Abstract
The research on antibiotics requires the integration of broad areas, such as microbiology, organic chemistry, biochemistry and pharmacology. It is similar to the field of chemical biology that is recently popular as an approach for drug discovery. When we isolate a new compound from a microorganism, we can pursue the interesting research on chemistry and biology. In this review, I would like to introduce our achievements in relation to reveromycin A.
Similar content being viewed by others
Introduction
Currently, antibiotics are widely used in medicine and agriculture as antimicrobial compounds. Selman A Waksman (1888–1973)1 originally defined an antibiotic as a substance produced by one microorganism that interferes with the growth or function of other microorganisms. Since the mid-1950s, several antitumor compounds and molecules with other bioactivities have been isolated from microorganisms, broadening the definition of antibiotics to include antitumor compounds, enzyme inhibitors and other compounds. Despite this broadening of the recognized biological activities of microbial metabolites, the term ‘antibiotic’ should be used to refer to an antimicrobial substance to eliminate confusion. From a different perspective, ‘antibiotics’ can be understood as the name of an academic field in the same sense that economics, physics, and mathematics are academic fields. Thus, we recognize ‘antibiotics’ as an academic area that integrates broader areas, such as microbiology, organic chemistry, biochemistry, and pharmacology. Recently, the field of chemical biology, which integrates chemistry and biology, has provided a productive approach to drug discovery. The concept of ‘antibiotics’ is similar to the concept of ‘chemical biology.’
In this review, I would like to introduce our achievements in relation to reveromycin A that was isolated from a soil microbial strain. The starting point of our research is always the isolation of new compounds. Once such a compound is identified, we can pursue unique lines of research, including studies of its mode of action and biosynthesis. Our chemical biology research thus originates in antibiotics research.
The origin of chemical biology in RIKEN
I would first like to describe RIKEN, the abbreviation for the Japanese name for the Institute of Physical and Chemical Research. RIKEN was modeled on the Kaiser Wilhelm Society and was formally founded in 1917. In 2003, Nobel laureate Dr Ryoji Noyori assumed the presidency of RIKEN. In 2015, RIKEN became one of the national research and development institutions and Dr Hiroshi Matsumoto succeeded Dr Noyori as the president. RIKEN is the largest comprehensive research institution in Japan, with a diverse range of scientific disciplines.
When we trace the history of chemical biology in RIKEN, we recall two pioneers of ‘Nogei Kagaku’, a discipline that is similar to agricultural chemistry and that aims to explain unsolved biological phenomena by understanding chemical compounds.2, 3 Umetaro Suzuki (1874–1943) aimed to cure beriberi and he isolated oryzanin from rice bran; this compound is now known as thiamine (vitamin B1). This was the first vitamin to be identified and its discovery initiated the field of vitaminology. Teijiro Yabuta (1888–1977) revealed the cause of bakanae (or ‘foolish seedling’) disease in rice plants. Bakanae disease is caused by a fungal infection by Gibberella fujikuroi. Yabuta isolated gibberellin from the fungus, a molecule that was subsequently detected ubiquitously in plants and was ultimately recognized as a plant hormone. Our chemical biology research at RIKEN has developed from the foundations established by these scientists. Kin-ichiro Sakaguchi (1897–1994) was an applied microbiologist who proposed the establishment of a culture collection to provide a central microorganism stock for Japan. This collection has ensured that important microorganisms are safely stored and can be readily accessed by researchers who want to use them for their investigations or inventions. Storage and distribution of these microorganisms support the work of both basic and industrial research. This pioneering work encouraged me to establish the chemical bank, a natural product depository (NPDepo) at RIKEN. Saburo Suzuki and Kiyoshi Isono were former directors of the RIKEN Antibiotics Laboratory that I currently direct. In the 1960s, they discovered polyoxin, an antifungal compound that was marketed worldwide.4 Moreover, the studies that resulted in the discovery of polyoxin initiated various lines of basic research, such as mode of action and total synthesis studies.5, 6
Bioprobes
When I became the director of the Antibiotics Laboratory at RIKEN, I focused on microbial metabolites that were active against cancer cells. We isolated novel compounds from microorganisms, and tested these using chemical and biological approaches.7
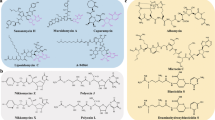
We have been investigating and controlling protein activities using small molecules. In molecular biology, gene mutation, gene knockout and RNA interference are the major strategies used to analyze protein function.8 Although these methods are reliable and specific to the gene of interest, they can take a long time to produce results. Chemical biology or chemical genetic approaches use small molecules that alter protein function rapidly and conditionally, just by their addition or removal. We named these ‘bioprobes’ and they provide useful tools to investigate the biological functions of proteins.8, 9 Some of these compounds are commercially available (Figure 1).
The mode of action of reveromycin A
During an antitumor compound screening study, reveromycin A was isolated from a soil bacteria, Streptomyces reveromyceticus SN-593, and found to inhibit mitogenic activity induced by epidermal growth factor.10 We subsequently examined the antitumor activity of reveromycin A in vivo and revealed its mode of action.11, 12, 13, 14
Reveromycin A showed potent in vivo antitumor activity against hormone-dependent tumors, such as ovarian cancer and prostate cancer, but was not as active against other types of tumors.14 However, even in mice where reveromycin A did not cure the tumors, it did protect from terminal-stage cachexia and hypercalcemia. This observation indicated that reveromycin A might be involved in calcium homeostasis. Bone provides a calcium reservoir and its remodeling is regulated by osteoblasts and osteoclasts. We evaluated the effects of reveromycin A on osteoclasts and osteoblasts. As shown in Table 1, reveromycin A was not strongly cytotoxic to tumor cells, but it did have potent and selective effects on osteoclasts. When reveromycin A was added to a coculture of osteoclasts and osteoblasts, the osteoclasts were killed and the osteoblasts survived. The data clearly demonstrated this selective toxicity of reveromycin A toward osteoclasts (Figure 2).15
Next, we wanted to know why reveromycin A selectively killed osteoclasts. We assumed that reveromycin A did not penetrate the cell membrane because of the acidity derived from three carboxylic acid residues (Figure 3). In general, acidic compounds show poor membrane permeability under normal conditions because the plasma membrane consists of negatively charged phospholipids. However, mature osteoclasts form an acidic environment to resorb bones. Under these acidic conditions, the protonation of carboxylic acid in reveromycin A is suppressed and reveromycin A can enter osteoclasts.
To test this hypothesis, we synthesized [3H]-labeled reveromycin A and demonstrated its selective incorporation into osteoclasts. As shown in Figure 4, reveromycin A was incorporated into the mature osteoclasts in their acidic environment, but not into the macrophages; these are the osteoclast precursor cells and they do not produce acid. Reveromycin A also failed to enter other cell lines. These observations suggested that the selective toxicity of reveromycin A related to its membrane permeability. The low membrane permeability of osteoblasts and osteoclast precursor cells rendered them resistant to reveromycin A, but the acidic environment formed by mature osteoclasts made them highly sensitive to reveromycin A.
Incorporation of [3H]reveromycin A into cells. BG-1, human ovarian carcinoma cells; Colon26, mouse rectal carcinoma cells; HeLa, human cervical carcinoma cells; Mφ, mouse macrophages; MC3T3-E1, mouse osteoblastic cells; MDCK, canine kidney epithelial cells; OCs, mouse osteoclasts; RAW264, mouse macrophages; UAMS, mouse osteoblastic cells.
To reveal the molecular target of reveromycin A, we collaborated with Dr Miyakawa’s group in Hiroshima University.16, 17 The target was identified as isoleucyl-transfer RNA (tRNA) synthetase using a genetic approach in Saccharomyces cerevisiae.17 One amino acid substitution, from asparagine to aspartic acid at position 660 in the yeast isoleucyl-tRNA synthetase gene (ILS1p), produced a reveromycin-resistant enzyme (Figure 5). Later, we carried out biochemical analyses that revealed that isoleucyl-tRNA synthetase is also the target of reveromycin A in osteoclasts. Isoleucyl-tRNA synthetase is an essential enzyme for protein synthesis in prokaryotes and eukaryotes. Analyses of one point mutation in this enzyme indicated that the reveromycin A-binding site was located at the editing domain of isoleucyl-tRNA synthetase. The amino acid sequence of this domain is conserved in eukaryotic organisms, but differs in prokaryotes. Therefore, reveromycin A inhibited the enzyme activity of eukaryotic isoleucyl-tRNA synthetase, but not of bacterial isoleucyl-tRNA synthetase. Moreover, the survival of osteoclasts is more dependent on isoleucine and glutamine than on proline and alanine. When isoleucine or glutamine is removed from the cell culture medium, osteoclasts rapidly enter apoptosis.
Taken together, we could propose two mechanisms underlying the specific effects of reveromycin A on mature osteoclasts. The first relates to the membrane permeability of reveromycin A that does not enter normal cells but can penetrate osteoclasts because of their acidic environment. The second relates to the greater dependency of osteoclasts on isoleucine for survival, as compared with other cell types.
In vivo efficacy of reveromycin A
Osteoporosis is caused by augmented osteoclast activity. Metastatic bone disease is also mediated by osteoclast function.18 Therefore, osteoclasts are the ideal therapeutic target to treat both osteoporosis and bone metastasis.
Antiosteoporotic activity
We first examined the antiosteoporotic activity of reveromycin A using an ovary-removed mouse model. The removal of the ovaries from female mice causes similar symptoms as those seen in postmenopausal human osteoporosis. Figure 6 shows the bone density of the trabecular bone area15 that was markedly reduced in the ovary-removed mice. However, this bone density loss was suppressed in mice treated with reveromycin A. Similar protective effects were observed in mice receiving a low-calcium diet.15, 19
Reveromycin A prevented osteoporosis in ovariectomized mice. The mouse trabecular bones are shown in the upper panel (soft X-ray). (a) Bone prepared from a mouse without intact ovaries. (b) Bone prepared from an ovariectomized mouse. (c) Bone from an ovariectomized mouse treated with reveromycin A. Reveromycin A was injected intravenously at the indicated dose twice daily. Quantification of trabecular density by peripheral quantitative computed tomography (pQCT) is shown in the lower panel. *P<0.05; **P<0.001.
Antimetastatic activity
We examined the antimetastatic activity of reveromycin A using a bone metastasis model system, in collaboration with Dr Sone’s group in Tokushima University.20 Human small-cell lung cancer SBC-5 cells were injected intravenously in natural killer cell-depleted SCID (severe combined immunodeficient) mice. In this model, the tumor cells formed metastatic foci in the bone, as well as in the lungs, liver and kidneys. Administration of reveromycin A inhibited the bone metastasis, but not those located in other organs. These findings clearly indicated that reveromycin A did not kill the tumor cells, but inhibited bone metastasis, presumably via its antiosteoclastic activity.
Research conducted by our collaborators, Dr Len Neckers and colleagues, at the National Cancer Institute, USA, found that 17-N-allylamino-17-demethoxygeldanamycin (17-AAG), which shows potent antitumor activity against various tumors, unexpectedly promoted bone metastasis of prostate tumors. These researchers speculated that inhibition of Src-protein by 17-AAG activated osteoclast function because the Src protein plays an important role in osteoclast homeostasis. A tyrosine kinase inhibitor, dasatinib, was found to inhibit the bone metastasis induced by 17-AAG in prostate tumor-bearing mice. A combination therapy study was conducted using 17-AAG and reveromycin A and this found that reveromycin A had a potent suppressive effect on the effects of 17-AAG on prostate tumor bone metastasis.21
Antiperiodontitis activity
When we reported the antiosteoclastic activity of reveromycin A, Drs Miyazawa and Goto in Aichi-Gakuin University were interested in using reveromycin A to treat periodontitis, a gum disease. They established a unique model of periodontitis in mice. It was previously thought that microbial flora caused human periodontal disease. However, even though these causal bacteria were absent from the mouse mouth, symptoms similar to those of periodontitis were observed in mice. This indicated that novel compounds that differed from antibacterial agents such as minocycline were required to treat periodontitis.22 At first, these researchers considered whether the suppression of osteoclast function could cure periodontitis and used bisphosphonates, analogs of pyrophosphate, to suppress osteoclast activity. However, this approach did not produce the expected results. Moreover, several reports describe adverse effects of bisphosphonates. Bisphosphonates are firstly incorporated into bone and then into osteoclasts. Therefore, these compounds can sometimes increase jaw fragility. Thus, they wanted to investigate whether reveromycin A provided effective treatment of periodontal disease, as an alternative to the bisphosphonates.
When an elastic module was inserted between the teeth, the number of osteoclasts increased and alveolar bone loss was observed. The bone loss in osteoprotegerin-deficient knockout mice was similar to that observed in periodontitis. Administration of reveromycin A to osteoprotegerin-deficient mice reduced osteoclast counts and suppressed alveolar bone loss.23, 24, 25
In summary, these in vivo evaluations of reveromycin A showed that it was active against osteoclasts and had potent activity in animal models of osteoporosis, bone metastasis of tumor cells and periodontal disease.
Biosynthesis of reveromycin A
To pursue further in vivo evaluation, a large amount of reveromycin A was required. Although we had succeeded in the total synthesis of reveromycin A, this chemical synthesis method was unable to supply sufficient quantities of reveromycin A.26, 27, 28 One of the difficulties associated with this synthesis was the instability of the spiroacetal ring; this 6,6-membered ring was easily converted to a 5,6-membered ring. It was also difficult to succinylate the tertiary hydroxyl residue (Figure 7). We therefore executed this reaction under ultrahigh pressure. To overcome these difficulties, we developed a biosynthetic method by gene engineering of the reveromycin gene cluster. This facilitated reveromycin A production by microorganisms at a normal pressure and temperature.
We succeeded in the cloning of the reveromycin A biosynthetic gene cluster among many similar polyketide synthase gene clusters.29 The key point was that we knew the specific medium that enhanced reveromycin A production. Because the production titer of reveromycin A was low initially, we tested several different kinds of fermentation media in order to optimize fermentation of the actinomycetes strain. This found that the V8 juice-containing medium was the most effective. We then investigated which ingredient of this juice had the greatest effect and found that tomato extract enhanced the production of reveromycin A. Next, we studied which specific polyketide synthase genes were induced by tomato extract. After exploring the expression of genes encoding ketosynthase–acyltransferase and enoylreductase by reverse transcription-PCR, we identified the reveromycin biosynthetic gene cluster that extended for 91 kb and contained 21 open reading frames (Figure 8).29
Polyketide compounds including a spiroacetal moiety
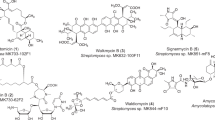
The spiroacetal structure is widely found in natural products. However, with the exception of several antibiotics, the biosynthetic routes for spiroacetal ring formation have not been fully elucidated. To date, two possible biosynthetic routes have been proposed: one involves cyclization of epoxide/ketone intermediates30, 31 and the other one involves dehydrative cyclization.32 A polyether compound, monensin A, is biosynthesized via cyclization of epoxide/ketone intermediates.30, 31 On the other hand, a protein phosphatase inhibitor, tautomycin, and an anthelmintic antibiotic, avermectin, were reported to be synthesized via dehydrative cyclization.32
Regarding reveromycin A biosynthesis, we used a combination of biochemical analyses and gene knockdown to identify two enzymes (RevG and RevJ) that were involved in spiroacetal formation (Figure 9).29 RevG acted as a dihydroxy ketone synthase and formed the keto-diol product. Interestingly, this keto-diol intermediate underwent dehydrative cyclization in the absence of the enzyme under acidic conditions. However, in this noncatalyzed situation, both the native (15S) and nonnative (15R) spiroacetal stereoisomers were detected in a 3:2 ratio. In the presence of RevJ, stereocontrolled cyclization occurred and only the native (15S) stereoisomer was produced.
Biochemical characterization of RevI
Next, I would like to describe hydroxylation at the C18 position. As RevI was annotated to be P450, we examined whether this enzyme catalyzed hydroxylation at the C18 position. P450 RevI was expressed in Escherichia coli and purified to homogeneity. The correct folding of the protein was confirmed by measuring the CO difference spectrum, and the resulting protein efficiently introduced the C18 hydroxyl residue, converting reveromycin T (a succinate-lacking reveromycin A derivative) to reveromycin T1 (C18 hydroxyreveromycin T).33 Figure 10 shows the cocrystal structure of RevI complexed with reveromycin T. This structure was determined at a resolution of 1.4 Å and solved by multi-wavelength anomalous diffraction. This crystallography revealed that RevI formed a relatively large substrate binding pocket that accommodated a bent reveromycin T conformation. Two arginine residues interacted with the two carboxyl groups of reveromycin T and the hydroxylated C18 faced the heme group.33
Succinylation of C18 tertiary alcohol
During chemical synthesis, the succinylation of the tertiary alcohol required ultrahigh pressure (1.5 Gpa).26 However, S. reveromyceticus easily performed this reaction under normal culture conditions. We investigated the genes required for tertiary alcohol succinylation in the reveromycin A biosynthetic gene cluster. Based on gene disruption and metabolites analyses, three genes (revK, revL and revM) were speculated to be involved in the succinylation of reveromycin T1.
Structure–activity relationship study
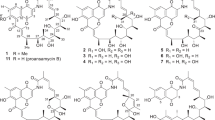
Biosynthetic production enabled us to generate a variety of reveromycin derivatives and to investigate their biological activities, as shown in Table 2 and Figure 11.34 Reveromycin A and its intermediate or shunt metabolites were obtained from the fermentation broth of the wild-type reveromycin A-producing strain. In the rev-deleted mutant, reveromycin T was accumulated. Reveromycin ethylester was accumulated in the presence of ethanol.34
Reveromycin A was most potent against osteoclasts, but was not cytotoxic to tumor cells. On the other hand, reveromycin T, which lacked a succinate moiety, had more potent effects than reveromycin A on human leukemia cells.
Conclusion
I want to emphasize that the discovery of a new compound can open a new door of scientific research. In this review article, I have introduced the story of reveromycin A as an example of this, describing its isolation, biological study and synthesis. Bioactive small molecules can be used both as therapeutic agents and as bioprobes7, 8, 9 to elucidate complex biological functions.
Target identification is important because without it, we cannot apply the compound to the appropriate disease. To my regret, we wasted a lot of time before the discovery of the specific effects of reveromycin A on osteoclasts. To avoid repeating this mistake, we have established new systems to identify the targets of bioactive compounds more rapidly. These integrated profiling systems using MorphoBase and ChemProteoBase provide a powerful strategy to elucidate bioactive compound targets.35, 36, 37, 38
Recently, compounds synthesized by combinatorial chemistry have been viewed as more suitable than natural products for investigation using high-throughput screening that can process a large number of samples. However, microbial metabolites still represent an important resource for drug discovery because of their unique chemical structures and potent biological activities. By overcoming the drawbacks associated with natural products, biosynthesis enables us to produce the desired derivatives and the most useful bioactive microbial metabolites.
Incidentally, the Nobel Prize in Physiology or Medicine 2015 was awarded to three scientists for the discovery of natural products with activities against parasites that cause tropical diseases. Namely, Drs Omura and Campbell were recognized for their discovery of anthelmintic antibiotics and Dr Tu was recognized for her discovery of a novel antimalarial compound. This award reminds us of the importance of drug discovery from natural sources.
References
Waksman, S. A. Antagonistic relations of microorganisms. Bacteriol. Rev. 5, 231–291 (1941).
Osada, H. The Japanese Society for Chemical Biology. ACS Chem. Biol. 1, 8 (2006).
Osada, H. Introduction of new tools for chemical biology research on microbial metabolites. Biosci. Biotechnol. Biochem. 74, 1135–1140 (2010).
Suzuki, S. et al. A new antibiotic, polyoxin. J. Antibiot. 18, 131 (1965).
Endo, A., Kakiki, K. & Misato, T. Mechanism of action of the antifungal agent polyoxin D. J. Bacteriol. 104, 189–196 (1970).
Kuzuhara, H., Ohrui, H. & Emoto, S. Total synthesis of polyoxin J. Tetrahedron. Lett. 14, 5055–5058 (1973).
Osada, H. Development and application of bioprobes for mammalian cell cycle analyses. Curr. Med. Chem. 10, 727–732 (2003).
Osada, H. in Bioprobes (ed. Osada, H.) 1–14 Springer, (2000).
Osada, H. Bioprobes for investigating mammalian cell cycle control. J. Antibiot. 51, 973–982 (1998).
Osada, H., Koshino, H., Isono, K., Takahashi, H. & Kawanishi, G. Reveromycin A, a new antibiotic which inhibits the mitogenic activity of epidermal growth factor. J. Antibiot. 44, 259–261 (1991).
Koshino, H., Takahashi, H., Osada, H. & Isono, K. Reveromycins, new inhibitors of eukaryotic cell growth. III. Structures. J. Antibiot. 45, 1420–1427 (1992).
Takahashi, H. et al. Reveromycins, new inhibitors of eukaryotic cell growth. I. Producing organism, fermentation, isolation and physico-chemical properties. J. Antibiot. 45, 1409–1413 (1992).
Takahashi, H. et al. Reveromycins, new inhibitors of eukaryotic cell growth. II. Biological activities. J. Antibiot. 45, 1414–1419 (1992).
Takahashi, H. et al. Inhibitory action of reveromycin A on TGF-alpha-dependent growth of ovarian carcinoma BG-1 in vitro and in vivo. Oncol. Res. 9, 7–11 (1997).
Woo, J.-T. et al. Reveromycin A, an agent for osteoporosis, inhibits bone resorption by inducing apoptosis specifically in osteoclasts. Proc. Natl Acad. Sci. USA 103, 4729–4734 (2006).
Cui, Z., Hirata, D., Tsuchiya, E., Osada, H. & Miyakawa, T. The multidrug resistance-associated protein (MRP) subfamily (Yrs1/Yor1) of Saccharomyces cerevisiae is important for the tolerance to a broad range of organic anions. J. Biol. Chem. 271, 14712–14716 (1996).
Miyamoto, Y. et al. Identification of Saccharomyces cerevisiae isoleucyl-tRNA synthetase as target of G1 -specific inhibitor reveromycin A. J. Biol. Chem. 277, 28810–28814 (2002).
Yin, J. J., Pollock, C. B. & Kelly, K. Mechanisms of cancer metastasis to the bone. Cell Res. 15, 57–62 (2005).
Kawatani, M. & Osada, H. Osteoclast-targeting small molecules for the treatment of neoplastic bone metastases. Cancer Sci. 100, 1999–2005 (2009).
Muguruma, H. et al. Reveromycin A inhibits osteolytic bone metastasis of small-cell lung cancer cells, SBC-5, through an antiosteoclastic activity. Clin. Cancer Res. 11, 8822–8828 (2005).
Yano, A. et al. Inhibition of Hsp90 activates osteoclast c-Src signaling and promotes growth of prostate carcinoma cells in bone. Proc. Natl Acad. Sci. USA 105, 15541–15546 (2008).
Umeda, M. et al. Microbial flora in the acute phase of periodontitis and the effect of local administration of minocycline. J. Periodontol. 67, 422–427 (1996).
Tanaka, M. et al. Effect of Reveromycin A on experimental tooth movement in OPG-/- mice. J. Dent. Res. 91, 771–776 (2012).
Yabumoto, T. et al. Stabilization of tooth movement by administration of reveromycin A to osteoprotegerin-deficient knockout mice. Am. J. Orthod. Dentofacial Orthop. 144, 368–380 (2013).
Mizuno, M. et al. Reveromycin A administration prevents alveolar bone loss in osteoprotegerin knockout mice treat periodontal disease. Sci. Rep. 5, 16510 (2015).
Shimizu, T., Kobayashi, R., Osako, K., Osada, H. & Nakata, T. Synthetic studies on reveromycin A: stereoselective synthesis of the spiroketal system. Tetrahedron Lett. 37, 6755–6758 (1996).
Shimizu, T. et al. Synthesis and biological activities of reveromycin A and spirofungin a derivatives. Bioorg. Med. Chem. Lett. 18, 3756–3760 (2008).
Shimizu, T. et al. Chemical modification of reveromycin A and its biological activities. Bioorg. Med. Chem. Lett. 12, 3363–3366 (2002).
Takahashi, S. et al. Reveromycin A biosynthesis uses RevG and RevJ for stereospecific spiroacetal formation. Nat. Chem. Biol. 7, 461–468 (2011).
Oliynyk, M. et al. Analysis of the biosynthetic gene cluster for the polyether antibiotic monensin in Streptomyces cinnamonensis and evidence for the role of monB and monC genes in oxidative cyclization. Mol. Microbiol. 49, 1179–1190 (2003).
Gallimore, A. R. et al. Evidence for the role of the monB genes in polyether ring formation during monensin biosynthesis. Chem. Biol. 13, 453–460 (2006).
Li, W., Ju, J., Rajski, S. R., Osada, H. & Shen, B. Characterization of the tautomycin biosynthetic gene cluster from Streptomyces spiroverticillatus unveiling new insights into dialkylmaleic anhydride and polyketide biosynthesis. J. Biol. Chem. 283, 28607–28617 (2008).
Takahashi, S. et al. Structure-function analyses of cytochrome P450revI involved in reveromycin A biosynthesis and evaluation of the biological activity of its substrate, reveromycin T. J. Biol. Chem. 289, 32446–32458 (2014).
Nogawa, T. et al. Creation of novel reveromycin derivatives by alcohol-added fermentation. J. Antibiot. (Tokyo) 66, 247–250 (2013).
Muroi, M. et al. Application of proteomic profiling based on 2D-DIGE for classification of compounds according to the mechanism of action. Chem. Biol. 17, 460–470 (2010).
Futamura, Y. et al. Morphobase, an encyclopedic cell morphology database, and its use for drug target identification. Chem. Biol. 19, 1620–1630 (2012).
Futamura, Y. et al. Identification of a molecular target of a novel fungal metabolite, pyrrolizilactone, by phenotypic profiling systems. Chembiochem 14, 2456–2463 (2013).
Futamura, Y., Muroi, M. & Osada, H. Target identification of small molecules based on chemical biology approaches. Mol. Biosyst. 9, 897–914 (2013).
Acknowledgements
I thank my collaborators for their great efforts, especially, K Isono for his helpful guidance at the beginning of this project. I acknowledge H Takahashi, H Koshino and M Ubukata (now Hokkaido Univ) for the isolation and structural determination of reveromycin A. Regarding the biological studies of reveromycin A, the collaborations with T Miyakawa (Hiroshima Univ), JT Woo, K Nagai (Chubu Univ), S Sone (Tokushima Univ), L Neckers (NCI), S Nagano (Tottori Univ), K Miyazawa and S Goto (Aichi-Gakuin Univ) were indispensable. I also thank T Usui (now Tsukuba Univ), N Kanoh (now Tohoku Univ), T Shimizu and T Nakata for the chemical studies of reveromycin A. I express my gratitude to the current members of our laboratory, particularly T Nogawa, M Kawatani and S Takahashi, for their contributions to the reveromycin study. I express my sincere thanks to the editorial office for giving me a chance to write this review article that is based on the award lecture for the Inhoffen Medal (23 April 2015), delivered in Braunschweig, Germany. I also thank the Helmholtz Centre for Infection Research and the Technical University Braunschweig for awarding me the Inhoffen Medal 2015. The research on reveromycin was supported by the MEXT Grant-in-Aid for Creative Scientific Research, JSPS KAKENHI and the Science and Technology Research Promotion Program for Agriculture, Forestry, Fishery and Food Industry.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no conflict of interest.
Rights and permissions
About this article
Cite this article
Osada, H. Chemical and biological studies of reveromycin A. J Antibiot 69, 723–730 (2016). https://doi.org/10.1038/ja.2016.57
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2016.57
This article is cited by
-
Studies on Streptomyces sp. SN-593: reveromycin biosynthesis, β-carboline biomediator activating LuxR family regulator, and construction of terpenoid biosynthetic platform
The Journal of Antibiotics (2022)
-
Inhibitory mechanism of reveromycin A at the tRNA binding site of a class I synthetase
Nature Communications (2021)