macrophages may exhibit functional redundancy during inflammation.
Macrophages share functional redundancy with MHCII+ monocytes for interleukin-23 (IL-23) production. CX3CR1GFP/+ mice were treated with a blocking anti-CSF-1R monoclonal antibody (mAb) or isotype (Iso) control and analyzed (a–e) 4 days or (f, g) 2 weeks after Helicobacter hepaticus (Hh) and anti-IL-10R mAb treatment. (a) Representative fluorescence-activated cell sorting (FACS) plots and (b) frequencies among CD45+ leukocytes of the indicated myeloid subsets, as defined in Figure 1c. (c) Total number of colonic lamina propria cells per mouse. (d) Frequencies of neutrophils among CD45+ cells. (e) Quantitative PCR (qPCR) analysis of Il23a mRNA expression by total colonic lamina propria cells analyzed at day 4. (f) Total number of colonic lamina propria cells per mouse 2 weeks after Hh and anti-IL-10R mAb treatment. (g) Colonic histopathology scores. Data points represent individual mice, bars indicate medians. Data are pooled from two experiments and representative of three independent experiments. *P<0.05, **P<0.01, as determined by Mann–Whitney U-test. CSF, colony-stimulating factor; DC, dendritic cell; Macs, macrophages; MHCII, major histocompatibility complex class II; NS, not significant; Uninf, uninfected.
Discussion
In this work, we show that intestinal CD11c+ mononuclear phagocytes play a nonredundant role in the inflammatory process triggered by Hh in the absence of IL-10R signaling through the production of IL-23. Both innate and adaptive immune responses were impaired in the absence of CD11c-restricted IL-23 production, preventing the characteristic intestinal infiltration of granulocytes, monocytes, and pathogenic T cells observed in this model. Multiple lamina propria MNPs express CD11c and analysis of Il23a gene expression from various colonic CD11c+ subsets identified MHCII+ monocytes and macrophages as the most abundant cellular sources of IL-23 during Hh+anti-IL-10R-induced inflammation, supporting a functional contribution of these cells during colitis. In contrast, IL-23-driven colitis was independent of Batf3-dependent CD103+ CD11b- DCs. These results are consistent with our previous studies showing a pathogenic role for infiltrating monocytes during T cell-driven colitis16 and further show that production of IL-23 is a major determinant of this function.
Others have identified monocytes/macrophages as a source of IL-23 upon intestinal inflammation.33, 34, 35 However, none of those studies have examined the relative expression of IL-23 by distinct MNP subsets in a systematic way nor have addressed the functional relevance of this expression in the context of chronic intestinal inflammation. This is mostly because of the difficulty of accurately identifying individual subsets of macrophages and DCs within the lamina propria owing to substantial overlap in their expression of surface markers.
Although CX3CR1hi macrophages have been described as a decisive factor determining gut health and inflammation,36 we have made the intriguing observation that intestinal macrophages dependent on CSF-1 for their survival were dispensable for the production of Il23a and development of colitis in our model. This indicates that newly recruited monocytes entering the lamina propria from the blood do not require full differentiation to launch a proinflammatory response and can drive colitis in the absence of macrophages. It further suggests that fully mature macrophages share some functional redundancy with MHCII+ monocytes regarding the production of IL-23. This observation is in line with the distinctive origin of intestinal macrophages, which are replenished exclusively from the blood monocyte pool through a series of developmental intermediates.10 Blood monocytes show induction of MHCII after or in concert with their arrival into tissues, possibly through interaction with the endothelium.37 Our observation that both MHCII- and CD64- subsets do not express Il23a suggests that the induction of IL-23 occurs in a particular, more mature monocyte developmental stage coinciding with the upregulation of MHCII and CD64.
DC-derived IL-23 has been shown to be involved in both homeostatic and inflammatory mucosal functions. Thus, interferon regulatory factor 4-dependent CD103+ CD11b+ intestinal DCs drive homeostatic Th17 cell differentiation,4, 38 a function associated with increased IL-12/IL-23 production.39 Similarly, we observed that CD11cIL-23- mice had strongly reduced levels of IL-17A+ cells in the colonic lamina propria and low levels of constitutive Il23a were detected among colonic CD103+ CD11b+ DCs and CD11b+ DCs during the steady state. In models of acute intestinal inflammation, Notch2-dependent CD103+ CD11b+ DCs are an obligate source of IL-23 for driving the IL-22-dependent antimicrobial response to infection with the attaching-and-effacing mouse pathogen Citrobacter rodentium.40 CD103+ CD11b+ DCs of the small intestine were also shown to produce a Toll-like receptor 5-dependent burst of IL-23 at ∼30 min following systemic flagellin administration, suggesting that this subset of DCs can induce IL-23-driven responses upon Toll-like receptor 5 stimulation.41 In contrast, others have shown that CD103+ CD11b+ DCs were dispensable for IL-22-mediated protection against C. rodentium,42 whereas depletion of CX3CR1+ MNPs resulted in more severe colitis and death in C. rodentium-infected mice.35 Together with our own data, these diverging results nicely illustrate that the functional sources of IL-23 that drive host protection or immune pathology vary and are not surprisingly highly context dependent. In our model, the development of colonic inflammation and histopathological changes induced by the noninvasive pathosymbiont Hh crucially depends on IL-23 production by CD11c+ cells. Induced Il23a expression is found at highest amounts in MHCII+ monocytes, which combined with their numerical dominance, suggests a major functional role of these cells in the pathogenesis of intestinal inflammation. However, given the ontogenic and phenotypical heterogeneity observed among the lamina propria CD11c+ MNP compartment during colitis and the difficulty to accurately distinguish CD11b+ DCs from macrophages because of their overlapping CD64 expression in inflammation, we cannot formally rule out a functional contribution of bona fide DCs in this model. Further studies specifically targeting the DC lineage will help to clarify this point.
Although IL-23 promotes Th17 responses, it is unclear whether IL-17 itself plays a pathogenic role in IL-10-deficient models of colitis. Indeed, IL-17A production by T cells was dispensable for T cell transfer colitis,43 and neutralization of IL-17 was not sufficient to inhibit colitis in IL-10-deficient mice, being only partially protective when combined with anti-IL-6 treatment.44 This suggests that IL-23 can drive IL-17-independent inflammatory pathways in the intestine. Our recent studies have suggested a key role for IL-23-driven granulocyte macrophage-CSF as a major downstream mediator that promotes chronic colitis through its ability to stimulate the myeloid cell response.45 Others have shown that granulocyte macrophage-CSF can promote IL-23 production by MNPs,46 suggesting a positive feedback loop that may contribute to T cell-mediated perpetuation of the innate inflammatory response.
Overall, our observation that intestinal CX3CR1-expressing CD11c+ cells are the major source of IL-23 in a physiological model of bacteria-driven colitis confirms the pathogenic potential of these cells during colitis and uncovers a novel mechanism through which they might perpetuate chronic inflammation. Further understanding of the functional sources of IL-23 in diverse forms of intestinal inflammation may lead to novel therapeutic strategies aimed at interrupting IL-23-driven immune pathology.
Methods
Mice. Itgax-cre-EGFP (Cd11c-cre)25 mice express a Cre recombinase gene, IRES, and an EGFP gene downstream of the 5.3 kb (full length) mouse CD11c promoter/enhancer. Expression of EGFP is expected to have equimolar expression with Cre recombinase. The promoter activity of CD11c-Cre-GFP strain 4097 has been described in detail elsewhere.26 Itgax-cre-EGFP,25 Il23afl/fl,27 Cd11c-cre.Il23afl/fl, CX3CR1GFP/+,47 and Batf3−/−30 mice were bred and maintained under specific pathogen-free conditions in accredited animal facilities at the University of Oxford. Experiments were conducted in accordance with the UK Scientific Procedures Act (1986) under a Project License (PPL) authorized by the UK Home Office. Mice were routinely screened for the absence of pathogens and Helicobacter species and were over 5 weeks of age when used.
Induction of colitis. Hh NCI-Frederick isolate 1A (strain 51449) was grown as described previously.48 Mice were fed on two consecutive days with Hh 1A (∼1.0 × 108 colony-forming units) by oral gavage and injected intraperitoneally with 1 mg anti-IL-10R (clone 1B1.2) mAb weekly, starting on day 0. Unless specified in the corresponding figures, mice were killed 2–3 weeks after infection.
Histology. Upon killing, samples of cecum and intact proximal, mid, and distal colon were fixed in phosphate-buffered 4% formalin. Next, 4–5 μm paraffin-embedded sections were stained with hematoxylin and eosin, and histopathology of the colon and cecum was assessed using a scoring system for epithelial hyperplasia, goblet cell depletion, leukocyte infiltration, and submucosal inflammation, as previously described.48 Photomicrographs were taken using a Nikon Coolscope (Nikon, Tokyo, Japan).
Isolation of leukocytes. For lamina propria leukocyte isolation, the colon was opened longitudinally, washed, and cut into pieces. Pieces were washed twice in Hanks’ balanced salt solution supplemented with 4% fetal bovine serum, 100 U ml−1 penicillin/streptomycin, and 5 mM EDTA at 37 °C with shaking to remove epithelial cells. Tissue was then digested at 37 °C in a shaking incubator with 1 mg ml−1 type VIII collagenase (Sigma-Aldrich, Gillingham, UK) and 0.5 mg ml−1 DNase I in RPMI-1640 medium supplemented with 4% fetal bovine serum and 100 U ml−1 penicillin/streptomycin. Cells were then layered on a 40/80% Percoll gradient, centrifuged, and the interface was recovered. Cells were counted using a Casy Cell Counter and Analyzer Model TT (Innovatis, Roche Diagnostics, Burgess Hill, UK).
Flow cytometry and cell sorting. For surface staining, cells were stained in phosphate-buffered saline/0.1% bovine serum albumin/5 mM EDTA buffer with a fixable viability dye and a combination of the following antibodies: CD45 (30-F11), CD11c (N418), MHCII (M5/114.15.2), F4/80 (BM8), and TCRβ (H57-597) (all from eBioscience, Hatfield, UK); CD103 (M290), CD11b (M1/70), and Siglec-F (E50-2440) (all from BD Biosciences, Oxford, UK); Ly6G (1A8), Ly6C (HK1.4), CD64 (X54-5/7.1), CD4 (RM4-5), CD24 (M1/69), and CD206 (C068C2) (all from BioLegend, London, UK). Fc block (anti-CD16/CD32, eBioscience) was included to minimize nonspecific antibody binding. For intracellular cytokine staining of T cells, cells were incubated for 3.5 h in complete IMDM medium containing 0.1 μM phorbol 12-myristate 13-acetate and 1 μM ionomycin with 1:1,000 GolgiPlug and GolgiStop solutions (BD Biosciences) at 37 °C in a humidified incubator with 5% CO2. Following surface staining, cells were fixed and permeabilized with the Cytofix/Cytoperm Fixation/Permeabilization Solution Kit (BD Biosciences) according to the manufacturer’s instructions. Cells were stained for 50 min with antibodies to IL-17A (17B7, eBioscience) and IFNγ (XMG1.2, eBioscience). All cells were analyzed on a SORP LSRII (BD Biosciences) or sorted on a FACSAriaIII (BD Biosciences) to a purity of >95%. Analysis was performed using FlowJo software (Tree Star, Ashland, OR).
Quantitative PCR. RNA was isolated from snap-frozen colonic tissue (combined proximal, mid, and distal colon sections), isolated lamina propria cells, BMDMs, or from FACS-sorted cells using the RNeasy Mini kit (QIAGEN, Manchester, UK) according to the manufacturer’s instructions, including an on-column DNase I digestion step. Complementary DNA synthesis was performed using Superscript III reverse transcriptase (QIAGEN) or High-Capacity cDNA Reverse Transcriptase (Applied Biosystems, Life Technologies, Paisley, UK). Quantitative PCR reactions for the candidate genes were performed using TaqMan gene expression assays (Life Technologies). Complementary DNA samples were analyzed in duplicate using the CFX96 detection system (Bio-Rad Laboratories, Hemel Hempstead, UK) or ViiA 7 Real-Time PCR System (Life Technologies), and gene expression levels for each sample were normalized to HPRT. Mean relative gene expression was determined, and the differences were calculated using the 2ΔC(t) method. Primer pairs and probes were as follows: TaqMan Gene Expression Assays for mouse Hprt (Mm01545399_m1), Il23a (Mm00518984_m1), and (Mm01160011_g1).
Quantitation of cytokine protein levels. For detection of cytokines by the Flowcytomix Multiplex system (Life Technologies), 5 × 105 leukocytes were cultured for 48 h in complete RPMI medium at 37 °C. Cytokines in the supernatant were detected using Simplex kits for mouse tumor necrosis factor-α, IL-1β, IL-17A, interferon-γ (IFNγ), and IL-22 according to the manufacturer’s instructions and analyzed by flow cytometry. IL-23 protein levels were quantified using the mouse IL-23 ELISA Ready-SET-Go! kit (eBioscience).
Quantitation of H. hepaticus. Hh colonization levels were quantified in cecal contents collected upon killing. DNA was isolated using the DNA Stool kit (QIAGEN) and quantitative PCR with Hh-specific primers against the cdtB gene was performed as previously described.48
Depletion of macrophages with anti-CSF-1R mAbs. Anti-CSF-1R mAb was purified from the culture supernatants of AFS98 hybridoma, kindly provided by Dr Miriam Merad (Mount Sinai School of Medicine, New York, NY). Mice were injected intraperitoneally with anti-CSF-1R mAb (AFS98) or rat IgG1 isotype control (HRPN, BioXcell, West Lebanon, NH) at doses of 1 mg per mouse on day −4 followed by 0.3 mg per mouse from day −3 to day 0. A group of mice was analyzed at day 0, whereas the remaining mice were infected with Hh and anti-IL-10R and continued to receive anti-CSF-1R treatment at 0.3 mg per mouse every second day until killing at day 4 or 14.
Statistics. Mann–Whitney U-test or one-way analysis of variance with Bonferroni’s post-test correction was performed using GraphPad Prism version 6.02 for Windows (GraphPad Software, La Jolla, CA) as indicated. Differences were considered statistically significant when P<0.05.
References
Kaser, A., Zeissig, S. & Blumberg, R.S. Inflammatory bowel disease. Annu. Rev. Immunol. 28, 573–621 (2010).
Geissmann, F., Manz, M.G., Jung, S., Sieweke, M.H., Merad, M. & Ley, K. Development of monocytes, macrophages, and dendritic cells. Science 327, 656–661 (2010).
Cerovic, V. et al. Intestinal CD103(-) dendritic cells migrate in lymph and prime effector T cells. Mucosal Immunol. 6, 104–113 (2013).
Persson, E.K. et al. IRF4 transcription-factor-dependent CD103(+)CD11b(+) dendritic cells drive mucosal T helper 17 cell differentiation. Immunity 38, 958–969 (2013).
Coombes, J.L. et al. A functionally specialized population of mucosal CD103+ DCs induces Foxp3+ regulatory T cells via a TGF-beta and retinoic acid-dependent mechanism. J. Exp. Med. 204, 1757–1764 (2007).
Denning, T.L. et al. Functional specializations of intestinal dendritic cell and macrophage subsets that control Th17 and regulatory T cell responses are dependent on the T cell/APC ratio, source of mouse strain, and regional localization. J. Immunol. 187, 733–747 (2011).
Edelson, B.T. et al. Batf3-dependent CD11b(low/-) peripheral dendritic cells are GM-CSF-independent and are not required for Th cell priming after subcutaneous immunization. PLoS One 6, e25660 (2011).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Jaensson, E. et al. Small intestinal CD103+ dendritic cells display unique functional properties that are conserved between mice and humans. J. Exp. Med. 205, 2139–2149 (2008).
Bain, C.C. et al. Constant replenishment from circulating monocytes maintains the macrophage pool in the intestine of adult mice. Nat. Immunol. 15, 929–937 (2014).
Bain, C.C. et al. Resident and pro-inflammatory macrophages in the colon represent alternative context-dependent fates of the same Ly6Chi monocyte precursors. Mucosal Immunol. 6, 498–510 (2013).
Rivollier, A., He, J., Kole, A., Valatas, V. & Kelsall, B.L. Inflammation switches the differentiation program of Ly6Chi monocytes from antiinflammatory macrophages to inflammatory dendritic cells in the colon. J. Exp. Med. 209, 139–155 (2012).
Zigmond, E. et al. Ly6C hi monocytes in the inflamed colon give rise to proinflammatory effector cells and migratory antigen-presenting cells. Immunity 37, 1076–1090 (2012).
Tamoutounour, S. et al. CD64 distinguishes macrophages from dendritic cells in the gut and reveals the Th1-inducing role of mesenteric lymph node macrophages during colitis. Eur. J. Immunol. 42, 3150–3166 (2012).
Gautier, E.L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Siddiqui, K.R., Laffont, S. & Powrie, F. E-cadherin marks a subset of inflammatory dendritic cells that promote T cell-mediated colitis. Immunity 32, 557–567 (2010).
Farache, J., Zigmond, E., Shakhar, G. & Jung, S. Contributions of dendritic cells and macrophages to intestinal homeostasis and immune defense. Immunol. Cell Biol. 91, 232–239 (2013).
Oppmann, B. et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity 13, 715–725 (2000).
Duerr, R.H. et al. A genome-wide association study identifies IL23R as an inflammatory bowel disease gene. Science 314, 1461–1463 (2006).
Hue, S. et al. Interleukin-23 drives innate and T cell-mediated intestinal inflammation. J. Exp. Med. 203, 2473–2483 (2006).
Kullberg, M.C. et al. Helicobacter hepaticus triggers colitis in specific-pathogen-free interleukin-10 (IL-10)-deficient mice through an IL-12- and gamma interferon-dependent mechanism. Infect. Immun. 66, 5157–5166 (1998).
Morrison, P.J. et al. Th17-cell plasticity in Helicobacter hepaticus-induced intestinal inflammation. Mucosal Immunol. 6, 1143–1156 (2013).
Kullberg, M.C. et al. IL-23 plays a key role in Helicobacter hepaticus-induced T cell-dependent colitis. J. Exp. Med. 203, 2485–2494 (2006).
Stein, M., Keshav, S., Harris, N. & Gordon, S. Interleukin 4 potently enhances murine macrophage mannose receptor activity: a marker of alternative immunologic macrophage activation. J. Exp. Med. 176, 287–292 (1992).
Stranges, P.B. et al. Elimination of antigen-presenting cells and autoreactive T cells by Fas contributes to prevention of autoimmunity. Immunity 26, 629–641 (2007).
Zhu, X.J., Yang, Z.F., Chen, Y., Wang, J. & Rosmarin, A.G. PU.1 is essential for CD11c expression in CD8(+)/CD8(-) lymphoid and monocyte-derived dendritic cells during GM-CSF or FLT3L-induced differentiation. PLoS One 7, e52141 (2012).
Thakker, P., Leach, M.W., Kuang, W., Benoit, S.E., Leonard, J.P. & Marusic, S. IL-23 is critical in the induction but not in the effector phase of experimental autoimmune encephalomyelitis. J. Immunol. 178, 2589–2598 (2007).
Chen, Y. et al. Anti-IL-23 therapy inhibits multiple inflammatory pathways and ameliorates autoimmune encephalomyelitis. J. Clin. Invest. 116, 1317–1326 (2006).
Ahern, P.P. et al. Interleukin-23 drives intestinal inflammation through direct activity on T cells. Immunity 33, 279–288 (2010).
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8alpha+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008).
Hashimoto, D. et al. Pretransplant CSF-1 therapy expands recipient macrophages and ameliorates GVHD after allogeneic hematopoietic cell transplantation. J. Exp. Med. 208, 1069–1082 (2011).
MacDonald, K.P. et al. An antibody against the colony-stimulating factor 1 receptor depletes the resident subset of monocytes and tissue- and tumor-associated macrophages but does not inhibit inflammation. Blood 116, 3955–3963 (2010).
Sheikh, S.Z., Matsuoka, K., Kobayashi, T., Li, F., Rubinas, T. & Plevy, S.E. Cutting edge: IFN-gamma is a negative regulator of IL-23 in murine macrophages and experimental colitis. J. Immunol. 184, 4069–4073 (2010).
Kamada, N. et al. Unique CD14 intestinal macrophages contribute to the pathogenesis of Crohn disease via IL-23/IFN-gamma axis. J. Clin. Invest. 118, 2269–2280 (2008).
Longman, R.S. et al. CX(3)CR1(+) mononuclear phagocytes support colitis-associated innate lymphoid cell production of IL-22. J. Exp. Med. 211, 1571–1583 (2014).
Zigmond, E. et al. Macrophage-restricted interleukin-10 receptor deficiency, but not IL-10 deficiency, causes severe spontaneous colitis. Immunity 40, 720–733 (2014).
Jakubzick, C. et al. Minimal differentiation of classical monocytes as they survey steady-state tissues and transport antigen to lymph nodes. Immunity 39, 599–610 (2013).
Schlitzer, A. et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 38, 970–983 (2013).
Scott, C.L. et al. CCR2CD103 intestinal dendritic cells develop from DC-committed precursors and induce interleukin-17 production by T cells. Mucosal Immunol. 8, 327–339 (2014).
Satpathy, A.T. et al. Notch2-dependent classical dendritic cells orchestrate intestinal immunity to attaching-and-effacing bacterial pathogens. Nat. Immunol. 14, 937–948 (2013).
Kinnebrew, M.A. et al. Interleukin 23 production by intestinal CD103(+)CD11b(+) dendritic cells in response to bacterial flagellin enhances mucosal innate immune defense. Immunity 36, 276–287 (2012).
Welty, N.E., Staley, C., Ghilardi, N., Sadowsky, M.J., Igyarto, B.Z. & Kaplan, D.H. Intestinal lamina propria dendritic cells maintain T cell homeostasis but do not affect commensalism. J. Exp. Med. 210, 2011–2024 (2013).
Noguchi, D. et al. Blocking of IL-6 signaling pathway prevents CD4+ T cell-mediated colitis in a T(h)17-independent manner. Int. Immunol. 19, 1431–1440 (2007).
Yen, D. et al. IL-23 is essential for T cell-mediated colitis and promotes inflammation via IL-17 and IL-6. J. Clin. Invest. 116, 1310–1316 (2006).
Griseri, T., McKenzie, B.S., Schiering, C. & Powrie, F. Dysregulated hematopoietic stem and progenitor cell activity promotes interleukin-23-driven chronic intestinal inflammation. Immunity 37, 1116–1129 (2012).
El-Behi, M. et al. The encephalitogenicity of T(H)17 cells is dependent on IL-1- and IL-23-induced production of the cytokine GM-CSF. Nat. Immunol. 12, 568–575 (2011).
Jung, S. et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Maloy, K.J., Salaun, L., Cahill, R., Dougan, G., Saunders, N.J. & Powrie, F. CD4+CD25+ T(R) cells suppress innate immune pathology through cytokine-dependent mechanisms. J. Exp. Med. 197, 111–119 (2003).
Acknowledgements
We thank Paresh Thakker for providing Il23afl/fl mice, Miriam Merad for providing the AFS98 hybridoma cell line, Fanny Franchini for technical assistance, Philip Ahern for his comments and suggestions regarding the manuscript, Helen Ferry for flow cytometry cell sorting, and Richard Stillion for histology. I.C.A. was supported by a Swiss National Science Foundation fellowship. S.M. was supported by a BBSRC Industrial CASE Studentship in association with UCB. A.N.H. was supported by a European Molecular Biology Organization (EMBO) long-term fellowship (ALTF 116-2012) and a Marie Curie fellowship (FP7-PEOPLE-2012-IEF, Proposal 330621). This work was supported by the Wellcome Trust UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Arnold, I., Mathisen, S., Schulthess, J. et al. CD11c+ monocyte/macrophages promote chronic Helicobacter hepaticus-induced intestinal inflammation through the production of IL-23. Mucosal Immunol 9, 352–363 (2016). https://doi.org/10.1038/mi.2015.65
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2015.65
This article is cited by
-
CX3CR1 modulates SLE-associated glomerulonephritis and cardiovascular disease in MRL/lpr mice
Inflammation Research (2023)
-
Identification of gut microbial species linked with disease variability in a widely used mouse model of colitis
Nature Microbiology (2022)
-
IL-23 signaling prevents ferroptosis-driven renal immunopathology during candidiasis
Nature Communications (2022)
-
Autophagy impairment in liver CD11c+ cells promotes non-alcoholic fatty liver disease through production of IL-23
Nature Communications (2022)
-
Intestinal epithelium-derived BATF3 promotes colitis-associated colon cancer through facilitating CXCL5-mediated neutrophils recruitment
Mucosal Immunology (2021)