Abstract
Phenylketonuria (PKU) is a heterogeneous metabolic disorder caused by a deficiency in hepatic phenylalanine hydroxylase (PAH). On the basis of phenotype/genotype correlations, determination of phenylketonuric genotype is important for classification of the clinical phenotype and treatment of PKU, including tetrahydrobiopterin therapy. We characterized the genotypes of 203 Japanese patients with PKU and hyperphenylalaninemia using the following systems: (1) denaturing high-performance liquid chromatography with a GC-clamped primer; (2) direct sequencing; and, (3) multiplex ligation-dependent probe amplification. Of 406 mutant alleles, 390 (96%) were genotyped; 65 mutations were identified, including 22 new mutations. R413P, R241C, IVS4-1g>a, R111X and R243Q were prevalent mutations. Mutations prevalent in the Japanese cohort are also common in Korean and Northern Chinese populations, suggesting same origin. The spectrum of prevalent mutations was not significantly different among six Japanese districts, indicating that Japan comprises a relatively homogeneous ethnic group. We classified the mutations by clinical phenotypes and in vivo PAH activity and estimated the mutations with potential tetrahydrobiopterin (BH4) responsiveness. The frequency of BH4 responsiveness based on the genotype was 29.1% in Japanese PKU patients. A catalog of PKU genotypes would be useful for predicting clinical phenotype, deciding on the subsequent treatment of PKU including BH4 therapy, and genetic counseling in East Asia.
Similar content being viewed by others
Introduction
Phenylketonuria (PKU) is an autosomal recessive disorder caused by a deficiency in hepatic phenylalanine hydroxylase (PAH; EC 1. 14.16.1). The disease causes mental retardation unless the affected child is maintained on a strict low-phenylalanine diet.1 Mass screening of newborns for PKU is performed in developed countries and patients show a wide spectrum of clinical severity. The incidence of PKU in Japan is 1/120 000,2 which is lower than in Caucasians (1/10 000),3 Chinese (1/18 000)4 and Korean (1/41 000)5 populations. The diagnosis of PKU is based on high phenylalanine levels and absence of tetrahydrobiopterin (BH4) deficiency, rather than on hepatic PAH activity. Phenylalanine levels also largely determine the clinical phenotype of PKU, although blood phenylalanine is influenced by dietary protein intake.
Studying the molecular genetics of PKU has proven clinically useful. PKU is a heterogeneous metabolic disorder at the genetic as well as clinical level, and more than 500 different mutations of the PAH gene have been recorded in the PAH database (PAHdb; http://www.PAHdb.mcgill.ca). PKU mutations in East Asian populations are relatively similar with each other and completely different from those in Caucasians.6, 7, 8, 9
Correlations between the genotype and clinical phenotype of PKU have been demonstrated using predicted PAH activity, which is the average in vitro PAH activity for both mutations,10 with several mutations related to non-classical PKU phenotypes (mild PKU and mild hyperphenylalaninemia (HPA)).7, 11, 12, 13
Recently, BH4-responsive PAH deficiency was defined by decreased blood phenylalanine after a BH4 loading test14 and patients with this deficiency were treated with long-term BH4.15, 16, 17, 18, 19, 20 Blood phenylalanine levels were relatively well controlled by BH4 therapy compared with dietary treatment involving improvement of patients’ quality of life. Such patients usually have mild PKU or mild HPA, but not all patients with mild PKU and HPA respond to BH4. BH4-responsive PAH deficiency is difficult at times to diagnose with the decrease of phenylalanine level by BH4 loading test, as BH4 loading test is affected by the factors such as the amount and duration of administered BH4, preload plasma phenylalanine levels and phenylalanine intake. Therefore, identification of genotype responsive to BH4 therapy can help in the clinical diagnosis and/or treatment of PKU patients.21
The genetic analysis of PKU involves screening the affected exons extensively by the following methods to precisely map the mutations: (1) single strand conformation polymorphism,22 (2) denaturing gradient-gel electrophoresis,23 (3) denaturing high-performance liquid chromatography (DHPLC)24, 25 and (4) thermal melt profiling.26 DHPLC detects mutations with high sensitivity and precision and at low cost compared with denaturing gradient-gel electrophoresis. However, it is sometimes difficult to achieve suitable peaks by the position of a mutation in the amplified DNA as the column is highly sensitive to temperature conditions. We developed and improved the standard DHPLC method by using GC-clamped primers, and in doing so we could identify the exon mutations more easily and stably.
Exon screens that do not identify a mutation in the coding region suggest a large deletion in exon 5 to 6 (E5_6 del), as reported in Japanese PKU patients tested by Southern hybridization.27, 28 This E5_6 del is also easily detected using multiplex ligation-dependent probe amplification (MLPA).29, 30
The present study analyzed the genetic data of 203 Japanese residents with PKU. We classified the mutations by clinical phenotype and estimated those with potential BH4-responsiveness. This mutation catalog could aid clinicians in East Asia significantly in predicting clinical phenotype of PKU, in planning subsequent treatments including BH4 therapy and in genetic counseling.
Materials and methods
Subjects
This study enrolled 203 Japanese residents with a serum phenylalanine level higher than 0.18 mM. PAH deficiency was diagnosed at the participating institutions on the basis of absence of neurologic deterioration on a low phenylalanine diet, analysis of dihydropteridine reductase activity in red blood cells, biopterin loading test and/or pteridine analysis in urine. The criterion for classical PKU is a serum phenylalanine concentration of 1.2 mM before initiation of a phenylalanine-restricted diet or in the absence of dietary restrictions later in life. Mild PKU and mild HPA are characterized by serum phenylalanine concentrations of 0.6–1.2 and 0.18–0.6 mM, respectively, without a phenylalanine-restricted diet. Genetic analysis of 41 of 203 patients has been reported elsewhere.7
The 20 patients with L52S/E5_6 del, A132V/R413P, R241C/(R111X, R252P, T278I, Q301H, R413P (7 patients) and R241C (3 patients)), A373T/IVS4nt-1, and P407S/(R111X, R158W and R252W) mutations were confirmed by BH4 loading test and/or phenylalanine breath test with BH4 administration.14, 17, 31 BH4 responsiveness was provided by a decrease of 20% in serum phenylalanine levels in the single-dose BH4 loading test (10 mg/kg), a decrease of 30% in serum phenylalanine levels in the four-dose BH4 loading test (total; 30 mg/kg) and/or two times increase in in vivo PAH activity after the four-dose of BH4 (total; 30 mg/kg) in phenylalanine breath test.
The Institutional Review Board of Osaka City University Graduate School of Medicine (Osaka, Japan) approved all study protocols and informed consent for the genetic analyses was obtained from all patients or their parents/guardians.
DHPLC analysis and sequencing
PCR reactions were performed in a 50-μl reaction mixture containing 0.5–1.0 μg genomic DNA, 2 mM of each dNTP, 50 mM KCl, 100 mM Tris-HCl (pH 8.3), 15 mM MgCl2, 0.1% gelatin, 50 ng of each PCR primer and KOD plus Taq DNA polymerase (Toyobo, Osaka, Japan). Thirty-five PCR cycles (94 °C for 30 s, 50–60 °C for 60 s and 72 °C for 60 s) were performed on a PTC-200 Thermal Cycler (Bio-Rad, Hercules, CA, USA). Table 1 lists the primer pairs used for amplification. A clamped 40-bp GC-sequence was used as the unilateral primer. Before DHPLC analysis, the PCR products were subjected to heteroduplex formation by heating for 5 min at 95 °C followed by cooling to 25 °C over 45 min. DHPLC analyses were performed using a WAVE DNA fragment analysis DHPLC system (Transgenomics, Glasgow, UK), according to the protocol recommended by the manufacturer.
Exons showing abnormal patterns on DHPLC were amplified with PCR primers, as described previously7 and purified using the Illustra GFX PCR DNA and gel-band purification kit (GE Healthcare, Buckinghamshire, UK). Samples were sequenced using the Applied Biosystems Dye-Terminator Kit on an ABI Prism 377 DNA sequencer (Applied Biosystems, Foster City, CA, USA), as described previously.7
MLPA analysis
MLPA (SALSA MLPA kit P055; MRC, Amsterdam, The Netherlands) was carried out for PKU patients with no identified PAH mutations on either or both alleles by DHPLC analysis and direct sequencing, according to the manufacturer's instructions. Genomic DNA was used for the ligation reactions and multiplex PCR amplification. PCR products were mixed with formamide, dextran-blue and the TAMRA-labeled internal-size standard (TAMRA-500, Applied Biosystems) and then analyzed on an ABI-377 gel sequencer (Applied Biosystems) using GeneScan software (Applied Biosystems). The peak height of each exon was compared with the internal-standard peak, and then the ratio of the peak area of each exon to the total peak area for 13 exons was compared between patients and control.
Results
PKU mutations in Japan
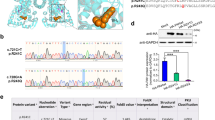
DHPLC with GC-clamped primer was useful to screen the exons with PKU mutations. When a GC-clamped primer was employed, more consistent and correct detection of four peaks with two heteroduplex and two homoduplex peaks were enabled, regardless of the column temperature. Several different PKU mutations were found in the same exon, especially exon 7 and it is sometimes difficult to detect the mutations by DHPLC using an unclamped primer for the optimum temperature differences among mutations, whereas use of a GC-clamped primer would allow the temperature conditions to be relaxed (Figure 1). The exons with negative results by DHPLC with GC-clamped primer showed no mutations by sequencing except for homozygous mutations. False-positive and false-negative for PKU mutation by DHPLC has not been seen.
Denaturing high-performance liquid chromatography (DHPLC) profiles of the T278I, R241C and R252W mutations in exon 7 (a) and the R413P and R408W mutations in exon 12 (b) using unclamped and GC-clamped primers. The mutations showed clear peaks at the same column temperatures using GC-clamped primers.
The 17 alleles without any identified mutations were subjected to MLPA analysis and one allele showed E5_6 del newly, which is known to have a large deletion involving exons 5 and 6 in Japanese patients with PKU. And no large deletion was found except for E5_6 del.
Table 2 details the gene analysis results for 203 patients with PAH deficiency living in Japan and with pre-treatment blood phenylalanine levels over 0.18 mM. Of 406 mutant alleles, 390 (96%) were genotyped and 65 mutations identified. Concerning 16 unknown PKU alleles, there were no patients with unidentified PKU mutations in both alleles. R413P mutation was the most frequently encountered (24.1%), followed by R241C (10.1%), IVS4-1g>a (8.9%), R111X (7.9%) and R243Q (5.4%). Eight prevalent mutations with an incidence over 3% accounted for 69.7% of the PKU alleles. As for private mutations, eight mutations were found in two PKU alleles and 39 mutations in a single PKU allele. These results indicated that PKU mutations in Japanese patients are heterogeneous. The mutations tended to appear in exon 7 (30.1%) and exon 12 (27.0%). Twenty-two novel mutations were identified newly, including A132V, V379A, E183fsX12, S16Y, S70F, V106A, H107R, I174N, Y198X, A202V, P219S, P225L, H271Q, Q267L, Q301H, F402I, F402C, S251fsX90, G312fsX29, K113_K114fsX81, IVS10-2a>t and IVS10-7c>a. R53H was found in the cis position, relative to other mutations in nine alleles. No particular mutation linked to R53H was noted. R53H is not a polymorphism32 and is associated with mild phenotype of PKU in Korean population.5
Comparison among East Asian countries
The distribution and frequency of PKU mutations were compared among Northern Chinese,33 Korean,5 Taiwanese34 and Japanese populations. Figure 2 shows PKU mutations with >5% frequency in at least one of these countries. The prevalent mutations were relatively similar across the analyzed populations, and particularly so between the Northern Chinese and Koreans, with the most frequent PKU mutation identified as R243Q, which accounted for 22.2 and 12.0% of all mutations, respectively. The second most prevalent was E6-96A>g, which accounted for 11.1 and 10.1%, respectively. This A-to-G transition is a splicing mutation for the disappearance of the consensus sequence of a 5′ donor splice site instead of a tyrosine to cysteine substitution of codon 204 in exon 6 and would give a 96 bp deletion.35, 36 The most frequent mutations in the Taiwanese population were R241C (32%) and R408Q (14%), each associated with a mild clinical phenotype. Private mutations, which are those seen in one or two alleles, differed among countries, suggesting a relatively recent origin.
Japanese PKU mutations were then compared among the six geographical districts in which the facilities requesting gene diagnosis were located; nine mutations with a frequency of >2.5% were compared (Figure 3). There was no major inter-district difference in the frequency of mutations. However, the pattern in the Kyushu District differed slightly from that in the other districts and was characterized by no IVS4-1g>a mutations and a relatively high frequency of R111X.
Correlation between clinical phenotype and genotype
Figure 4 shows the relationship between pre-treatment blood phenylalanine level and mild PKU/HPA-associated mutations. R408Q, V106A, P407S, R241C and L52S mutations were suspected to correlate with mild PKU, whereas A132V, R297C, A373T, D415N, A403V, F402I, T380M and V379A mutations were more associated with mild HPA. Only the P407S, R241C and T380M mutations were present in sufficient cases (n>2) for analysis. Fifty-nine (29.1%) patients showed those genotypes associated with mild PKU and HPA in 203 patients. Fifty-two (25.6%) patients showed pre-treatment blood phenylalanine, as shown in Figure 4. The blood phenylalanine level in patients with R243Q/R243Q, reflecting 10% in vitro PAH activity, was lower than in those patients with compound heterozygote of R243Q/severe mutation. The blood phenylalanine level in the patients with compound heterozygote of R243Q/R241C or P407S/R241C was lower than in the patients with compound heterozygote of R241C/severe mutation, R243Q/severe mutation or P407S/severe mutation. The mutations in these cases (Figure 4) were associated with mild PKU and mild HPA in Japanese patients and were assumed to be BH4-responsive mutations. The frequency of BH4-responsive PAH deficiency was predicted to be 29.1% among PKU patients living in Japan.
Correlation between PAH mutations and pretreatment phenylalanine levels. The R243Q to V379A mutations were compound heterozygote with severe mutation. The mean in vitro PAH activity (% of control) of two alleles is shown on the top of bars. D415N and A403V mutations indicated two different in vitro PAH activities. The blue letters show the mutations related with mild PKU. The green letters show the mutations related with mild HPA. The red letters show the mutation between mild PKU and classical PKU; sm indicates severe mutation, n indicates the number of cases analyzed.
Discussion
Genetic diagnosis is useful for evaluating the clinical phenotype of PKU and the diagnosis of BH4-responsive PAH deficiency. It is therefore important to establish a more reliable PKU-associated genetic assay. PKU genes are quite diverse and more than 60 different mutations have been detected in patients with PKU living in Japan. Innovations in genetic engineering have enabled the development of techniques for more rapid and large-scale gene analysis. However, assaying genes involved in single-gene disorders require accuracy and reliability over a small number of samples. The analysis system described herein was designed to (1) detect the exon mutation using GC-clamped DHPLC, (2) sequence the affected exon or all exons for undetectable mutations by DHPLC and (3) identify large deletions using MLPA analysis if mutations are not detected at steps (1) and (2). This method detects gene mutations at a rate of 96%.
The GC-clamped DHPLC shows clear peaks originating from mutations with minimal influence from column temperature and PCR byproducts and heteroduplex peaks are detected at high sensitivity. Only samples amplified from patient DNA are used for DHPLC to maximize sensitivity and to simplify the manipulation. Therefore, patients with homozygous mutations cannot be identified with this system. As incidences in which mutations are not detectable in both alleles are very rare (0.16%), the homozygous mutations are estimated in the patients. In such cases, the shortcoming can be resolved by sequencing exon 7 and 12 and/or all exons. In the current cohort, the actual percentage of homozygous patients was 13.3% in contrast to the theoretical percentage of 9.3% (calculated for Japanese patients on the basis of the data for each mutation). This difference seems to reflect the influence of relative marriage in Japan.
To elucidate the origin of PKU genes in East Asia, we compared PKU genes among the East Asian countries of China, Korea, Taiwan and Japan. The distribution of immunoglobulin G heavy-chain allotype suggested that the Mongoloid in East Asia originates from the northern Mongoloid (around the Lake Baikal) and the southern Mongoloid (Guangxi area in Southwestern China).37 The frequency and distribution of PKU mutation in Southern China were proposed to differ from those in Northern China.6 As shown in Figure 2, the pattern of prevalent mutations was similar among Northern China, Korea and Japan, and particularly so between Northern China and Korea. The prevalent mutation might thus originate from the northern Mongoloid ancestors, before spreading among the populations of Korea and Japan through Northern China.
The origin of Japanese people is considered a mixing of Jomon- and Yayoi-period populations that moved from the East Asian continent to Japan 12 000 and 2000 years ago, respectively. The present genetic analysis suggests that Japanese PKU mutations were affected strongly by PKU mutations of northern Mongoloid origin with additional influence of the geographical characteristics (an isolated country made of islands), thus exerting a ‘founder effect’ leading to a specifically high incidence of R413P (24.1%).
Analysis among six Japanese districts revealed no major differences in the spectrum of PKU mutations, suggesting that Japanese populations comprise a relatively homogeneous ethnic group. The Kyushu District differed slightly from the other districts in the absence of IVS4-1g>a mutations, although available data were not sufficient to analyze possible ancestral contributions underlying this finding (that is, relative influence of Jomon-period people and Yayoi-period people). The lack of major inter-district differences in Japan in this analysis is probably attributable also to the method used for comparison in that the district was defined by the facility requesting the assay rather than the patient's birthplace.
One major objective of the present genetic analysis was to identify mutations associated with the responsiveness to BH4 therapy in East Asian patients. Previous reports cite a BH4 responsiveness of 30–45% in the general PKU population, with reported associations to clinical phenotype and the frequency of responsive cases being 80–100% for mild HPA, 40–60% for mild PKU and <10% for classical PKU.21, 38, 39, 40 Most of the patients in the current study who showed BH4 responsiveness suffer from mild PKU or mild HPA; these patients responded to BH4 loading and showed an increase in in vivo PAH activity by phenylalanine breath test,15, 31 which is proportional to the residual PAH activity by BH4 dosing under the regulation system of BH4 and phenylalanine in in vivo PAH activity.31, 41 Therefore, a residual PAH activity resulting from a combination of both alleles is useful to predict the BH4 responsiveness in PKU patients.42
In PKU, clinical phenotype is basically determined by genotype. BH4 responsiveness is also related to genotype. The frequency of BH4-responsive patients was 29.1% among the present cohort of PKU patients living in Japan. This frequency is not high in comparison with European and North American populations, possibly because of the strict criteria used here to judge BH4 responsiveness. Zurflüh et al.43 listed BH4-responsive mutations using the BIOPKUdb (http://www.bh4.org/BH4Databases.asp) similar to our data in Figure 4. R408Q, P407S, R241C, A373T, D415N, A403V and T380M were confirmed to BH4-responsive mutations in their data. A132V was listed among the unclear mutations. V106A, L52S, R297C and F402I were not listed. Concerning R413P, V388M and R243Q in our Japanese patients, the homozygote of R413P and compound heterozygote of R413P/severe mutation were linked to classical PKU clinically, showed no BH4 responsiveness in BH4 loading test, and had very low-level in in vivo PAH activity in the phenylalanine breath test.31 Therefore, R413P is a BH4-unresponsive mutation. Two patients with compound heterozygote mutations of V388M/R413P showed classical PKU as did 8 patients with R243Q/severe mutation. R243Q and V388M mutations may have residual PAH activity,7, 44 but not to a sufficient level for a BH4 response to reduce in blood phenylalanine levels. These two mutations might be the border between BH4-responder and unresponder. For these reasons, R243Q and V388M were not included in the BH4-responsive mutations. Nevertheless, 1 patient with R243Q/R243Q was expected to be BH4 responsive. For the prediction of BH4-responsive, the combination of the mutations in both alleles is important as described by Karacic et al.42
Another potential factor in the incidence of low-frequency, BH4-responsive mutations in Japan is regional differences. In Europe, the frequency of BH4-responsive cases was higher in Southern Europe and lower in Eastern Europe.45 In East Asia, a high frequency of mild phenotype (84%) was noted in Taiwan,34 whereas the frequency of R413P (a BH4-unresponsive mutation) in Japan was specifically high (24%). This factor might contribute to the relatively low frequency of BH4-responsive mutations in this study.
In addition, blood phenylalanine levels measured by the BH4 loading test might be affected by various environmental factors like food and infection, as well as genotype used to determine the BH4-responsive PAH deficiency. The determination of patient genotype in PKU patients is clearly useful for characterizing clinical and biochemical phenotypes and should permit further optimization of therapy including BH4 therapy, determination of long-term prognosis and intensive support for patients and their families.
References
Scriver, C. R. & Kaufman, S. in The Metabolic and Molecular Bases Of Inherited Disease 8th edn (eds Scriver, C.R., Baudet, A.L., Valle, D., & Sly, W.S.) 1667–1724 (McGraw-Hill, New York, 2001).
Aoki, K. & Wada, Y. Outcome of the patients detected by newborn screening in Japan. Acta Paediatr. Jpn. 30, 429–434 (1988).
Bickel, H., Bachmann, C., Beckers, R., Brandt, N. J., Clayton, B. E., Corrado, G. et al. Neonatal mass screening for metabolic disorders: summary of recent sessions of the committee of experts to study inborn metabolic diseases, public health committee, council of Europe. Eur. J. Pediatr. 137, 133–139 (1981).
Liu, S. R. & Zuo, Q. H. Newborn screening for phenylketonuria in eleven districts. Chin. Med. J. 99, 113–118 (1986).
Lee, D. H., Koo, S. K., Lee, K. S., Yeon, Y. J., Oh, H. J., Kim, S. W. et al. The molecular basis of phenylketonuria in Koreans. J. Hum. Genet. 49, 617–621 (2004).
Okano, Y., Hase, Y., Lee, D. H., Furuyama, J., Shintaku, H., Oura, T. et al. Frequency and distribution of phenylketonuric mutations in Orientals. Hum. Mutat. 1, 216–220 (1992).
Okano, Y., Asada, M., Kang, Y., Nishi, Y., Hase, Y., Oura, T. et al. Molecular characterization of phenylketonuria in Japanese patients. Hum. Genet. 103, 613–618 (1998).
Sueoka, H., Nagao, M. & Chiba, S. Rapid mutation screening of phenylketonuria by polymerase chain reaction-linked restriction enzyme assay and direct sequence of the phenylalanine hydroxylase gene: clinical application in northern Japan and northern China. Genet. Test. 4, 249–256 (2000).
Eisensmith, R. C., Okano, Y., Dasovich, M., Wang, T., Güttler, F., Lou, H. et al. Multiple origins for phenylketonuria in Europe. Am. J. Hum. Genet. 51, 1355–1365 (1992).
Okano, Y.,, Eisensmith, R. C., Gütler, F., Lichter-Konecki, U., Konecki, D. S., Trefz, F. K. et al. Molecular basis of phenotypic heterogeneity in phenylketonuria. N. Engl. J. Med. 324, 1232–1238 (1991).
Guldberg, P., Rey, J., Zschocke, F., Romano, V., Francois, B., Michiels, L. et al. European multicenter study of phenylalanine hydroxylase deficiency: classification of 105 mutations and a general system for genotype-based prediction of metabolic phenotype. Am. J. Hum. Genet. 63, 71–79 (1998).
Kayaalp, E., Treacy, E., Waters, P. J., Byck, S., Nowacki, P. & Scriver, C. R. Human phenylalanine hydroxylase mutations and hyperphenylalaninemia phenotypes: a metanalysis of genotype-phenotype correlations. Am. J. Hum. Genet. 61, 1309–1317 (1997).
Zschocke, J., Graham, C. A., Stewart, F. J., Carson, D. J. & Nevin, N. C. Non-phenylketonuria hyperphenylalaninemia in northern Ireland: frequent mutation allows screening and early diagnosis. Hum. Mutat. 4, 114–118 (1994).
Kure, S., Hou, D. C., Ohura, T., Iwamoto, H., Suzuki, S., Sugiyama, N. et al. Tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. J. Pediatr. 135, 375–378 (1999).
Muntau, A. C., Röschinger, W., Habich, M., Dmmelmair, H., Hoffmann, B., Sommerhoff, C. P. et al. Tetrahydrobiopterin as an alternative treatment for mild phenylketonuria. N. Engl. J. Med. 347, 2122–2132 (2002).
Cerone, R., Schiaffino, M. C., Fantasia, A. R., Perfumo, M., Birk Moller, L. & Blau, N. Long-term follow-up of a patient with mild tetrahydrobiopterin -responsive phenylketonuria. Mol. Genet. Metab. 81, 137–139 (2004).
Shintaku, H., Kure, S., Ohura, T., Okano, Y., Ohwada, M., Sugiyama, N. et al. Long-term treatment and diagnosis of tetrahydrobiopterin-responsive hyperphenylalaninemia with a mutant phenylalanine hydroxylase gene. Pediatr. Res. 55, 425–430 (2004).
Trefz, F. K., Scheible, D., Frauendienst-Egger, G., Korall, H. & Blau, N. Long-term treatment of patients with mild and classical phenylketonuria by tetrahydrobiopterin. Mol. Genet. Metab. 86, S75–S80 (2005).
Hennermann, J. B., Bührer, C., Blau, N., Vetter, B. & Mönch, E. Long-term treatment with tetrahydrobiopterin increases phenylalanine tolerance in children with severe phenotype of phenylketonuria. Mol. Genet. Metab. 86, S86–S90 (2005).
Levy, H. L., Milanowski, A., Chakrapani, A., Cleary, M., Lee, P., Trefz, F. K. et al. Efficacy of sapropterin dihydrochloride (tetrahydrobiopterin, 6R-BH4) for reduction of phenylalanine concentration in patients with phenylketonuria: a phase III randomised placebo-controlled study. Lancet 370, 504–510 (2007).
Blau, N. & Erlandsen, H. The metabolic and molecular bases of tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. Mol. Genet. Metab. 82, 101–111 (2004).
Yao, Y., Matsubara, Y. & Narisawa, K. Rapid detection of phenylketonuria mutations by non-radioactive single-strand conformation polymorphism analysis. Acta Paediatr. Jpn. 36, 231–235 (1994).
Guldberg, P., Henriksen, K. F. & Güttler, F. Molecular analysis of phenylketonuria in Denmark: 99% of the mutations detected by denaturing gradient gel electrophoresis. Genomics 17, 141–146 (1993).
Narayanaswami, G. & Taylor, P. D. Improved efficiency of mutation detection by denaturing high-performance liquid chromatography using modified primers and hybridization procedure. Genet. Test. 5, 9–16 (2001).
Bräutigam, S., Kujat, A., Kirst, P., Seidel, J., Lüleyap, H. U. & Froster, U. G. DHPLC mutation analysis of phenylketonuria. Mol. Genet. Metab. 78, 205–210 (2003).
Dobrowolski, S. F., Ellingson, C., Coyne, T., Grey, J., Martin, R., Naylor, E. W. et al. Mutations in the phenylalanine hydroxylase gene identified in 95 patients with phenylketonuria using novel systems of mutation scanning and specific genotyping based upon thermal melt profiles. Mol. Genet. Metab. 91, 218–227 (2007).
Okano, Y., Hase, Y., Shintaku, H., Araki, K., Furuyama, J- I., Oura, T. et al. Molecular characterization of phenylketonuric mutations in Japanese by analysis of phenylalanine hydroxylase mRNA from lymphoblasts. Hum. Mol. Genet. 4, 659–660 (1994).
Gable, M., Williams, M., Stephenson, A., Okano, Y., Ring, S., Hurtubise, M. et al. Comparative multiplex dosage analysis detects whole exon deletions at the phenylalanine hydroxylase locus. Hum. Mutat. 21, 379–386 (2003).
Desviat, L. R., Pérez, B. & Ugarte, M. Identification of exonic deletions in the PAH gene causing phenylketonuria by MLPA analysis. Clin. Chim. Acta 373, 164–167 (2006).
Kozak, L., Hrabincova, E., Kintr, J., Horky, O., Zapletalova, P., Blahakova, I. et al. Identification and characterization of large deletions in the phenylalanine hydroxylase (PAH) gene by MLPA: evidence for both homologous and non-homologous mechanisms of rearrangement. Mol. Genet. Metab. 89, 300–309 (2006).
Okano, Y., Hase, Y., Kawajiri, M., Nishi, Y., Inui, K., Sakai, N. et al. In vivo studies of phenylalanine hydroxylase by phenylalanine breath test: diagnosis of tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. Pediatr. Res. 56, 714–719 (2004).
Pey, A.L., Stricher, F., Serrano, L . & Martinez, A. Predicted effects of missense mutations on native-state stability account for phenotypic outcome in phenylketonuria, a paradigm of misfolding diseases. Am. J. Hum. Genet. 81, 1006–1024 (2007).
Song, F., Qu, Y. J., Zhang, T., Jin, Y. W., Wang, H. & Zheng, X. Y. Phenylketonuria mutations in Northern China. Mol. Genet. Metab. 86, S107–S118 (2005).
Chien, Y. H., Chiang, S. C., Huang, A., Chou, S. P., Tseng, S. S., Huang, Y. T. et al. Mutation spectrum in Taiwanese patients with phenylalanine hydroxylase deficiency and a founder effect for the R241C mutation. Hum. Mutat. 23, 206 (2004).
Wang, T., Okano, Y., Eisensmith, R. C., Lo, W. H. Y., Huang, S. Z., Zeng, Y. T. et al. Missense mutations prevalent in Orientals with phenylketonuria: molecular characterization and clinical implications. Genomics 10, 449–456 (1991).
Ellingsen, S., Knappskog, P. M. & Eiken, H. G. Phenylketonuria splice mutation (Exon 6nt-96A-g) masquerading as missense mutation (Y204C). Hum. Mutat. 9, 88–90 (1997).
Matsumoto, H. Characterization of Mongoloid and neighboring populations based on the genetic markers of human immunoglobulins. Hum. Genet. 80, 207–218 (1998).
Desviat, L. R., Pérez, B., Bèlanger-Quintana, A., Castro, M., Aguado, C., Sánchez, A. et al. Tetrahydrobiopterin responsiveness: results of the BH4 loading test in 31 Spanish PKU patients and correlation with their genotype. Mol. Genet. Metab. 83, 157–162 (2004).
Leuzzi, V., Carducci, C., Carducci, C, Chiarotti, F., Artiola, C., Giovanniello, T. et al. The spectrum of phenylalanine variations under tetrahydrobiopterin load in subjects affected by phenylalanine hydroxylase deficiency. J. Inherit. Metab. Dis. 29, 38–46 (2006).
Fiege, B. & Blau, N. Assessment of tetrahydrobiopterin (BH4) responsiveness in phenylketonuria. J. Pediatr. 150, 627–630 (2007).
Okano, Y., Takatori, K., Kudo, S., Sakaguchi, T., Asada, M., Kajiwara, M. et al. Effects of tetrahydrobiopterin and phenylalanine on in vivo human phenylalanine hydroxylase by phenylalanine breath test. Mol. Genet. Metab. 92, 308–314 (2007).
Karacić, I., Meili, D., Sarnavka, V., Heintz, C., Thöny, B., Ramadza, D. P. et al. Genotype-predicted tetrahydrobiopterin (BH4)-responsiveness and molecular genetics in Croatian patients with phenylalanine hydroxylase (PAH) deficiency. Mol. Genet. Metab. 97, 165–171 (2009).
Zurflüh, M. R., Zschocke, J., Lindner, M., Feillet, F., Chery, C., Burlina, A. et al. Molecular genetics of tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency. Hum. Mutat. 29, 167–175 (2008).
Wang, T., Okano, Y., Eisensmith, R. C., Lo, W. H. Y., Huang, S. Z., Zeng, Y. T. et al. Missense mutations prevalent in Orientals with phenylketonuria: molecular characterization and clinical implication. Genomics 10, 449–456 (1991).
Zschocke, J. Phenylketonuria mutations in Europe. Hum. Mutat. 21, 345–356 (2003).
Acknowledgements
This study was supported in part by grants from the Ministry of Education, Culture, Sports, Science and Technology of Japan, the Ministry of Health, Labor and Welfare of Japan, by funds from Morinaga Houshikai Association and from Suyama Research Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Okano, Y., Kudo, S., Nishi, Y. et al. Molecular characterization of phenylketonuria and tetrahydrobiopterin-responsive phenylalanine hydroxylase deficiency in Japan. J Hum Genet 56, 306–312 (2011). https://doi.org/10.1038/jhg.2011.10
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2011.10
Keywords
This article is cited by
-
Identification of deep intronic variants of PAH in phenylketonuria using full-length gene sequencing
Orphanet Journal of Rare Diseases (2023)
-
The spectrum of phenylalanine hydroxylase variants and genotype–phenotype correlation in phenylketonuria patients in Gansu, China
Human Genomics (2023)
-
Characterization of phenylalanine hydroxylase gene variants and analysis of genotype–phenotype correlation in patients with phenylalanine hydroxylase deficiency from Fujian Province, Southeastern China
Molecular Biology Reports (2022)
-
Neonatal screening and genotype-phenotype correlation of hyperphenylalaninemia in the Chinese population
Orphanet Journal of Rare Diseases (2021)
-
Phenylketonuria
Nature Reviews Disease Primers (2021)