Abstract
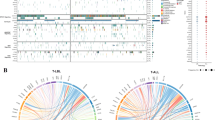
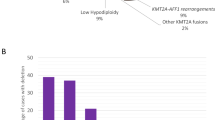
It has been generally acknowledged that the diagnosis, treatment and prognosis evaluation of leukemia largely rely on an adequate identification of genetic abnormalities. A systemic analysis of genetic aberrations was performed in a cohort of 1346 patients with newly diagnosed acute lymphoblastic leukemia (ALL) in China. The pediatric patients had higher incidence of hyperdiploidy and t(12;21) (p13;q22)/ETV6–RUNX1 than adults (P<0.0001); in contrast, the occurrence of Ph and Ik6 variant of IKZF1 gene was much more frequent in adult patients (all P<0.0001). In B-ALL, the existence of Ik6 and that of BCR–ABL were statistically correlated (P<0.0001). In comparison with Western cohorts, the incidence of t(9;22) (q34;q11)/BCR–ABL (14.60%) in B-ALL and HOX11 expression in T-ALL (25.24%) seemed to be much higher in our group, while the incidence of t(12;21) (p13;q22)/ETV6–RUNX1 (15.34%) seemed to be lower in Chinese pediatric patients. The occurrence of hyperdiploidy was much lower either in pediatric (10.61% vs 20–38%) or adult patients (2.36% vs 6.77–12%) in our study than in Western reports. In addition, the frequencies of HOX11L2 in adult patients were much higher in our cohort than in Western countries (20.69% vs 4–11%). In general, it seems that Chinese ALL patients bear more adverse prognostic factors than their Western counterparts do.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Harrison CJ . Cytogenetics of paediatric and adolescent acute lymphoblastic leukaemia. Br J Haematol 2009; 144: 147–156.
Pui CH, Relling MV, Downing JR . Acute lymphoblastic leukemia. N Engl J Med 2004; 350: 1535–1548.
Szczepanski T, Harrison CJ, van Dongen JJ . Genetic aberrations in paediatric acute leukaemias and implications for management of patients. Lancet Oncol 2010; 11: 880–889.
Pui CH, Evans WE . Treatment of acute lymphoblastic leukemia. N Engl J Med 2006; 354: 166–178.
Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR et al. Proposals for the classification of the acute leukaemias. French-American-British (FAB) co-operative group. Br J Haematol 1976; 33: 451–458.
Bene MC, Castoldi G, Knapp W, Ludwig WD, Matutes E, Orfao A et al. Proposals for the immunological classification of acute leukemias. European Group for the Immunological Characterization of Leukemias (EGIL). Leukemia 1995; 9: 1783–1786.
EGIL (European Group for the Immunological Classification of Leukaemias). The value of c-kit in the diagnosis of biphenotypic acute leukemia. Leukemia 1998; 12: 2038.
ISCN 2005. An International System for Human Cytogenetic Nomenclature 2005, In: Shaffer LG, Tommerup N, (eds) S Karger: Basel.
Pallisgaard N, Hokland P, Riishoj DC, Pedersen B, Jorgensen P . Multiplex reverse transcription-polymerase chain reaction for simultaneous screening of 29 translocations and chromosomal aberrations in acute leukemia. Blood 1998; 92: 574–588.
Zhu YM, Zhao WL, Fu JF, Shi JY, Pan Q, Hu J et al. NOTCH1 mutations in T-cell acute lymphoblastic leukemia: prognostic significance and implication in multifactorial leukemogenesis. Clin Cancer Res 2006; 12: 3043–3049.
Asnafi V, Radford-Weiss I, Dastugue N, Bayle C, Leboeuf D, Charrin C et al. CALM-AF10 is a common fusion transcript in T-ALL and is specific to the TCRgammadelta lineage. Blood 2003; 102: 1000–1006.
Meleshko AN, Movchan LV, Belevtsev MV, Savitskaja TV . Relative expression of different Ikaros isoforms in childhood acute leukemia. Blood Cells Mol Dis 2008; 41: 278–283.
Martinelli G, Iacobucci I, Storlazzi CT, Vignetti M, Paoloni F, Cilloni D et al. IKZF1 (Ikaros) deletions in BCR-ABL1-positive acute lymphoblastic leukemia are associated with short disease-free survival and high rate of cumulative incidence of relapse: a GIMEMA AL WP report. J Clin Oncol 2009; 27: 5202–5207.
Cario G, Zimmermann M, Romey R, Gesk S, Vater I, Harbott J et al. Presence of the P2RY8-CRLF2 rearrangement is associated with a poor prognosis in non-high-risk precursor B-cell acute lymphoblastic leukemia in children treated according to the ALL-BFM 2000 protocol. Blood 2010; 115: 5393–5397.
Yoda A, Yoda Y, Chiaretti S, Bar-Natan M, Mani K, Rodig SJ et al. Functional screening identifies CRLF2 in precursor B-cell acute lymphoblastic leukemia. Proc Natl Acad Sci USA 2010; 107: 252–257.
Russell LJ, Capasso M, Vater I, Akasaka T, Bernard OA, Calasanz MJ et al. Deregulated expression of cytokine receptor gene, CRLF2, is involved in lymphoid transformation in B-cell precursor acute lymphoblastic leukemia. Blood 2009; 114: 2688–2698.
Mullighan CG, Collins-Underwood JR, Phillips LA, Loudin MG, Liu W, Zhang J et al. Rearrangement of CRLF2 in B-progenitor- and Down syndrome-associated acute lymphoblastic leukemia. Nat Genet 2009; 41: 1243–1246.
Harvey RC, Mullighan CG, Chen IM, Wharton W, Mikhail FM, Carroll AJ et al. Rearrangement of CRLF2 is associated with mutation of JAK kinases, alteration of IKZF1, Hispanic/Latino ethnicity, and a poor outcome in pediatric B-progenitor acute lymphoblastic leukemia. Blood 2010; 115: 5312–5321.
Poppe B, Dastugue N, Vandesompele J, Cauwelier B, De Smet B, Yigit N et al. EVI1 is consistently expressed as principal transcript in common and rare recurrent 3q26 rearrangements. Genes Chromosomes Cancer 2006; 45: 349–356.
La Starza R, Aventin A, Crescenzi B, Gorello P, Specchia G, Cuneo A et al. CIZ gene rearrangements in acute leukemia: report of a diagnostic FISH assay and clinical features of nine patients. Leukemia 2005; 19: 1696–1699.
De Keersmaecker K, Graux C, Odero MD, Mentens N, Somers R, Maertens J et al. Fusion of EML1 to ABL1 in T-cell acute lymphoblastic leukemia with cryptic t(9;14) (q34;q32). Blood 2005; 105: 4849–4852.
Dube ID, Raimondi SC, Pi D, Kalousek DK . A new translocation, t(10;14) (q24;q11), in T cell neoplasia. Blood 1986; 67: 1181–1184.
Soudon J, Bernard O, Mathieu-Mahul D, Larsen CJ . c-myc gene expression in a leukemic T-cell line bearing a t(8;14) (q24;q11) translocation. Leukemia 1991; 5: 60–65.
Barriga F, Bertin P, Legues E, Risueno C, Andrade W, Cabrera E et al. t(1;5) (q23;q33) in a patient with high-risk B-lineage acute lymphoblastic leukemia. Cancer Genet Cytogenet 1996; 87: 4–6.
Bousquet M, Broccardo C, Quelen C, Meggetto F, Kuhlein E, Delsol G et al. A novel PAX5-ELN fusion protein identified in B-cell acute lymphoblastic leukemia acts as a dominant negative on wild-type PAX5. Blood 2007; 109: 3417–3423.
Akasaka T, Balasas T, Russell LJ, Sugimoto KJ, Majid A, Walewska R et al. Five members of the CEBP transcription factor family are targeted by recurrent IGH translocations in B-cell precursor acute lymphoblastic leukemia (BCP-ALL). Blood 2007; 109: 3451–3461.
Pan Q, Zhu YJ, Gu BW, Cai X, Bai XT, Yun HY et al. A new fusion gene NUP98-IQCG identified in an acute T-lymphoid/myeloid leukemia with a t(3;11) (q29q13;p15)del(3) (q29) translocation. Oncogene 2008; 27: 3414–3423.
Ariffin H, Chen SP, Kwok CS, Quah TC, Lin HP, Yeoh AE . Ethnic differences in the frequency of subtypes of childhood acute lymphoblastic leukemia: results of the Malaysia-Singapore Leukemia Study Group. J Pediatr Hematol Oncol 2007; 29: 27–31.
Liang DC, Shih LY, Yang CP, Hung IJ, Liu HC, Jaing TH et al. Frequencies of ETV6-RUNX1 fusion and hyperdiploidy in pediatric acute lymphoblastic leukemia are lower in far east than west. Pediatr Blood Cancer 2010; 55: 430–433.
Moorman AV, Ensor HM, Richards SM, Chilton L, Schwab C, Kinsey SE et al. Prognostic effect of chromosomal abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: results from the UK Medical Research Council ALL97/99 randomised trial. Lancet Oncol 2010; 11: 429–438.
Gaynon PS, Trigg ME, Heerema NA, Sensel MG, Sather HN, Hammond GD et al. Children's Cancer Group trials in childhood acute lymphoblastic leukemia: 1983–1995. Leukemia 2000; 14: 2223–2233.
Mancini M, Scappaticci D, Cimino G, Nanni M, Derme V, Elia L et al. A comprehensive genetic classification of adult acute lymphoblastic leukemia (ALL): analysis of the GIMEMA 0496 protocol. Blood 2005; 105: 3434–3441.
Moorman AV, Harrison CJ, Buck GA, Richards SM, Secker-Walker LM, Martineau M et al. Karyotype is an independent prognostic factor in adult acute lymphoblastic leukemia (ALL): analysis of cytogenetic data from patients treated on the Medical Research Council (MRC) UKALLXII/Eastern Cooperative Oncology Group (ECOG) 2993 trial. Blood 2007; 109: 3189–3197.
A Collaborative Study of the Group Francais de Cytogenetique Hematologique. Cytogenetic abnormalities in adult acute lymphoblastic leukemia: correlations with hematologic findings outcome. Blood 1996; 87: 3135–3142.
Secker-Walker LM, Prentice HG, Durrant J, Richards S, Hall E, Harrison G . Cytogenetics adds independent prognostic information in adults with acute lymphoblastic leukaemia on MRC trial UKALL XA. MRC Adult Leukaemia Working Party. Br J Haematol 1997; 96: 601–610.
van Grotel M, Meijerink JP, van Wering ER, Langerak AW, Beverloo HB, Buijs-Gladdines JG et al. Prognostic significance of molecular-cytogenetic abnormalities in pediatric T-ALL is not explained by immunophenotypic differences. Leukemia 2008; 22: 124–131.
Cave H, Suciu S, Preudhomme C, Poppe B, Robert A, Uyttebroeck A et al. Clinical significance of HOX11L2 expression linked to t(5;14) (q35;q32), of HOX11 expression, and of SIL-TAL fusion in childhood T-cell malignancies: results of EORTC studies 58881 and 58951. Blood 2004; 103: 442–450.
Ferrando AA, Neuberg DS, Dodge RK, Paietta E, Larson RA, Wiernik PH et al. Prognostic importance of TLX1 (HOX11) oncogene expression in adults with T-cell acute lymphoblastic leukaemia. Lancet 2004; 363: 535–536.
Asnafi V, Beldjord K, Libura M, Villarese P, Millien C, Ballerini P et al. Age-related phenotypic and oncogenic differences in T-cell acute lymphoblastic leukemias may reflect thymic atrophy. Blood 2004; 104: 4173–4180.
Baak U, Gokbuget N, Orawa H, Schwartz S, Hoelzer D, Thiel E et al. Thymic adult T-cell acute lymphoblastic leukemia stratified in standard- and high-risk group by aberrant HOX11L2 expression: experience of the German multicenter ALL study group. Leukemia 2008; 22: 1154–1160.
Klein F, Feldhahn N, Herzog S, Sprangers M, Mooster JL, Jumaa H et al. BCR-ABL1 induces aberrant splicing of IKAROS and lineage infidelity in pre-B lymphoblastic leukemia cells. Oncogene 2006; 25: 1118–1124.
Fuxa M, Busslinger M . Reporter gene insertions reveal a strictly B lymphoid-specific expression pattern of Pax5 in support of its B cell identity function. J Immunol 2007; 178: 3031–3037.
Adams B, Dorfler P, Aguzzi A, Kozmik Z, Urbanek P, Maurer-Fogy I et al. Pax-5 encodes the transcription factor BSAP and is expressed in B lymphocytes, the developing CNS, and adult testis. Genes Dev 1992; 6: 1589–1607.
Taki T, Kano H, Taniwaki M, Sako M, Yanagisawa M, Hayashi Y . AF5q31, a newly identified AF4-related gene, is fused to MLL in infant acute lymphoblastic leukemia with ins(5;11) (q31;q13q23). Proc Natl Acad Sci USA 1999; 96: 14535–14540.
Collins-Underwood JR, Mullighan CG . Genomic profiling of high-risk acute lymphoblastic leukemia. Leukemia 2010; 24: 1676–1685.
Hess JL . MLL: a histone methyltransferase disrupted in leukemia. Trends Mol Med 2004; 10: 500–507.
Garcia-Manero G, Yang H, Kuang SQ, O’Brien S, Thomas D, Kantarjian H . Epigenetics of acute lymphocytic leukemia. Semin Hematol 2009; 46: 24–32.
Garcia-Manero G, Daniel J, Smith TL, Kornblau SM, Lee MS, Kantarjian HM et al. DNA methylation of multiple promoter-associated CpG islands in adult acute lymphocytic leukemia. Clin Cancer Res 2002; 8: 2217–2224.
Wetzler M, Dodge RK, Mrozek K, Carroll AJ, Tantravahi R, Block AW et al. Prospective karyotype analysis in adult acute lymphoblastic leukemia: the cancer and leukemia group B experience. Blood 1999; 93: 3983–3993.
Jabbour EJ, Faderl S, Kantarjian HM . Adult acute lymphoblastic leukemia. Mayo Clin Proc 2005; 80: 1517–1527.
Borkhardt A, Cazzaniga G, Viehmann S, Valsecchi MG, Ludwig WD, Burci L et al. Incidence and clinical relevance of TEL/AML1 fusion genes in children with acute lymphoblastic leukemia enrolled in the German and Italian multicenter therapy trials. Associazione Italiana Ematologia Oncologia Pediatrica and the Berlin-Frankfurt-Munster Study Group. Blood 1997; 90: 571–577.
Schrappe M, Reiter A, Ludwig WD, Harbott J, Zimmermann M, Hiddemann W et al. Improved outcome in childhood acute lymphoblastic leukemia despite reduced use of anthracyclines and cranial radiotherapy: results of trial ALL-BFM 90. German-Austrian-Swiss ALL-BFM Study Group. Blood 2000; 95: 3310–3322.
Hann I, Vora A, Harrison G, Harrison C, Eden O, Hill F et al. Determinants of outcome after intensified therapy of childhood lymphoblastic leukaemia: results from Medical Research Council United Kingdom acute lymphoblastic leukaemia XI protocol. Br J Haematol 2001; 113: 103–114.
Acknowledgements
This work was supported in part by the Mega project of Ministry of Science and Technology (2008ZX09312-026, 2009ZX09103-431), the National Basic Research Program (973, 2010CB529200), the National Natural Science Foundation of China (30871106, 30772744, 30821063) and Science and Technology Commission Foundation of Shanghai (09dZ1974500).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, B., Wang, YY., Shen, Y. et al. Newly diagnosed acute lymphoblastic leukemia in China (I): abnormal genetic patterns in 1346 childhood and adult cases and their comparison with the reports from Western countries. Leukemia 26, 1608–1616 (2012). https://doi.org/10.1038/leu.2012.26
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2012.26
Keywords
This article is cited by
-
Ectopic expression of a combination of 5 genes detects high risk forms of T-cell acute lymphoblastic leukemia
BMC Genomics (2022)
-
Cytogenetic Characteristics of Childhood Acute Lymphoblastic Leukemia: A Study of 1541 Chinese Patients Newly Diagnosed between 2001 and 2014
Current Medical Science (2022)
-
Anti-CD19 CAR-T cell therapy bridge to HSCT decreases the relapse rate and improves the long-term survival of R/R B-ALL patients: a systematic review and meta-analysis
Annals of Hematology (2021)
-
Genetic mutational analysis of pediatric acute lymphoblastic leukemia from a single center in China using exon sequencing
BMC Cancer (2020)
-
Treatment response, survival, safety, and predictive factors to chimeric antigen receptor T cell therapy in Chinese relapsed or refractory B cell acute lymphoblast leukemia patients
Cell Death & Disease (2020)