Abstract
Sandhoff disease (SD) is an autosomal recessive neurodegenerative lysosomal storage disorder caused by mutations in HEXB gene. Molecular pathology is unknown in Indian patients with SD. The present study is aimed to determine mutations spectrum and molecular pathology leading to SD in 22 unrelated patients confirmed by the deficiency of β-hexosaminidase-A and total-hexosaminidase in leukocytes. To date, nearly 86 mutations of HEXB have been described, including five large deletions. Over all we have identified 13 mutations in 19 patients, eight of which were novel, including two missense mutations [c.611G>A (p.G204E), c. 634A>T (p.H212Y)], two nonsense mutations [c.333G>A (p.W111X), c.298C>T (p.R100X)], one splice site mutation c.1082+5 G>T, two small in-frame deletions [c.534_541delAGTTTATC (p.V179RfsX10), c.1563_1573delTATGGATGACG (p.M522LfsX2)] and one insertion c.1553_1554insAAGA (p.D518EfsX8). We have also identified previously known, five sequence variations leading to amino acid changes [c.926G>A (p.C309Y), c.1597C>T (p.R533C)], one nonsense mutation c.850 C>T (p.R284X), one splice site mutation c.1417+1 G-A and one insertion c.1591_1592insC (p.R531TfsX22). Mutation was not identified in three patients. We observed from this study that mutation c.850C>T (p.R284X) was identified in 4/19 (21%) patients which is likely to be the most common mutation in the country. This is the first study providing insight into the molecular basis of SD in India.
Similar content being viewed by others
Introduction
GM2 gangliosidoses, the second most common lipid storage disorder, occurs due to deficiency of β-Hexosaminidase enzyme (EC 3.2.1.52) and is among the majority of lysosomal storage disorders (LSDs) studies in India.1 The disorders included are Tay–Sachs disease (TSD) (MIM 272800), Sandhoff disease (SD) (MIM 268800) and GM2 activator deficiency or GM2 gangliosidosis AB variant (GM2AD) (MIM 272750). They are caused by mutations in the respective genes HEXA (MIM 606869; locus 15q23), HEXB (MIM 606873; locus 5q13.3) and GM2A (MIM 613109; locus 5q33.1). Recently, our group identified mutations in HEXA gene in Indian patients with TSD.2, 3 This study demonstrated novel molecular basis for TSD in India. There is an allelic heterogeneity amongst infantile TSD patients with the exception of the founder mutation p.E462V in patients from Gujarat. SD has phenotype similar to TSD and clinical features include progressive neurodegeneration, seizures, macrocephaly, spasticity and cherry red spots on fundus examination. Liver and spleen enlargement is unique to SD. Several studies on lysosomal storage disorders have been reported from India.1, 4, 5, 6, 7, 8, 9 However, the molecular basis of SD is unknown in Indian patients. We report the clinical, biochemical and molecular profile in patients of SD from twenty three unrelated families from India.
Materials and Methods
Subject and sample collection
Twenty two families included in the study. The institutional ethics committees from both the participating institutes have approved the study. Clinical details were filled up in the case record forms and blood samples were collected after an informed written consent obtained from the guardian of the study subjects. The clinical inclusion criteria included progressive loss of acquired milestones, seizures, macrocephaly, liver/spleen enlargement and cherry red spot on fundus examination. Exclusion criteria included 'Hurler' facies, corneal haziness, dysotosis multiplex which pointed toward other storage disorders such as GM1 gangliosidosis, Niemann Pick disease, mucopolysaccharidoses and mucolipidoses. Neuroimaging criteria included presence of gray matter disease shown by hyperintensity in basal ganglia and/or hypointensity in thalamus in T2 weighted images of magnetic resonance imaging of the brain.
Enzyme assay
Six milliliter peripheral blood was collected in EDTA vacutainer from all the patients for leukocyte enzyme assay and DNA extraction. The enzyme activity was determined by fluorimetric method using specific synthetic substrate. The total-hexosaminidase (total Hex) was measured from the hydrolysis of the synthetic substrate 4-methylumbelliferyl-N-acetyl-β-d-glucos amine (MUG), which releases fluorescent 4-methylumbelliferone when acted upon by β-hexosaminidase. Hexosaminidase-B (Hex-B) was determined as the activity left after the samples were heated for 3 hours at 50 °C. This procedure led to loss of Hex-A which is heat-labile but not of Hex-B or intermediate isoenzyme. Therefore, Hex-A activity was obtained after subtracting Hex-B activity from total Hex activity. Hex-A was also assayed with the more specific sulphated substrate 4-methylumbelliferyl-N-acetyl-β-d-glucos amine-6-sulphate (MUGS).10 Hex-A activity (obtained from MUGS lysis) was expressed as percentage of total Hex activity (obtained from MUG lysis).
Molecular analysis of HEXB gene by sanger sequencing
Genomic DNA was extracted by the standard salting out method.11 The exonic and intronic flanking sequence of the HEXB gene was PCR amplified in 14 fragments using primers designed using Primer3 software (http://bioinfo.ut.ee/primer3-0.4.0/).12 DNA amplification was performed for each fragment in a 10 μl final volume containing 100 ng genomic DNA, 1 mm dNTPs, 10 pmol of each primer, 1 U Taq polymerase and 1 μl 10 × PCR buffer. Thirty cycles of amplification were used, each consisting of 1 min denaturation at 94 °C, 45 s annealing at 60–65 °C suitable for each exons and 45 s extension at 72 °C in a thermal cycler. Final extension time was 10 min. Negative control PCR tubes contained all of the above ingredients except DNA. PCR products were then electrophoresed in 2% agarose along with the appropriate negative controls and a 100 base-pair DNA ladder. Products that passed this quality check were purified by treatment with Exo-SAP-IT (USB Corporation, Cleveland, OH, USA) and then sequenced using BigDye Terminator v3.1 and capillary electrophoresis was performed using an automated sequencer ABI 3730XL (Applied Biosystems, Carlsbad, CA, USA). Mutations were described according to mutation nomenclature, considering nucleotide +1 the A of the first ATG translation initiation codon. Nucleotide numbers are derived from cDNA HEXB sequence (RefSeq cDNA NM_000521.3). Putative mutations were confirmed in two separate PCR products from the patient’s DNA. Heterozygosity for these mutations was confirmed in the parents. The mutations identified were then looked up in public databases like The Human Gene Mutation Database (http://www.hgmd.cf.ac.uk), dbSNP (http://www.ncbi.nlm.nih.gov/SNP/index.html) and McGill University database (http://www.hexdb.mcgill.ca). Novel variants were also ruled out as polymorphism by sequencing the corresponding exons/introns in 100 normal unrelated individuals.
In silico analysis
Prediction of functional effects of nonsynonymous single-nucleotide substitutions was done using softwares SIFT (Sorting Intolerant From Tolerant) (available at http://sift.jcvi.org/), Polyphen2 (Polymorphism Phenotyping v2) (available at http://genetics.bwh.harvard.edu/pph2/) and MutationT@ster (available at http://www.mutationtaster.org/).13, 14, 15 HumVar-trained prediction model of Polyphen2 was used for distinguishing mutations with drastic effects from all the remaining human variation, including abundant mildly deleterious alleles. Evolutionary conservation of the amino acid residues of Hex B was analyzed using ClustalW program available online at (http://www.uniprot.org/help/sequence-alignments).
Results
Twenty-two unrelated families with equal number of probands were included in the study. State wise representation was as follow: nine families from Maharashtra, four families from Karnataka, two families each from Andhra Pradesh, Gujarat, Rajasthan, Kerala and one family from Delhi. Consanguinity in parents was present in 9/22 (40.9%) families. The mean age at presentation was 17.3 months (±8.06). Most patients presented with neuroregression (20/22) and some with developmental delay (2/22) and bilateral cherry red spot was present in all. The mean age of onset of symptoms was 9.17±2.14 months; however, patients with developmental delay presented earlier (mean age of onset 7 months). Seizure was present in 17/22 cases (77.3%). Liver and/or spleen enlargement was present in 8/22 cases (36.4%). Neuroimaging in form of computed tomography or magnetic resonance imaging of the brain of the proband was available in 10/22 patients and showed changes such as cortical atrophy (n=10), hyperdense thalami (n=10) and dysmyelination (n=5).
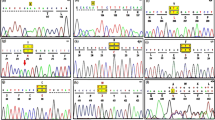
We confirmed 18/22 cases with SD (deficient activity of Hex-A and total-hex activity was observed in the leukocytes), with carrier detection (% Hex-A) in 4/22 parents where the proband was not alive but was earlier diagnosed as SD by enzyme study (Table 1). The mean total-hex value in an affected cases (n=18) was 176.69±101.36 nmol h−1 mg−1, while the mean Hex-A value was 55.54±27.76 nmol h−1 mg−1 (n=18). The mean Hex-A % value in carrier parents was 71.47% (±4.57). Molecular analysis was carried out in all affected patients with SD by bi-directional sequencing. Exon sequencing analysis revealed 13 mutations in 19 patients, while mutation was not identified in three patients. We have identified four missense mutation c.611G>A (p.G204E), c. 634A>T (p.H212Y), c.926G>A (p.C309Y), c.1597C>T (p.R533C), three nonsense mutations c.333G>A (p.W111X), c.298C>T (p.R100X), c.850C>T (p.R284X), two splice site mutation c.1082+5 G>T, c.1417+1 G>A, two deletion c.534_541delAGTTTATC (p.V179RfsX10), c.1563_1573delTATGGATGACG (p.M522LfsX2) and two insertion c.1591_1592insC (p.R531TfsX22), c.1553_1554insAAGA (p.D518EfsX8) in 19 patients. The recurrent intronic mutations c.1417+1 G>A was identified in two unrelated patients from Hyderabad and c.333 G>A (p.W111X) in two unrelated patients from Maharashtra suggesting possible founder effect. The mutation c.850 C>T (p.R284X) was also identified in 4/19 (21%) patients (two from Rajasthan and one each from Delhi and Kerala) suggesting them as a possible common mutation in the country. The clinical features, biochemical findings and results of mutation analysis are summarized in Table 1.
Discussion
To date, several clinically and biochemically diagnosed SD cases have been reported from India; however, the molecular characterization is lacking. Our study includes 19 cases of classical infantile Sandhoff disease variety from different geographic parts of India presented between 3 and 9 months of age.16
The unusual clinical features observed in our cases were the presence of mongolian spots on the back, legs of six unrelated probands. They are reported in association with Hurler syndrome, Hunter disease, Mucolipidosis-II and GM1 gangliosidosis.17 However, there is only one report of this association in Sandhoff disease.18 The Mongolian spots result from entrapment of melanocytes in the dermis because of arrested transdermal migration from the neural crest into the epidermis. It is likely to be due to enhancement of nerve growth factor activity from interaction of abnormal intracellular metabolites with tyrosine kinase type receptors.19
The distinguishing feature of SD versus TSD is presence of liver, spleen enlargement and cerebral white matter changes on neuroimaging in SD patients. Cerebral dysmyelination on neuroimaging was observed in five unrelated cases. Although, SD has been traditionally considered to be a gray matter disease, neuroimaging studies have characterized abnormalities within both the white and gray matter. Bilateral thalamic changes have been observed specially in GM2 gangliosidosis.20
In Sandhoff’s disease, total-hex activity is reduced to less than 2–3% of normal, unlike TSD, where total-hex activity is preserved because of functional Hex-B activity. However, the β isoenzyme is unable to hydrolyze the GM2 ganglioside, which accumulates in the brain and also within the liver, spleen, and kidneys, that may become enlarged.21 In two cases, the proband was referred to as having TSD on clinical referral elsewhere; however, our biochemical and molecular diagnosis confirmed the case as having SD.
The molecular profile was different in SD cases versus TSD cases from India (Mistri et al.). The majority of mutations in SD are indels whereas in TSD, missense mutations predominate. Similar to TSD, allelic heterogeneity was identified in SD cases. Present study also suggests that c.850 C>T (p.R284X) mutation was identified in 4/19 (21%) patients and it could be a common mutation in Indian population similar to what was observed in TSD for c.1385 A>T (E462V) in 21.42% of patients as founder mutation in six unrelated cases from Gujarat, c.964G>T (p. D322Y) in 10.7% and c.1277_1278insTATC in 14.28% of patients.2, 3
Allelic heterogeneity is reported in SD from different parts of the world. Till date, 86 mutations have been identified, including five large deletions (Human Gene Mutation Database, Cardiff, UK, 2014). In particular, the 16 kb deletion, accounting for 27% of the SD mutant alleles, has been reported as the most frequent mutation in patient affected from SD with different ethnic backgrounds.22, 23 However, multiplex ligase-dependent amplification assay was unavailable to us. The mutation p.R533C was found in present study with unknown second allele in one patient. It has been reported previously from Iran.24 Turkey and Saudi Arabia25 in classical infantile cases. Another mutation at 533 amino acid position viz. pR533H has been previously demonstrated in compound heterozygous state with IVS2-1G>A in an adult Sandhoff patient with motor neuron disease phenotype.26 The p.R284X mutation present in a CpG hotspot has been reported from different parts of the world. In the Italian series by Zampieri et al.27 the p.R284X mutation represented 29% of mutant alleles. The mutation (p.R284X) was also found in 4/20 (20%) patients in our study. The p.C309Y mutation has been identified before in compound heterozygous state in an adult onset Sandhoff disease patient.28 This mutation is predicted to alter processing of the pro-β chain. However in homozygous state this mutation leads to severe disease and in our study the patient presented around late infantile period. The following mutations identified in our study are novel mutations viz. two missense mutations [c.611G>A (p.G204E), c. 634 A>T (p.T212S)], two nonsense mutations [c.333G>A (p.W111X), c.298C>T (p.R100X)], one splice site mutation c.1082+5 G>T, two small in-frame deletions [c.534_541delAGTTTATC (p.V179RfsX10), c.1563_1573delTATGGATGACG (p.M522LfsX2)] and one insertion c.1553_1554insAAGA (p.D518EfsX8). Their mechanism of pathogenesis is nonsense-mediated decay of mRNA leading to no protein formation.
In conclusion, our results confirmed the remarkable heterogeneity of the mutational spectrum of the HEXB gene and provided new insight into molecular pathology of SD. In addition p.R284X mutation is common in Indian SD patients and can be used as a first line molecular screen in our patients. Though very few mutations occur in exons 1–5, due to lack of common mutations, complete gene sequencing will be required in most of the SD cases in India.
References
Sheth, J., Mistri, M., Sheth, F., Shah, R., Bavdekar, A., Godbole, K. et al. Burden of lysosomal storage disorders in India: experience of 387 affected children from a single diagnostic facility. JIMD Rep. 12, 51–63 (2013).
Mistri, M., Tamhankar, P. M., Sheth, F., Sanghavi, D., Kondurkar, P., Patil, S. et al. Identification of novel mutations in HEXA gene in children affected with Tay Sachs disease from India. PLoS ONE 7, e39122 (2012).
Sheth, J., Mistri, M., Datar, C., Kalane, U., Patil, S., Kamate, M. et al. Expanding the spectrum of HEXA mutations in Indian patients with Tay–Sachs disease. MGM Rep. 1, 425–430 (2014).
Seshadri, R., Christopher, R. & Arvinda, H. R. Teaching NeuroImages: MRI in infantile Sandhoff disease. Neurology 77, e34 (2011).
Jain, A., Kohli, A. & Sachan, D. Infantile Sandhoff disease with peripheral neuropathy. Pediatr. Neurol. 42, 459–461 (2010).
Nalini, A. & Christopher, R. Cerebral glycolipidoses: clinical characteristics of 41 pediatric patients. J. Child Neurol. 19, 447–452 (2004).
Unnikrishnan, A. G., Danda, S. & Seshadri, M. S. Juvenile Sandhoff Dease. Indian Pediatr. 38, 89–92 (2001).
Kaur, M. & Verma, I. C. Enzyme studies in GM2 gangliosidiosis, and their application in prenatal diagnosis. Ind. J. Pediatr. 62, 485–489 (1995).
Mittal, K., Mittal, A., Shelja, Raikwar, P. & Aggarwal, S. Infantile Sandhoff disease with hypothyroidism: a rare presentation. Int. J. Sci. Eng. Res. 5, 1455–1456 (2014).
Wendeler, M. & Sandhoff, K. Hexosaminidase assays. Glycoconj. J. 26, 945–952 (2009).
Miller, S. A., Dykes, D. D. & Polesky, H. F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Untergrasser, A., Cutcutache, I., Koressaar, T., Ye, J., Faircloth, B. C. & Remm, M. Primer3 - new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. A., Schmidt, S., Peshkin, L., Ramensky, V. E., Gerasimova, A. & Bork, P. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Schwarz, J. M., Rödelsperger, C., Schuelke, M. & Seelow, D. Mutation taster evaluates disease-causing potential of sequence alterations. Nat. Methods 7, 575–576 (2010).
Maegawa, G. H., Stockley, T., Tropak, M., Banwell, B., Blaser, S., Kok, F. et al. The natural history of juvenile or subacute GM2 gangliosidosis: 21 new cases and literature review of 134 previously reported. Pediatrics 118, e1550–e1562 (2006).
Luna, P. C., Abad, M. E., Boggio, P., Figueredo, A. M., Salgo, M., Schenone, A. et al. Lysosomal storage diseases, diagnosis from skin lesions. Dermatol. Argent. 17, 221–229 (2011).
Ashrafi, M. R., Tavasoli, A., Shiva, S., Parvaneh, N. & Tamizifar, B. Diffuse dermal melanocytosis in two patients with Sandhoff disease and mucopolysaccharidosis VI. Int. J. Dermatol. 53, 736–738 (2014).
Hackbart, B. A., Arita, J. H., Pinho, R. S., Masruha, M. R. & Vilanova, L. C. Mongolian spots are not always a benign sign. J. Pediatr. 162, 1070 (2013).
Sass, K., Wiebe, S. & Lemire, E. Infantile Sandhoff disease. SA J. Radiol. 13, 66 (2009).
Brismar, J., Brismar, G., Coates, R., Gascon, G. & Ozand, P. Increased density of the thalamus on CT scans in patients with GM2 gangliosidoses. Am. J. Neuroradiol. 11, 125–130 (1990).
Bolhuis, P. A. & Bikker, H. Deletion of the 5’-region in one or two alleles of HEXB in 15 out of 30 patients with Sandhoff disease. Hum. Genet. 90, 328–329 (1992).
Zampieri, S., Cattarossi, S., Oller Ramirez, A. M., Rosano, C., Lourenco, C. M., Passon, N. et al. Sequence and copy number analyses of HEXB gene in patients affected by Sandhoff disease: functional characterization of 9 novel sequence variants. PLoS ONE 7, e41516 (2012).
Aryan, H., Aryani, O., Banihashemi, K., Zaman, T. & Houshmand, M. Novel mutations in Sandhoff disease: a molecular analysis among Iranian cohort of infantile patients. Iran J. Public Health 41, 112–118 (2012).
Kaya, N., Al-Owain, M., Abudheim, N., Al-Zahrani, J., Colak, D., Al-Sayed, M. et al. GM2 gangliosidosis in Saudi Arabia: multiple mutations and considerations for future carrier screening. Am. J. Med. Genet A 155, 1281–1284 (2011).
Yoshizawa, T., Kohno, Y., Nissato, S. & Shoji, S. Compound heterozygosity with two novel mutations in the HEXB gene produces adult Sandhoff disease presenting as a motor neuron disease phenotype. J. Neurol. Sci. 195, 129–138 (2002).
Zampieri, S., Filocamo, M., Buratti, E., Stroppiano, M., Vlahovicek, K., Rosso, N. et al. Molecular and functional analysis of the HEXB gene in Italian patients affected with Sandhoff disease: identification of six novel alleles. Neurogenetics 10, 49–58 (2009).
Gomez-Lira, M., Sangalli, A., Mottes, M., Perusi, C., Pignatti, P. F., Rizzuto, N. et al. A common beta hexosaminidase gene mutation in adult Sandhoff disease patients. Hum. Genet. 96, 417–422 (1995).
Acknowledgements
Ethical consideration and the state of informed consent: all procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation (institutional and national) and with the Helsinki Declaration of 1975, as revised in 2000. Informed consent was obtained from all patients for being included in the study.
Author contributions
PT, JS and MM designed the experiment and standardized the protocols. MM, PK and DS were involved in processing of the samples. PT, JS and MM prepared the manuscript. All the authors read and approved the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tamhankar, P., Mistri, M., Kondurkar, P. et al. Clinical, biochemical and mutation profile in Indian patients with Sandhoff disease. J Hum Genet 61, 163–166 (2016). https://doi.org/10.1038/jhg.2015.130
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.130