Abstract
POGZ, the gene encoding pogo transposable element-derived protein with zinc-finger domain, has been implicated in autism spectrum disorder and it is widely expressed in the human tissues, including the brain. Intellectual disability (ID) is highly heterogeneous neurodevelopment disorder and affects ~2–3% of the general population. Here we report the identification of a novel frameshift mutation in the coding region of the POGZ gene (c.1277_1278insC), which occurred de novo in a Chinese patient with ID. In silico analysis and western blotting revealed this frameshift mutation generating truncated protein in peripheral blood lymphocytes, and this may disrupt several important domains of POGZ gene. Our finding broadens the spectrum of POGZ mutations and may help to understand the molecular basis of ID and aid genetic counseling.
Similar content being viewed by others
Introduction
Intellectual disability (ID) is a heterogeneous group of neurodevelopmental disorders and may be caused by genetic and environmental factors. About estimate 2–3% of the general population is affected in ID.1 The etiology of ID is genetically heterogeneous, and de novo point mutations may account for 22% in severe ID cases.2 Recently, hundreds of candidate ID genes were revealed by large-scale trio-based next-generation sequencing studies,2, 3 suggesting de novo mutations are strongly enriched in ID patients.
POGZ is a gene that codes for the pogo transposable element-derived protein with zinc finger domain, which plays a role in mitotic cell cycle progression, and is involved in kinetochore assembly and mitotic sister chromatid cohesion.4 Recently researches highlight the gene as a chromatin modifier that has significant role in psychiatric disorders and autism spectrum disorder (ASD).5, 6, 7 Here we report a novel de novo frameshift mutation in POGZ gene in Chinese patient with ID.
Materials and Methods
Case report
A female patient aged 5 years, the first child of healthy non-consanguineous parents, was born at 39 weeks of gestation after a normal pregnancy. Prenatal period was normal. Her birth weight was 3013 g (+3.39 s.d.). Family history was negative for ID and other neurological disorder. She could raise her head at 4 months and walked alone at 18 months. Later her speech and physical development was delayed. She could only count numbers within 20, and her developmental quotient was 54 at 3.5 years of age.
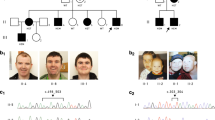
On examination at 5 years of age, her weight was 18 kg (+0.04 s.d.), height was 108 cm (+0.24 s.d.) and occipital frontal circumference was 47cm (+0.08 s.d.). Patient was assessed by using Wechsler Preschool and Primary Scale of Intelligence-4th edn, autism behavior checklist and Achenbach system of empirically based assessment. Patient could only speak simple sentences, had a mild ID (intelligent quotient=63). Patient had a low frustration tolerance and displayed intermittent tantrum behaviors. The craniofacial features consisting of high-arched eyebrows, a broad, low nasal bridge, anteverted nares and a thin upper lip (Figure 1). Single-nucleotide polymorphism array analysis (Illumina Human660 BeadsChip, Illumina Inc., San Diego, CA, USA) confirmed no pathogenic copy-number variation change in the patient.
Clinical photographs of patient harboring a de novo mutation (c.1277_1278insC) in POGZ. (a) Frontal view and (b) lateral view demonstrate high-arched eyebrows, broad and flat nasal bridge and thin upper lip. A full color version of this figure is available at the Journal of Human Genetics journal online.
Next-generation sequencing
In this study, we used molecular inversion probes to sequence in 764 patients (ID 729, epilepsy 27 and ASD 8) and 772 of parents’ samples. Informed consent from all the parents and approval from the local institutional review board were obtained for the molecular studies. DNA was extracted from peripheral blood samples.
Molecular inversion probes were designed to cover coding region of POGZ gene. The libraries were amplified during 22 cycles of PCR, during which an 8-bp sample barcode was introduced. After the barcode library was purified and quantified, it was sequenced via paired-end 100-bp reads with an 8-bp barcode read on Illumina HiSeq2000 sequencer. The reads were performed arm trimming (MIPGen v.1.0), alignment (BWA v.0.7.8), multi-sample genotype calling (GATK Unified Genotyper v.3.2-2) and variants were annotated with SeattleSeq138. Variants with an alternative allele frequency <0.005 in the NHLBI Exome Sequencing Project Exome Variant Server (ESP6500), or not present in our exome database of ~500 individuals and the 1000 Genomes Browser were included before the analysis as previously described.8 The variants were validated by Sanger sequencing.
Western blot analysis
In the patient, the c.1277_1278insC frameshift mutation in the exon 9 disrupts the zinc-finger domain. To validate the functionality of de novo mutation in POGZ, we detected the POGZ encode protein by western blot analysis in peripheral blood lymphocytes. The protein lysates were electrophoresed on 8% SDS–polyacrylamide gel electrophoresis gel and transferred onto a polyvinylidene difluride membrane. The membranes were incubated with anti-human POGZ polyclonal antibody (rabbit polyclonal; Abcam, Cambridge, MA, USA), followed by detection with secondary horseradish peroxidase-conjugated goat anti-rabbit IgG antibody. Proteins were visualized using the enhanced chemiluminescence system. Densitometry was analyzed by Quality One Image software (Biorad, Hercules, CA, USA).
Results and Discussion
In this study, we found one novel de novo mutation of POGZ in patient with mild ID, leading to the cause of a premature truncated protein in peripheral blood lymphocytes (Figure 2a). The antibody detected wild-type POGZ at the correct size (predicted molecular weight ~155 kD), and also detected the truncated protein in patient (Figure 3). The expression of wild-type POGZ protein in the patient was reduced respectively (n=3, P<0.05, Student's t-test).
(a) Electropherograms of the patient and her parents showing the presence of the de novo mutation in PO GZ gene. (b) Schematic overview of the POGZ full-length and truncating protein, the frameshift variation was marked by black arrow. In silico analysis the frameshift mutation (c.1277_1278insC) was predicted causing a premature truncated protein in POGZ with a short N-terminal domain.
Western blot analysis of POGZ protein variation from patient and his parents in peripheral blood lymphocytes. Anti-POGZ antibodies were used to recognize the wild type and novel POGZ mutant. Densitometric results shows wild-type POGZ level was significantly reduced in the patient (n=3; P<0.05, Student's t-test).
ID is a multifarious neurodevelopmental disorder and may exhibit overlapping phenotypes with autism and epilepsy.9 The next-generation sequencing provides us a great opportunity to study the genetic cause of psychiatric disorder. Recently studying the de novo variations offered more insights into genetic-determining genes in ID.10 In previous studies, loss of function mutations in POGZ gene were found significantly enriched in patients with ASD and ID by large-scale next-generation sequencing.11, 12 Our case provides evidence for the important role of POGZ gene in the development of ID.
The POGZ gene is located on chromosome 1q21.3, which encodes several domains including multiple C2H2-type zinc fingers, a centromere protein (CENP) B-like DNA-binding domain, and a DDE domain, which may have an important role in mitosis and neuronal proliferation.4, 13 The DDE domain is characterized by a catalytic site composed of two or three aspartic acid, which allows DNA-modifying reactions such as strand cleaving, nicking and ligation.14 Recent research suggests those DNA-binding domain protein may function in the role of human neurodevelopmental disorders.15
For now, eight de novo POGZ mutations have been identified through next-generation sequencing, including missense, frameshift and stop-gain mutations, which have been associated with ID and ASD.16, 17 Here we report the first de novo frameshift mutation in the zinc-fingers domain, yielding a premature truncated protein that may disrupt DNA and protein binding activation (Figure 2b).
Moreover, a recent report described a Japanese patient with a de novo missense mutation identified by trio-based exome sequencing. The mutation was located in the CENP B-like DNA-binding domain and the patient was described having ASD, severe ID and global developmental delay. After the identification of this de novo mutation in POGZ, we tried to collect all available patient's clinical data and images to reevaluate the diagnosis of psychiatric disorder. Psychiatrist evaluated the patient and found mild ID with stereotyped behaviors in this patient; however, ASD was absent (autism behavior checklist total score 19; sensory 3, relating 0, body and object use 7, language 3, social and self-help skills 6). Photos of the patient at the age of 5 years clearly showed such as broad and flat nasal bridge and thin upper lip consistent with previously described in the Japanese patient.17 Those evidence were found in our patient suggest phenotypic heterogeneity may exist among patients with mutation in POGZ gene.
In summary, the patient features are consistent with mild ID caused by a de novo frameshift mutation in POGZ gene. Our research supports the hypothesis that de novo variants represent significant causal factors in ID and extend the phenotype of patient with POGZ haploinsufficiency.
References
de Vries, B. B., Pfundt, R., Leisink, M., Koolen, D. A., Vissers, L. E., Janssen, I. M. et al. Diagnostic genome profiling in mental retardation. Am. J. Hum. Genet. 77, 606–616 (2005).
de Ligt, J., Willemsen, M. H., van Bon, B. W., Kleefstra, T., Yntema, H. G., Kroes, T. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Rauch, A., Wieczorek, D., Graf, E., Wieland, T., Endele, S., Schwarzmayr, T. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Nozawa, R. S., Nagao, K., Masuda, H. T., Iwasaki, O., Hirota, T., Nozaki, N. et al. Human POGZ modulates dissociation of HP1alpha from mitotic chromosome arms through Aurora B activation. Nat. Cell Biol. 12, 719–727 (2010).
Cotney, J., Muhle, R. A., Sanders, S. J., Liu, L., Willsey, A. J., Niu, W. et al. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat. Commun. 6, 6404 (2015).
Martinez de Paz, A., Vicente Sanchez-Mut, J., Samitier-Marti, M., Petazzi, P., Saez, M., Szczesna, K. et al. Circadian cycle-dependent MeCP2 and brain chromatin changes. PloS ONE 10, e0123693 (2015).
Turner, T. N., Sharma, K., Oh, E. C., Liu, Y. P., Collins, R. L., Sosa, M. X. et al. Loss of delta-catenin function in severe autism. Nature 520, 51–56 (2015).
O'Roak, B. J., Vives, L., Fu, W., Egertson, J. D., Stanaway, I. B., Phelps, I. G. et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 338, 1619–1622 (2012).
Karam, S. M., Riegel, M., Segal, S. L., Felix, T. M., Barros, A. J., Santos, I. S. et al. Genetic causes of intellectual disability in a birth cohort: a population-based study. Am. J. Med. Genet. A 167, 1204–1214 (2015).
Neale, B. M., Kou, Y., Liu, L., Ma'ayan, A., Samocha, K. E., Sabo, A. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
De Rubeis, S., He, X., Goldberg, A. P., Poultney, C. S., Samocha, K., Cicek, A. E. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Deciphering Developmental Disorders Study Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
Hormozdiari, F., Penn, O., Borenstein, E. & Eichler, E. E. The discovery of integrated gene networks for autism and related disorders. Genome Res. 25, 142–154 (2015).
Bartholomeeusen, K., Christ, F., Hendrix, J., Rain, J. C., Emiliani, S., Benarous, R. et al. Lens epithelium-derived growth factor/p75 interacts with the transposase-derived DDE domain of PogZ. J. Biol. Chem. 284, 11467–11477 (2009).
Micucci, J. A., Sperry, E. D. & Martin, D. M. Chromodomain helicase DNA-binding proteins in stem cells and human developmental diseases. Stem Cells Dev. 24, 917–926 (2015).
Iossifov, I., O'Roak, B. J., Sanders, S. J., Ronemus, M., Krumm, N., Levy, D. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Fukai, R., Hiraki, Y., Yofune, H., Tsurusaki, Y., Nakashima, M., Saitsu, H. et al. A case of autism spectrum disorder arising from a de novo missense mutation in POGZ. J. Hum. Genet. 60, 277–279 (2015).
Acknowledgements
This study was supported by grants from the National Key Basic Research Program of China (2012CB944600) and the National Key Technology R&D Program of China (2012BAI09B05). We thank the family members of the patient for their participation in our study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tan, B., Zou, Y., Zhang, Y. et al. A novel de novo POGZ mutation in a patient with intellectual disability. J Hum Genet 61, 357–359 (2016). https://doi.org/10.1038/jhg.2015.156
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.156
This article is cited by
-
White-Sutton syndrome and congenital heart disease: case report and literature review
BMC Pediatrics (2023)
-
Autism-associated protein POGZ controls ESCs and ESC neural induction by association with esBAF
Molecular Autism (2022)
-
Pogz deficiency leads to transcription dysregulation and impaired cerebellar activity underlying autism-like behavior in mice
Nature Communications (2020)
-
Pathogenic POGZ mutation causes impaired cortical development and reversible autism-like phenotypes
Nature Communications (2020)