Abstract
Many single-nucleotide polymorphisms (SNPs) have been reported to be associated with lipid concentrations in recent genome-wide association studies. The aim of this study was to validate the associations of rs2197089 in the lipoprotein lipase (LPL) gene with serum lipid concentrations and gene expression levels in the Chinese Han population and examine the potential interactions. A total of 9339 participants were recruited and genotyped for rs2197089. Gene expression levels of LPL in blood cells of 309 participants were evaluated by real-time PCR. We observed significant associations between rs2197089 and decreased triglycerides (TG) (P=0.0006), but not high-density lipoprotein cholesterol (HDL-C) concentration (P=0.0881). However, weak evidence of interaction between cigarette smoking and rs2197089 was detected (P=0.0362). In smokers, significant association between rs2197089 and increased HDL-C concentration was found (P=0.0068). Participants with the minor allele A had higher expression levels of LPL (P=0.0243). The results of our study indicated that rs2197089 was significantly associated with TG but it was associated with HDL-C only in smokers. This SNP seemed to have influence on the expression level of LPL.
Similar content being viewed by others
Introduction
Lipoprotein lipase (LPL) is a key enzyme in the metabolism of lipoproteins. It hydrolyzes plasma lipoprotein triglycerides (TG) into free fatty acids and glycerol, converts very-low-density lipoprotein to low-density lipoprotein cholesterol.1 The LPL gene is located at 8p22 and comprises 10 exons spanning about 30 kb.2, 3 Serum lipid concentrations are affected by both lifestyle factors, such as diet, obesity, physical activity, and genetic factors.4 Several functional polymorphisms in LPL have been studied with regard to their associations with lipid profile and risk of coronary heart disease in candidate gene association studies.5, 6
In recent years, many single-nucleotide polymorphisms (SNPs) in or near LPL associated with plasma levels of high-density lipoprotein cholesterol (HDL-C) and TG have been reported in several genome-wide association studies (GWAS)7, 8, 9, 10, 11, 12 and most of them are in high linkage disequilibrium. The SNP rs2197089, in the 3′ downstream of the LPL gene, has been reported to be associated with HDL-C and TG.8, 9, 12 Some later studies successfully replicate the associations in different ethnicities, such as African American, American Indian, Mexican American and Hispanic populations.13, 14, 15 However, not all the GWAS-identified associations can be replicated in diverse ethnic groups. For example, the association between rs2197089 and TG failed to generalize in African Americans.14
It is suggested that the true differences in the associations between the genetic polymorphism and serum lipid concentrations between ethnic groups may result from differing prevalence of other factors involved in gene–gene and gene–environment interactions.16 In support of this hypothesis, significant interactions have been observed between LPL gene polymorphisms and environmental factors, such as smoking,17, 18, 19, 20, 21 gender,17, 22 alcohol consumption,20 dietary,23, 24, 25 physical activity18, 26 and obesity.24, 27, 28, 29, 30, 31, 32 However, results from other studies did not support some of these findings for interaction between LPL SNPs and smoking,24, 33, 34 gender,24 alcohol consumption,24, 33 dietary,35, 36 physical activity20, 25 and obesity.20, 37, 38
The association between rs2197089 and lipid concentrations was less reported in the Chinese Han population, and so was the interaction of this SNP with risk factors on lipid concentrations. The aim of this study was to validate the associations between this SNP and serum lipid concentrations in the Chinese Han population and examine the potential effect modification of these associations by interactions with cigarette smoking, alcohol consumption, hypertension, gender and obesity. Furthermore, the relationship between rs2197089 and gene expression levels of LPL was also evaluated.
Materials and methods
Study population
A group of 9339 participants were randomly selected from the International Collaborative Study of Cardiovascular Disease in Asia (InterASIA in China)39 and judged to be free of coronary heart disease, stroke, renal or hepatic diseases, type 2 diabetes and cancer by medical history. Standard questionnaire were used by trained interviewers to obtain information on demographic characteristics including age, gender, ethnicity, details of medical history, smoking and alcohol consumption. Blood pressure, weight and height were recorded. This study was approved by the local bioethics committee, and all subjects gave written informed consent.
Biochemical measurements
Overnight fasting blood samples were drawn by venipuncture to measure serum biochemical measurements including total cholesterol, HDL-C, TG and glucose. Blood specimens were processed in the central clinical laboratory at the Department of Population Genetics at Fuwai Hospital of the Chinese Academy of Medical Sciences in Beijing. This laboratory participates in the Lipid Standardization Program of the US Centers for Disease Control and Prevention. Total cholesterol, HDL-C, TG and glucose were analyzed enzymatically on a Hitachi 7060 Clinical Analyzer (Hitachi High-Technologies Corp., Tokyo, Japan). The low-density lipoprotein cholesterol concentrations were calculated by use of the Friedewald equation.
DNA extraction and genotyping
Genomic DNA was isolated from white blood cells according to a standard procedure using a DNA extraction kit (Tiangen Biotech, Beijing, China). SNP was genotyped using genomic DNA with the fluorogenic 5′-nuclease TaqMan allelic discrimination assay system (Applied Biosystems, Foster City, CA, USA). The assays were performed under standard conditions on a 7900 HT Fast Real-Time PCR instrument (Applied Biosystems). Successful genotyping rate was over 98%. For quality control, 2% of the samples were randomly selected and genotyped in duplicate. A comparison of the duplicate results showed that the discrepancy was not >1%.
RNA extraction and real-time PCR
To analyze the mRNA level of the LPL, we separated peripheral blood mononuclear cells from fresh blood samples of 309 healthy individuals. Total RNA was extracted with TRIzol (Invitrogen, Carlsbad, CA, USA). After quantification, 1 μg of total RNA was reverse transcribed using Moloney Murine Leukemia Virus Reverse Transcriptase (Tiangen Biotech) with oligo dT according to the manufacturer's instructions.
Real-time PCR was performed using the Applied Biosystems 7900 HT Fast Real-Time PCR System. Expression of all assays was measured in triplicates and average values of the triplicates were used for the analysis. Gene expression was quantified using the comparative Ct method, which standardized the Ct values to an internal housekeeping gene (ACTB) and calculated the relative expression values (ΔCt method).
Statistical analysis
Statistical power was calculated by using Quanto software (http://hydra.usc.edu/gxe/). Statistical analyses were performed with the SAS (Version 9.0, SAS Institute, Cary, NC, USA) software package. The distributions of HDL-C and TG were evaluated by skewness and kurtosis statistics. TG values were log-transformed to approximate a normal distribution, and the log-transformed values were used in all statistical tests. Deviation from Hardy–Weinberg equilibrium was tested by χ2-test. The SNP was analyzed as 0, 1 or 2 copies of the minor allele in an additive genetic model. Linear regression was used to test for association between the number of minor alleles and the concentrations of HDL-C and TG. All models were adjusted for age, gender, hypertension, body mass index and smoke status. To detect the potential gene–environment interactions, an extra interaction term about the genotype and variables for exposure was included in the regression model. The trend of the mean LPL expression levels in different genotype groups was tested using simple linear regression test. In all analysis a P-value<0.05 was considered statistically significant. Multiple testing was adjusted using Bonferroni correction.
Results
Association with lipid concentrations
Table 1 showed the general characteristics of the participants included in this study. We investigated the associations between rs2197089 and lipid concentrations in the 9339 study samples. Genotype frequency for rs2197089 and the results of the stepwise linear regression analyses were shown in Table 2. The minor allele frequency of rs2197089 was 33.8% in our samples. No significant deviation from Hardy–Weinberg equilibrium was observed for this polymorphism (P=0.2608). We have 96% power to detect the association between rs2197089 and lipid concentration under the additive model with a main effect of 0.02 in our study. Under an additive model, we observed significant associations between rs2197089 and decreased TG (Effect=−0.028 (±0.008), P=0.0006), but not HDL-C concentration (Effect=0.009 (±0.005), P=0.0881).
However, weak evidence of interaction between cigarette smoking and rs2197089 was detected, as the corresponding product term in the regression model had significant contribution for HDL-C (P=0.0362). The interaction did not reach a significant level of 0.05 if we considered the multiple testing (two times of evaluations for interactions).
We performed subgroup analyses by separating the study population into smokers and nonsmokers (Table 2). In smokers, we observed significant association between rs2197089 and increased HDL-C concentration (Effect=0.023 (±0.008), P=0.0068), even after considering multiple testing (six times for testing association of rs2197089 with HDL-C and TG in all, smokers and nonsmokers). In nonsmokers, the association was not significant. The effect of rs2197089 on TG concentration seemed not to be modified by smoking, as minor allele A carriers had lower TG concentration than GG homozygotes in both smokers and nonsmokers.
We attempted to find additional interactions between rs2197089 and other factors, such as alcohol consumption, hypertension, obesity and gender, but no significant result was found, neither for HDL-C nor TG.
LPL gene expression
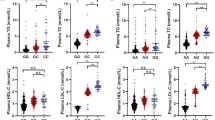
Figure 1 showed the result of gene expression analysis. The standardized expression level of the minor allele A carriers were higher than that of the major allele homozygotes. The expression levels significantly increased as the copy of the A allele increased (P-value for linear trend test was 0.0243). The AA homozygotes and AG heterozygotes had 1.41 and 1.22-fold higher expression levels than the GG homozygotes, respectively.
Expression levels of LPL gene in different genotypes of rs2197089. The standardized expression levels of the minor allele A carriers were higher than that of the major allele homozygotes. P-value for linear trend test was 0.0243. The AA homozygotes and AG heterozygotes had 1.41 and 1.22-fold higher expression levels than the GG homozygotes, respectively. BMI, body mass index; DBP, diastolic blood pressure; GLU, glucose; HWE, Hardy–Weinberg Equilibrium; LD, linkage disequilibrium; LDL-C, low-density lipoprotein cholesterol; SBP, systolic blood pressure.
Discussion
In the present study, we first evaluated the associations between rs2197089 and lipid concentrations in a group of 9339 samples of the Chinese Han population. We found that rs2197089 was associated with decreased TG concentration, but not HDL-C. However, significant association between rs2197089 and increased HDL-C concentration was observed in smokers. The expression levels of LPL gene in different genotype groups seemed to be discrepant. Participants with the minor allele A had higher expression levels of LPL.
Three GWASs reported the associations between rs2197089 and HDL-C and TG concentrations.8, 9, 12 However, some studies showed that not all the GWAS-identified associations can be replicated. The Population Architecture using Genomics and Epidemiology study was established to determine whether GWAS-identified variants generalized to diverse groups drawn from population-based studies.40 Forty-nine GWAS-identified SNPs, which were associated with one or more lipid traits, were genotyped in a recent study and only 16 associations could be generalized across African American, American Indian, Mexican American and Hispanic populations. The association between rs2197089 and HDL-C was successfully generalized across the four populations, but the association between rs2197089 and TG failed to generalize in African Americans.14
There was quite a difference for the frequency of rs2197089 between different populations. The A allele frequency was 55% in Europeans, and much higher in African Americans (78%), but the frequency in East Asian, American Indian, Mexican American and Hispanic populations was <50%. It was 33.8% in our study. The linkage disequilibrium structure was slightly different between CEU (Utah Residents with Northern and Western European Ancestry) and CHB (Han Chinese in Beijing) in HapMap data as well. Allele frequency and linkage disequilibrium structure differences might be explanations for non-replication for the association between rs2197089 and HDL-C concentrations in our samples.
The associations between genotypes and phenotypes can be modified by gene–gene and gene–environment interactions. Effect modification of the associations between LPL SNPs, rs263 in intron,5, 25 HindIII (rs320) in intron8, 17, 18 and S447X (rs328) in exon,9, 17, 20, 21 and serum lipid concentrations by interactions with cigarette smoking has been reported. The minor allele of rs263 was reported to be associated with higher HDL-C and lower TG levels through an interaction with smoking, and the effect was stronger in smokers than in nonsmokers.25 For the HindIII polymorphism, Peacock et al.17 found that smokers with the H+ allele had higher levels of TG than those with one or more H- alleles. This effect was larger and significant only in smoking women and was marginally significant in smoking men, but effects were smaller and opposite in the nonsmokers. Sentí et al.18 reported that in smokers, sedentary H+H+ homozygotes showed significantly higher TG and lower HDL-C concentrations than sedentary H– carriers. No statistically significant differences were observed in lipid levels of active or sedentary nonsmokers between H– carriers and H+H+ homozygotes. For S447X, the association of the X447 allele with higher HDL-C was reported to be significant only in non-smoking women but not in smoking women by Komurcu-Bayrak et al.21 However, Lee et al.20 showed that in both men and women, X447 allele carriers had higher HDL-C concentration, but the effect was greater in smokers than in nonsmokers. However, some studies found that the interaction of rs263 Andreott et al.24 and rs328 Talmnd et al.33 and Skoglund-Anderson et al.34 with smoking was not significant. Weak evidence of interaction between rs2197089 and smoking on HDL-C concentration, which has not been reported in previous studies, was detected in our study. SNP rs2197089 might influence HDL-C concentration by interacting with smoking. In smokers, rs2197089 was detected to be significantly associated with HDL-C concentration. HDL-C concentration increased as the copy of the minor allele A increased. But in nonsmokers, AA homozygotes did not have higher HDL-C concentration. Then, another explanation for non-replication for the association between rs2197089 and HDL-C concentration in Chinese Han population might be gene–environment interactions. Smoking was reported to be related with reduced LPL activity,41 but the precise mechanisms by which the interaction of LPL polymorphisms with smoking affects lipid concentrations were still unknown.
As most GWAS-identified SNPs were present in non-coding regions, a functional SNP would be most likely affecting gene regulation by altering the binding of some transcription factors. Smith et al.42 carried out a study to validate the candidate functional SNPs in LPL. The electrophoretic mobility-shift assay analysis revealed potential binding differences between the wild-type and variant alleles of rs2197089 using the Huh7 cell line. We thought that rs2197089 might have some functional effects on gene expression. Our expression study showed that the expression levels of LPL gene in different genotypes seemed to be different. Participants with the minor allele A had higher expression levels of LPL. Overall, the previous in vitro study and our expression study suggested that rs2197089 might influence the expression of the LPL gene by affecting the binding of some transcription factors, and then influenced the lipid concentrations.
In summary, the results of our study demonstrated that rs2197089 was associated with TG concentration. The effect of this SNP on HDL-C concentration might be modified by smoking. The SNP genotypes seemed to have influence on the expression levels of LPL. In this paper, however, only one SNP in the 3'-end region of LPL was reported, other polymorphisms in and around this region should also be investigated in the future and further studies are needed to elucidate the underlying mechanisms.
References
Goldberg, I. J. Lipoprotein lipase and lipolysis: central roles in lipoprotein metabolism and atherogenesis. J. Lipid Res. 37, 693–707 (1996).
Sparkes, R. S., Zollman, S., Klisak, I., Kirchgessner, T. G., Komaromy, M. C., Mohandas, T. et al. Human genes involved in lipolysis of plasma lipoproteins: mapping of loci for lipoprotein lipase to 8p22 and hepatic lipase to 15q21. Genomics 1, 138–144 (1987).
Monsalve, M. V., Henderson, H., Roederer, G., Julien, P., Deeb, S., Kastelein, J. J. et al. A missense mutation at codon 188 of the human lipoprotein lipase gene is a frequent cause of lipoprotein lipase deficiency in persons of different ancestries. J. Clin. Invest. 86, 728–734 (1990).
Heller, D. A., de Faire, U., Pedersen, N. L., Dahlen, G. & McClearn, G. E. Genetic and environmental influences on serum lipid levels in twins. N. Engl. J. Med. 328, 1150–1156 (1993).
Wittrup, H. H., Andersen, R. V., Tybjaerg-Hansen, A., Jensen, G. B. & Nordestgaard, B. G. Combined analysis of six lipoprotein lipase genetic variants on triglycerides, high-density lipoprotein, and ischemic heart disease: cross-sectional, prospective, and case-control studies from the Copenhagen City Heart Study. J. Clin. Endocrinol. Metab. 91, 1438–1445 (2006).
Sagoo, G. S., Tatt, I., Salanti, G., Butterworth, A. S., Sarwar, N., van Maarle, M. et al. Seven lipoprotein lipase gene polymorphisms, lipid fractions, and coronary disease: a HuGE association review and meta-analysis. Am. J. Epidemiol. 168, 1233–1246 (2008).
Aulchenko, Y. S., Ripatti, S., Lindqvist, I., Boomsma, D., Heid, I. M., Pramstaller, P. P. et al. Loci influencing lipid levels and coronary heart disease risk in 16 European population cohorts. Nat. Genet. 41, 47–55 (2009).
Kathiresan, S., Melander, O., Guiducci, C., Surti, A., Burtt, N. P., Rieder, M. J. et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nat. Genet. 40, 189–197 (2008).
Kooner, J. S., Chambers, J. C., Aguilar-Salinas, C. A., Hinds, D. A., Hyde, C. L., Warnes, G. R. et al. Genome-wide scan identifies variation in MLXIPL associated with plasma triglycerides. Nat. Genet. 40, 149–151 (2008).
Sabatti, C., Service, S. K., Hartikainen, A. L., Pouta, A., Ripatti, S., Brodsky, J. et al. Genome-wide association analysis of metabolic traits in a birth cohort from a founder population. Nat. Genet. 41, 35–46 (2009).
Teslovich, T. M., Musunuru, K., Smith, A. V., Edmondson, A. C., Stylianou, I. M., Koseki, M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Willer, C. J., Sanna, S., Jackson, A. U., Scuteri, A., Bonnycastle, L. L., Clarke, R. et al. Newly identified loci that influence lipid concentrations and risk of coronary artery disease. Nat. Genet. 40, 161–169 (2008).
Deo, R. C., Reich, D., Tandon, A., Akylbekova, E., Patterson, N., Waliszewska, A. et al. Genetic differences between the determinants of lipid profile phenotypes in African and European Americans: the Jackson Heart Study. PLoS Genet. 5, e1000342 (2009).
Dumitrescu, L., Carty, C. L., Taylor, K., Schumacher, F. R., Hindorff, L. A., Ambite, J. L. et al. Genetic determinants of lipid traits in diverse populations from the population architecture using genomics and epidemiology (PAGE) study. PLoS Genet. 7, e1002138 (2011).
Rotger, M., Bayard, C., Taffe, P., Martinez, R., Cavassini, M., Bernasconi, E. et al. Contribution of genome-wide significant single-nucleotide polymorphisms and antiretroviral therapy to dyslipidemia in HIV-infected individuals: a longitudinal study. Circ. Cardiovasc. Genet. 2, 621–628 (2009).
Ordovas, J. M. Cardiovascular disease genetics: a long and winding road. Curr. Opin. Lipidol. 14, 47–54 (2003).
Peacock, R. E., Temple, A., Gudnason, V., Rosseneu, M. & Humphries, S. E. Variation at the lipoprotein lipase and apolipoprotein AI-CIII gene loci are associated with fasting lipid and lipoprotein traits in a population sample from Iceland: interaction between genotype, gender, and smoking status. Genet. Epidemiol. 14, 265–282 (1997).
Senti, M., Elosua, R., Tomas, M., Sala, J., Masia, R., Ordovas, J. M. et al. Physical activity modulates the combined effect of a common variant of the lipoprotein lipase gene and smoking on serum triglyceride levels and high-density lipoprotein cholesterol in men. Hum. Genet. 109, 385–392 (2001).
Corella, D., Guillen, M., Saiz, C., Portoles, O., Sabater, A., Folch, J. et al. Associations of LPL and APOC3 gene polymorphisms on plasma lipids in a Mediterranean population: interaction with tobacco smoking and the APOE locus. J. Lipid Res. 43, 416–427 (2002).
Lee, J., Tan, C. S., Chia, K. S., Tan, C. E., Chew, S. K., Ordovas, J. M. et al. The lipoprotein lipase S447X polymorphism and plasma lipids: interactions with APOE polymorphisms, smoking, and alcohol consumption. J. Lipid Res. 45, 1132–1139 (2004).
Komurcu-Bayrak, E., Onat, A., Poda, M., Humphries, S. E., Acharya, J., Hergenc, G. et al. The S447X variant of lipoprotein lipase gene is associated with metabolic syndrome and lipid levels among Turks. Clin. Chim. Acta 383, 110–115 (2007).
Hegele, R. A., Evans, A. J., Tu, L., Ip, G., Brunt, J. H. & Connelly, P. W. A gene-gender interaction affecting plasma lipoproteins in a genetic isolate. Arterioscler. Thromb. 14, 671–678 (1994).
Nettleton, J. A., Steffen, L. M., Ballantyne, C. M., Boerwinkle, E. & Folsom, A. R. Associations between HDL-cholesterol and polymorphisms in hepatic lipase and lipoprotein lipase genes are modified by dietary fat intake in African American and White adults. Atherosclerosis 194, e131-140 (2007).
Andreotti, G., Menashe, I., Chen, J., Chang, S. C., Rashid, A., Gao, Y. T. et al. Genetic determinants of serum lipid levels in Chinese subjects: a population-based study in Shanghai, China. Eur. J. Epidemiol. 24, 763–774 (2009).
Pyun, J. A., Kim, S., Park, K., Baik, I., Cho, N. H., Koh, I. et al. Interaction effects of lipoprotein lipase polymorphisms with lifestyle on lipid levels in a Korean population: a cross-sectional study. Genomics Inform. 10, 88–98 (2012).
Boer, J. M., Kuivenhoven, J. A., Feskens, E. J., Schouten, E. G., Havekes, L. M., Seidell, J. C. et al. Physical activity modulates the effect of a lipoprotein lipase mutation (D9N) on plasma lipids and lipoproteins. Clin. Genet. 56, 158–163 (1999).
Fisher, R. M., Mailly, F., Peacock, R. E., Hamsten, A., Seed, M., Yudkin, J. S. et al. Interaction of the lipoprotein lipase asparagine 291-->serine mutation with body mass index determines elevated plasma triacylglycerol concentrations: a study in hyperlipidemic subjects, myocardial infarction survivors, and healthy adults. J. Lipid Res. 36, 2104–2112 (1995).
Gerdes, C., Fisher, R. M., Nicaud, V., Boer, J., Humphries, S. E., Talmud, P. J. et al. Lipoprotein lipase variants D9N and N291S are associated with increased plasma triglyceride and lower high-density lipoprotein cholesterol concentrations: studies in the fasting and postprandial states: the European Atherosclerosis Research Studies. Circulation 96, 733–740 (1997).
Wittekoek, M. E., Pimstone, S. N., Reymer, P. W., Feuth, L., Botma, G. J., Defesche, J. C. et al. A common mutation in the lipoprotein lipase gene (N291S) alters the lipoprotein phenotype and risk for cardiovascular disease in patients with familial hypercholesterolemia. Circulation 97, 729–735 (1998).
Garenc, C., Perusse, L., Gagnon, J., Chagnon, Y. C., Bergeron, J., Despres, J. P. et al. Linkage and association studies of the lipoprotein lipase gene with postheparin plasma lipase activities, body fat, and plasma lipid and lipoprotein concentrations: the HERITAGE Family Study. Metabolism 49, 432–439 (2000).
Senti, M., Bosch, M., Aubo, C., Elosua, R., Masia, R. & Marrugat, J. Relationship of abdominal adiposity and dyslipemic status in women with a common mutation in the lipoprotein lipase gene. The REGICOR investigators. Atherosclerosis 150, 135–141 (2000).
Huang, A. Q., Hu, Y. H., Zhan, S. Y., Xu, B., Pang, Z. C., Cao, W. H. et al. Lipoprotein lipase gene S447X polymorphism modulates the relation between central obesity and serum lipids, a twin study. Int. J. Obes. 30, 1693–1701 (2006).
Talmud, P. J., Hawe, E., Robertson, K., Miller, G. J., Miller, N. E. & Humphries, S. E. Genetic and environmental determinants of plasma high density lipoprotein cholesterol and apolipoprotein AI concentrations in healthy middle-aged men. Ann. Hum. Genet. 66, 111–124 (2002).
Skoglund-Andersson, C., Ehrenborg, E., Fisher, R. M., Olivecrona, G., Hamsten, A. & Karpe, F. Influence of common variants in the CETP, LPL, HL and APO E genes on LDL heterogeneity in healthy, middle-aged men. Atherosclerosis 167, 311–317 (2003).
Friedlander, Y., Leitersdorf, E., Vecsler, R., Funke, H. & Kark, J. The contribution of candidate genes to the response of plasma lipids and lipoproteins to dietary challenge. Atherosclerosis 152, 239–248 (2000).
Thu, N. N., Mai, T. T., Ohmori, R., Kuroki, M., Chuyen, N. V., Hung, N. T. et al. Plasma triglyceride and HDL-cholesterol concentrations in Vietnamese girls are affected by lipoprotein lipase, but not apolipoprotein CIII polymorphism. J. Nutr. 136, 1488–1492 (2006).
Kastelein, J. J., Ordovas, J. M., Wittekoek, M. E., Pimstone, S. N., Wilson, W. F., Gagne, S. E. et al. Two common mutations (D9N, N291S) in lipoprotein lipase: a cumulative analysis of their influence on plasma lipids and lipoproteins in men and women. Clin. Genet. 56, 297–305 (1999).
Arca, M., Campagna, F., Montali, A., Barilla, F., Mangieri, E., Tanzilli, G. et al. The common mutations in the lipoprotein lipase gene in Italy: effects on plasma lipids and angiographically assessed coronary atherosclerosis. Clin. Genet. 58, 369–374 (2000).
Gu, D., Reynolds, K., Wu, X., Chen, J., Duan, X., Reynolds, R. F. et al. Prevalence of the metabolic syndrome and overweight among adults in China. Lancet. 365, 1398–1405 (2005).
Matise, T. C., Ambite, J. L., Buyske, S., Carlson, C. S., Cole, S. A., Crawford, D. C. et al. The Next PAGE in understanding complex traits: design for the analysis of Population Architecture Using Genetics and Epidemiology (PAGE) Study. Am. J. Epidemiol. 174, 849–859 (2011).
Freeman, D. J., Caslake, M. J., Griffin, B. A., Hinnie, J., Tan, C. E., Watson, T. D. et al. The effect of smoking on post-heparin lipoprotein and hepatic lipase, cholesteryl ester transfer protein and lecithin:cholesterol acyl transferase activities in human plasma. Eur. J. Clin. Invest. 28, 584–591 (1998).
Smith, A. J., Palmen, J., Putt, W., Talmud, P. J., Humphries, S. E. & Drenos, F. Application of statistical and functional methodologies for the investigation of genetic determinants of coronary heart disease biomarkers: lipoprotein lipase genotype and plasma triglycerides as an exemplar. Hum. Mol. Genet. 19, 3936–3947 (2010).
Acknowledgements
We thank Ying Li, Liancheng Zhao, Donghua Liu, Jingping Chen, Fanghong Lu, Zhendong Liu, Cailiang Yao, Chong Shen, Xiaodong Pu, Lin Yu, Xianping Wu, Dongsheng Hu, Dongshuang Guo, Chenlong Dou, Qingjie Meng, Jun Yang, Zili Yang and the many people who helped with the field and laboratory work. This work was supported by National Basic Research Program of China (973 Plan) (grant no. 2011CB503901 and 2012CB517501), National Natural Science Foundation of China (grant no. 30930047).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mo, X., Liu, X., Wang, L. et al. Association of lipoprotein lipase polymorphism rs2197089 with serum lipid concentrations and LPL gene expression. J Hum Genet 58, 160–164 (2013). https://doi.org/10.1038/jhg.2012.151
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.151
Keywords
This article is cited by
-
Common variants in the genes of triglyceride and HDL-C metabolism lack association with coronary artery disease in the Pakistani subjects
Lipids in Health and Disease (2017)
-
Effect of obesity on the association between MYL2 (rs3782889) and high-density lipoprotein cholesterol among Korean men
Journal of Human Genetics (2016)