Abstract
Bacteria in the 16S rRNA clade SAR86 are among the most abundant uncultivated constituents of microbial assemblages in the surface ocean for which little genomic information is currently available. Bioinformatic techniques were used to assemble two nearly complete genomes from marine metagenomes and single-cell sequencing provided two more partial genomes. Recruitment of metagenomic data shows that these SAR86 genomes substantially increase our knowledge of non-photosynthetic bacteria in the surface ocean. Phylogenomic analyses establish SAR86 as a basal and divergent lineage of γ-proteobacteria, and the individual genomes display a temperature-dependent distribution. Modestly sized at 1.25–1.7 Mbp, the SAR86 genomes lack several pathways for amino-acid and vitamin synthesis as well as sulfate reduction, trends commonly observed in other abundant marine microbes. SAR86 appears to be an aerobic chemoheterotroph with the potential for proteorhodopsin-based ATP generation, though the apparent lack of a retinal biosynthesis pathway may require it to scavenge exogenously-derived pigments to utilize proteorhodopsin. The genomes contain an expanded capacity for the degradation of lipids and carbohydrates acquired using a wealth of tonB-dependent outer membrane receptors. Like the abundant planktonic marine bacterial clade SAR11, SAR86 exhibits metabolic streamlining, but also a distinct carbon compound specialization, possibly avoiding competition.
Similar content being viewed by others
Introduction
The term SAR86 refers to a 16S rRNA clade of γ-proteobacteria observed in clone libraries constructed from surface ocean microbial communities (Britschgi and Giovannoni, 1991; Schmidt et al., 1991; Mullins et al., 1995). Subsequent 16S-based surveys have verified the ubiquity of this clade in the global ocean (Gonzalez et al., 2000; Malmstrom et al., 2007; Schattenhofer et al., 2009), whereas studies at open ocean and coastal time-series sites show that SAR86 is most abundant in the surface ocean following the onset of thermal stratification (Morris et al., 2005; Mary et al., 2006; Treusch et al., 2009). Multiple phylogenetic sub-classes are present within the SAR86 16S clade, though it is not known if this is related to differentiations in ecophysiology (Treusch et al., 2009).
Sequencing of ∼40 kbp bacterial artificial chromosome (BAC) and fosmid clones generated from a variety of locations revealed that some SAR86 genomes contain a gene coding for a rhodopsin capable of generating a proton motive force (pmf) using light energy (proteorhodopsin, Beja et al., 2000; Sabehi et al., 2004, 2005). Subsequently, proteorhodopsin was observed in many other marine bacteria and, in some cases, shown to provide an ancillary source of energy under some conditions (DeLong and Beja, 2010). As SAR86 remains resistant to cultivation, the role of proteorhodopsin in SAR86 remains unknown, as does SAR86's overall metabolic capabilities.
Shotgun sequencing metagenomic studies like the Global Ocean Sampling (GOS) have provided an unbiased and quantitative survey of genome fragments from microbial communities inhabiting the surface ocean (Venter et al., 2004; Rusch et al., 2007). Without genomic context, it is difficult to subdivide this collection of genes into the metabolic units that are organisms. Thus, it was sobering to find that the ∼200 genomes available for marine microbes provide a reference for only ∼25% of the GOS dataset at a nucleotide identity of 50% (Yooseph et al., 2010), a deficiency particularly severe for non-photosynthetic microbes. Among the uncultivated majority, SAR86 16S clades were the most abundant lineage (Yooseph et al., 2010), indicating that genomes from this lineage could add a great amount of contextual information to metagenomes from the marine environment.
Materials and methods
Single cell collection, screening and multiple displacement amplification (MDA)
Seawater was collected at the Scripps Institution of Oceanography (SIO) Pier on 9 April 2009 at 8:30 am. The water temperature was 13.5 °C with a salinity of 33.5 PSU and a chlorophyll a fluorescence of 2.7 μg l−1 (SIO Automated Shore station equipment, www.sccoos.org). The sample was filtered (0.8 μm pore size), amended with glycerol (final concentration 15% v/v), flash frozen and stored at −80 °C. Prior to sorting, the sample was thawed and stained with SYBR Green I nucleic acid stain (at 10 × , Invitrogen, Carlbad, CA, USA). Single cells were sorted using a FACS Aria II flow cytometer equipped with a 488 nm laser and custom forward scatter (FSC-PMT) (BD Biosciences, San Jose, CA, USA) using detection by the side scatter (SSC-PMT) and green fluorescence (512 nm), with the highest purity setting and the lowest flow rate to avoid sorting of coincident events. Phosphate-buffered saline, sterilized by 0.2 μm filtration, was used as sheath fluid. Single cells were sorted into 384-well plates containing 4 μl of TE (10 mM Tris, 0.1 mM EDTA, pH 8.0) buffer in each well and stored at −80 °C until MDA. MDA is described in the Supplementary Material, as is PCR screening of MDA reactions. Two MDAs with 16S sequences with >97% nucleotide identity to SAR86 were selected for 454 sequencing.
Preparation of pyrosequencing libraries
FLX Titanium 3 kb paired end libraries were generated and sequenced using 10 μg of pooled, re-amplified MDA as template according to manufacturer's specifications (Life Technologies, Bradford, PA, USA). Two single cell MDA reactions were barcoded and sequenced on the same plate, generating 700 000+ reads of >200 bp.
Assembly of SAR86 single cell genomes
Data derived from single cells should confirm the gene content found in the metagenomic assemblies and could potentially provide information about the variable and hyper-variables segments associated with microbial genomes that cannot be easily acquired from metagenomic assemblies. Assembly of the single cells was carried out using the Celera Assembler available at http://sourceforge.net/apps/mediawiki/wgs-assembler with error cutoffs set to 0.05, a word size of 14, and the bog unitiger. Assemblies will be deposited to the NCBI genome project database before the publication.
GOS metagenomic dataset
The data set consists of 10.97 million reads, of which 8.4 million were produced using Sanger sequencing and 2.54 million were produced using 454 Titanium sequencing. All sequence data used here are available at NCBI (under project ID 13694) and have also been submitted to CAMERA (Seshadri et al., 2007). The Sanger data used here include published data from the following GOS sites: GS00b, GS00c, GS00d, GS01a, GS01b, GS01c, GS02, GS03, GS04, GS05, GS06, GS07, GS08, GS09, GS10, GS13, GS14, GS15, GS16, GS17, GS18, GS19, GS21, GS22, GS23, GS25, GS26, GS27, GS28, GS29, GS30, GS31, GS34, GS35, GS36, GS37 and GS51. For the purposes of this study, we excluded the GS00a sample from the analysis because it contained an exceptionally high abundance of the non-planktonic Burkholderia and Shewanella-like genomic sequences (Rusch et al., 2007).
The following recently published Sanger and 454 GOS data (Rusch et al., 2010; Yooseph et al., 2010) are also used here. Sanger data from: GS41-GS47, GS48, GS49, GS108, GS109, GS110, GS111, GS112, GS113, GS114, GS115, GS116, GS117, GS119, GS120, GS121, GS122, GS123, GS148 and GS149. 454 pyrosequencing data sets were generated from 0.1 μm filters for GS108 (library identifier GOS108XLRVAL-4F-1-400) and GS112 (library identifier GOS112XLRVAL-4F-1-400).
Assembly and binning of SAR86 metagenomes
A global assembly of the GOS dataset was performed with the Celera Assembler. Metagenomic assemblies were executed aggressively by using default parameters, except error cutoffs were increased to 15% and the utgGenomeSize set to 150 000 bps as described previously (Rusch et al., 2007). Some of these scaffolds were highly similar in gene organization (synteny) with previously sequenced SAR86 BAC sequences and at 60–70% nucleotide identity implied that these scaffolds were closely related to the SAR86 lineage. In an effort to isolate more of the SAR86 genomes, we constructed proportional sample distribution of reads recruited above 90% identity for all the GOS scaffolds over 5 kbp in length. This distribution was used to calculate a distance matrix (using the R statistical package) between all the GOS scaffolds. Manual inspection of scaffolds closest to the already identified SAR86B scaffolds produced a set of 36 scaffolds that had a consistent distribution with the scaffolds that were syntenic with known SAR86 BACs. The total span of these scaffolds was under 2 Mbp, which was consistent with the sizes of other known planktonic microbes. In an effort to find more complete assemblies and order and orient these smaller putative SAR86 scaffolds, we searched for other GOS scaffolds that were syntenic to these putative SAR86 scaffolds. As many scaffolds were identified, two much larger scaffolds (806 kb and 443 kb) were syntenic with the previously identified putative SAR86 scaffolds and known SAR86 BACs while sharing a similar sample distribution and coverage characteristics with each other. We were unable to discover any other scaffolds that had similar coverage and sample distribution with this second set of putative SAR86 scaffolds, suggesting that any missing sequence is either small or associated with variable regions that would not be identified using these approaches. The two long SAR86 scaffolds and 36 smaller contigs were used to largely order and orient each other, producing a meta-scaffold that we designated SAR86A. A separate set of SAR86-like scaffolds were identified by their unique GOS sample distribution and binned to produce the SAR86B genome. Reads recruited to the SAR86B scaffolds are derived almost from exclusively GS-100 and GS-102 and to a lesser extent from GS-148 and GS-149 (Figure 2c). The validation of the assemblies is discussed in the results and the Supplementary Material.
The assemblies and annotation are deposited in the NCBI genome project collection under accession numbers AHBG00000000–AHBJ00000000.
Genome completeness estimates
Using the Comprehensive Microbial Resource as a database, 107 hidden Markov models (HMMs) that hit only one gene in greater than 95% of bacterial genomes were identified (Supplementary Table S1). Trusted cutoff scores for the TIGRFAMs and Pfam HMMs were those supplied by the TIGRFAMs and Pfam libraries (Haft et al., 2003; Finn et al., 2010). These HMMs were then used to search the SAR86 genomes, and the percentage of the total found was extrapolated for the entire genome.
Phylogenetic analyses
All predicted proteins from 196 completed γ-proteobacteria genomes, 37 marine γ-proteobacteria genomes from the Gordon and Betty Moore Marine Microbial Genome project (Yooseph et al., 2010) and the SAR86 assemblies were used as a starting dataset. The non-γ-proteobacterial genomes Ralstonia solanacearum GMI1000, Chromobacterium violaceum ATCC 12472, Rhodospirillum rubrum ATCC 11170 and Sinorhizobium meliloti 1021 were included as proteobacterial outgroups. The AMPHORA HMMs were used to screen for 31 single-copy proteins that are useful as phylogenetic markers, as identified by Wu and Eisen. In the case of duplicate genes, the HMM scores were used to find the best match. Each genus was trimmed to a single representative with the greatest number of markers found and, in the cases where multiple best options were available; preference was given to closed genomes. For the remaining genomes, markers aligned to the HMMs, and the resulting alignments were refined with MUSCLE (Edgar, 2004) and visual inspection. It was not possible to use all 31 AMPHORA markers, as a mosaic of missing markers exists across γ-proteobacterial genomes. Markers were discarded if there was no representative found for either one of the SAR metagenomic assemblies or for a phylogenetically close clade. Remaining markers’ alignments were judged based on the quality of the alignment and a resulting single protein maximum likelihood phylogeny. Markers that generated gapped alignments or aberrant phylogenies (mostly internal placement of non-γ-proteobacteria outgroups) were discarded. In the end, 12 markers were concatenated to create a global alignment; rplB, rplD, rplE, rplF, rplL, rplP, rplS, rpoB, rpsE, rpsI, rpsS and tsf. A maximum likelihood phylogeny was generated using Approximate Likelihood Ratio Test PhyML.
Abundance calculations for SAR86
As per Rusch et al. (2010), the relative abundance of each genome in each GOS sample was derived from the fragment recruitment data for reads that recruited to the given assembly at 90% identity or greater. The final value is reported as percentage of reads from a sample recruited per Mbp of reference sequence. These numbers are derived from the number of reads recruited to the reference after normalizing for the length of the reference that was observed and the relative number of reads collected per sample.
Genome comparisons and annotation
The SAR86A and B genomes were annotated using the Prokaryotic Annotation Pipeline (Davidsen et al., 2010), BLAST, KASS (Moriya et al., 2007) and signalP (Emanuelsson et al., 2007). Approximately 75% of the proteins in SAR86A were manually curated and orthologs in the SAR86B-D genomes were identified using a best blast hit approach using a non-redundant amino acid database from NCBI combined with the annotations of SAR86A.
A reciprocal best blast hit approach was used to examine the abundances of the SAR86 proteins in the GOS dataset. To determine the total abundance of SAR86 in each sample 96 proteins found in single copy in both SAR86A and B genomes were counted using reciprocal best blast hit. To determine the relative proportion of the total population containing specific biosynthetic pathways, the abundance of the homologs found in the SAR86A and B genomes was determined using reciprocal best blast hit.
16S phylogenetic analysis
Infernal (Nawrocki et al., 2009), a sequence- and structure-based alignment program, was used to align the 16S sequences using a bacterial alignment model obtained from the Ribosomal Database Project (Cole et al., 2009). Alignment columns with >90% gaps were removed and the subsequent multiple sequence alignment was used to construct a maximum likelihood phylogeny using RAxML (Stamatakis, 2006).
Results and discussion
Overview of genome assemblies
Metagenomic assembly and single-cell sequencing provide a cultivation-independent approach to generating genome sequences (Woyke et al., 2009; Rusch et al., 2010; Hess et al., 2011), and the two techniques were used to produce four SAR86 assemblies. With GOS samples as a metagenomic dataset, a combination of aggressive assembly followed by binning based on sample distribution, similarity to SAR86 BACs, oligonucleotide frequencies and manual curation resulted in the assembly of two nearly complete genomes (SAR86A and B). The SAR86A consensus genome consists of two scaffolds in 41 contigs with a total length of 1.25 Mbp. The SAR86B consensus genome consists of 31 scaffolds and is substantially larger at 1.70 Mbp. The two genomes contain 1316 and 1712 open reading frames, respectively (Table 1). These consensus genomes are not equivalent to assemblies from a clonal isolate, thus single cell techniques (Raghunathan et al., 2005; Lasken, 2007) were used to acquire further data. Sequences from each amplified single cell should represent an independent assessment of the gene content, and furthermore should not suffer from the biases and limitations that might be introduced in a metagenomic assembly. Single cells were isolated from coastal waters (San Diego, CA, USA) using flow cytometry, followed by amplification of genomic DNA with MDA (Dean et al., 2001, 2002). Two of the amplified genomes identified as SAR86 by 16S rRNA sequencing were further sequenced using Titanium 454 pyrosequencing. This resulted in two partial genomes (SAR86C and D) with a total length of 750 and 925 Kbp split among 142 and 194 contigs, respectively (Table 1). All four genomes have low proportion of guanine-cytosine (%GC).
Several analyses show that the metagenomic assemblies represent naturally occurring SAR86 populations in terms of gene content and genome structure. Pairwise alignments reveal substantial similarity in genome content and organization or synteny between the metagenomic assemblies and large (20 kbp+) SAR86 genome fragments acquired using single cell methods or molecular cloning (Supplementary Figure S1). While not insightful to genome structure, the smaller single cell contigs contain genes found on the metagenomic assemblies, establishing a consistent gene content. Recruitment of metagenomic reads to the assemblies is uniform over the length of the SAR86 genomes in terms of depth of coverage and the frequency of recruited reads among the different samples (Figure 1). Greater than 90% of the high identity Sanger mate pairs were recruited in the proper orientation and distance (Figure 1). Finally, scatter plots of the first three components of a principle component analysis of tetranucleotide usage form tight clusters that are consistent with a single genomic source ((Teeling et al., 2004), Supplementary Figure S2).
Recruitment of Global Ocean Sampling metagenomic data to the SAR86 assembled genomes. In the top panels, only the metagenomic reads that aligned best to the SAR86A or SAR86B reference (to the exclusion of all complete and draft microbial and viral genomes available at NCBI and each other) are shown as a dot whose color is determined by metagenomic sample from which it was identified. The pattern of recruitment of reads at greater than 90% across the entire genome (note that artificial gaps have been introduced between any two scaffolds) is largely consistent over the length of all scaffolds, and is qualitatively similar to recruitment plots seen for complete genomes (Rusch et al., 2007). In the middle panels, mate-pairing information is presented to indicate the wealth of information supporting the orientation of the assembled scaffolds. In these plots lines have been drawn connecting two good mated reads, where good is defined as mates in the correct orientation and that are separated by no more and no less than two standard deviations from the expected insert distance. The ends of the lines indicate the percent identity of each mate to the reference genome. The bottom panels indicate the number of good mate pairs found for each base pair of the genome assembly (coverage). Note that the gaps between contigs will reduce the number of good mate pairs, driving down coverage for the SAR86B genome. Based on our understanding of the Celera Assembler, we interpret these plots as indicating that these assemblies are the best-supported layout of the contigs given the available information. As these assemblies represent data from many different cells, it is possible that there are other valid layouts that are less prevalent in the data.
To estimate actual genome size from our partial genomes, a catalog of 107 single copy genes, including nearly all ribosomal proteins and tRNA synthases found in nearly all free-living bacteria, was compiled using the Comprehensive Microbial Resource (see methods). A total of 100 and 99 of these were found spread across the SAR86A and B genomes, respectively, suggesting they are greater than 90% complete. Seven of the missing proteins are the same: the signal recognition protein, signal recognition docking protein, initiation factor 3, ribosomal proteins L20 and L35, and both subunits of phenylalanyl tRNA synthase. SAR86B appears to lack dimethyladenosine transferase, though a manual search detected a putative protein that just missed the HMM cutoff for ksgA in Supplementary Table S1. The same seven proteins were not found in SAR86C and D, which are estimated to be 54% and 48% complete, respectively (Table 1). Each genome contains one complete rRNA loci. While tentative, it is possible that SAR86 has dispensed with each of these seven proteins; for example, as the signal recognition proteins were thought to be essential, this has been recently challenged (Hasona et al., 2005).
In abundant marine bacteria, such as SAR11, Prochlorococcus and Synechococcus, high variability of gene content at several locations comprising ∼10% of the genomes is expected to prevent complete assembly from metagenomic datasets (Rusch et al., 2007, 2010). Based on an examination of mated reads recruited to ends of the SAR86A scaffolds, there is at least one hypervariable region in natural SAR86A genomes. The high variability results in low coverage, thus preventing both assembly of the hypervariable region and circularization of the two scaffolds. Unfortunately, the single cells contigs do not extend into or clarify this hypervariable region.
Phylogenetic and biogeographic characterization of the genomes assemblies
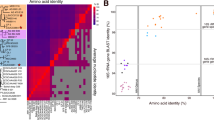
The 16S rRNA sequences from the SAR86 A–D group in SAR86 clusters I and IIa are >98% similar to each other in nucleotide sequence (Figure 2a). The 16S rRNA sequence cannot resolve global phylogenies of γ-proteobacteria (22) or even SAR86 confidently (Figure 2a), thus conserved proteins were used to construct maximum likelihood phylogenies (Figure 2b). A 12 protein phylogeny recapitulated the topology of a recent multi-protein phylogeny of γ-proteobacteria (Williams et al., 2010) while loosely pairing the SAR86A and B assemblies with Francisella tularensis as basal nodes (Supplementary Figure S3). A reduced seven-protein phylogeny allowed inclusion of the SAR86C and D genomes and retains a consistent global γ-proteobacterial topology (Figure 2b). The large phylogenetic distance from SAR86 to any cultivated γ-proteobacteria suggests a relatively ancient divergence or an artifact caused by rapid rates of evolution. For example, both SAR86 and F. tularensis exhibit metabolic streamlining (Larsson et al., 2005), which is accompanied by rapid protein sequence evolution (Dufresne et al., 2005). The loose association of these genomes in our phylogenies thus might be an artifact caused by long-branch attraction.
Phylogeny and emergent ecotypes of SAR86. (a) 16S rRNA RAxML phylogeny of the four SAR86 genome assemblies presented here, several BACs and fosmids, and the closest marine microbial genomes. Node values are bootstrap support for 100 iterations. (b) A maximum likelihood phylogeny of seven concatenated proteins found in nearly all γ-proteobacterial genomes and the four SAR86 genomes. Here, node values are approximate likelihood ratio test support for each branch point (values below 0.5 were removed). These values provide the probability (from 0–1) that such a branch point exists in the real tree.
Fragment recruitment (Rusch et al., 2007) is akin to in silico whole-genome DNA hybridization with a known nucleotide identity. The presence of genomes highly similar to the SAR86A assembly within multiple metagenomic datasets is confirmed by the abundance of reads at greater than 95% nucleotide identity (Figure 1). By visualizing only metagenomic reads that are a best match to SAR86A, a separate genome that could not be assembled can be observed at 80% nucleotide identity (SAR86A-like, Figure 1). Similar trends are observed for SAR86B (Figure 1). SAR86A and B are phylogenetically distinct, at least at the protein level, but we still have not captured the full diversity of even these individual SAR86 lineages.
The most abundant genomes in a GOS dataset of 10 million Sanger reads were determined using fragment recruitment at 90% and 50% nucleotide identity (Table 2). The SAR86 assemblies recruit more of the GOS dataset than any other non-photosynthetic bacteria except organisms in the SAR11 clade (Pelagibacteraceae) (Table 2), including the flavobacteria genomes sequenced by Woyke et al. (2009). At 50% nucleotide identity, the SAR86 and Pelagibacter genomes recruit ∼50 × more metagenomic data than at 90% nucleotide identity. This is in sharp contrast to Prochlorococcus and Synechococcus genomes where a relaxation of the identity cutoff only increased recruitment 10–50% (Table 2). Our interpretation is that there are numerous and abundant subclades of SAR86 and Pelagibacteraceae for which genomic representation is lacking.
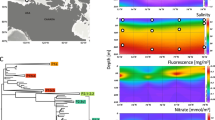
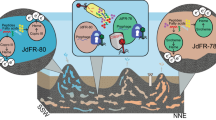
Across the GOS dataset, the four genomes exhibit a biogeography that may be related to physiology (Figure 3, Supplementary Table S2). For example, the two most related genomes, SAR86 C and D, are found at colder coastal sites, consistent with their isolation in coastal California, whereas SAR86A is found at all open ocean locations (Figure 3). SAR86B appears to have a very specific geographic distribution as it only recruits metagenomes collected from a small subset of warm coastal sites, specifically Zanzibar and the Gulf of Panama.
Emergent ecotypes of SAR86: recruitment of metagenomic reads at 90% nucleotide identity to each SAR86 assembly is shown for 73 GOS metagenomes. Also shown is a trace of the seawater temperature measured at the time of sampling and the geographic region. The complete dataset is available in Supplementary Table S2.
Metabolic streamlining in SAR86
Metabolic analyses of genomic information can provide information on the physiological capabilities of an organism, though conclusions are speculative even when a strain is cultivated and a completely finished genome is available. However, the hypotheses presented will certainly be useful for future physiological experiments conducted with a cultivated strain or natural microbial communities. For the sake of a comprehensive analysis, we will note when genes or pathways were not found, though only when the trends are consistent across all four genomes or genome alignments provide ancillary evidence of absence. The combined datasets catalog a portion of the SAR86 core genome and some of accessory proteins associated with specific lineages. Due to the cohabitation of the planktonic fraction on the surface ocean (Table 2), many of the putative features of SAR86 are directly compared with Pelagibacteraceae (Figure 4).
Metabolism of SAR86. A simplified and non-comprehensive schematic showing the salient metabolic features of SAR86 discussed in the text and determined with manual curations. Metabolic pathways where the genes are missing from SAR11 genomes are shown in red. DMSP, dimethylsulfoniopropionate; GBT, glycine betaine; GPX, glutathione peroxidase; GSH, glutathione.
A loss of biosynthetic pathways was observed in the Pelagibacteraceae Candidatus Pelagibacter ubique, presumably resulting from metabolic streamlining as a way to reduce nutrient requirements in the oligotrophic ocean (Giovannoni et al., 2005). Consistent with the modest size and low %GC of SAR86 genomes, all the genes in several vitamin biosynthesis pathways, including B12, B6, biotin, pantothenate, thiamine and retinol, are absent from SAR86A, C and D. These pathways are also missing from SAR86B, with the exception of thiamine biosynthesis (Table 1). Failure to find any of the genes for the vitamin synthesis pathways in four different genomes (excepting thiamine) provides strong, but not conclusive, evidence for auxotrophy of these vitamins in SAR86. SAR86B contains a putative B12 transporter, consistent with auxotrophy, though transporters for the other vitamins could not be identified. It is also possible that alternative biosynthetic routes are used (Webb et al., 2007). The potential for vitamin auxotrophy should be considered in future cultivation efforts.
SAR86A lacks the proteins required for methionine (Met), histidine (His), and arginine (Arg) synthesis. SAR86B contains the genes for the synthesis of His and Arg within contigs that are otherwise syntenic to the SAR86A genome (Supplementary Figures S4a and b), suggesting the absence in SAR86A is not an artifact. It is possible that these proteins are found in different genomic locations of SAR86A that we did not recover. SAR86D also contains the His synthesis operon. The SAR86B and D amino-acid synthesis genes were used to estimate the presence of these pathways in natural populations. The proteins for the Met, His and Arg synthesis pathways are always less abundant than a set of 107 core genes across 73 metagenomes after normalization (Supplementary Figure S4c). If most natural SAR86 populations retained the synthesis pathways, normalized recruitment of the genes for the amino-acid synthesis genes should be roughly equivalent to that of the core genes, which was not observed. Instead, it appears that substantial portions of natural SAR86 populations are auxotrophs for Met, His and Arg.
Each genome contains the demethylase dmdA that produces methyl-mercaptopropionate from the algal osmolyte dimethyl-sulfoniopropionate, providing reduced sulfur (Howard et al., 2006). Each genome also contains putative transporters for glycine-betaine that may facilitate DMSP uptake. In contrast to SAR11, all SAR86 genomes contain genes for putative transporters of glutathione (γ-glutamate-cysteine-glycine) and γ-glutamyl transferases, which are required to break the otherwise recalcitrant γ-bond in glutathione, expanding the diversity of organic sulfur available to SAR86 relative to SAR11 (Figure 4). The concentrations of dissolved glutathione and γ-Glu-Cys in the oligotrophic north Pacific reach 10–15 nM, with turnover times on the order of days to weeks (Dupont et al., 2006). These known metal-binding compounds likely influence Cu, Fe and Hg speciation; thus, consumption of this pool by SAR86 adds an additional dimension to models of trace metal biogeochemistry. All of the various SAR86 assemblies lack the enzymes required for sulfate uptake or assimilatory reduction, as was also observed in SAR11 (Tripp et al., 2008).
SAR86B does have a much larger genome than SAR86A. Many of the SAR86B-specific genes could not be functionally annotated, though there are expansions in the numbers of TonB receptors, ABC transporters and beta-lactamases (Table 1). Several glycosyl hydrolases found in the SAR86A genome are also duplicated in the SAR86B genome.
Proteorhodopsin phototrophy
Proteorhodopsin was originally found on a BAC of SAR86 origin and, when heterologously expressed in E. coli and provided with exogenous retinol, functioned as a light-driven proton pump (Beja et al., 2000). In some marine bacteria, proteorhodopsin can facilitate energy generation under nutrient-limiting conditions (Steinder et al., 2011). SAR86A and C contain one putative green-light tuned proteorhodopsin, whereas SAR86B and D each contain two. Autotrophic carbon-fixation pathways are lacking in the SAR86 genomes, so the proteorhodopsin-generated pH gradient across the cytoplasmic membrane may be used for phosphorylation or transport. Proteorhodopsin requires retinol for functionality, and in many proteorhodopsin-containing marine BACs and genomes, genes for pigment synthesis are colocalized with proteorhodopsin and exhibit parallel phylogenetic differentiation (McCarren and DeLong, 2007). Thus, it is surprising that the five proteins for the retinol biosynthesis pathway are lacking from all four SAR86 genome assemblies and the numerous SAR86 BACs and fosmids. Like some SAR86 BACs (Sabehi et al., 2004, 2005), the proteorhodopsins in SAR86 A-D are flanked by a short-chain dehydrogenase gene that might be used to convert retinal or β-carotene to retinol. This could only catalyze the conversion of an already synthesized hydrophobic pigment, with the initial five steps of carotenoid synthesis missing. Either retinol biosynthesis pathways are part of hypervariable genomic regions, making them rare, or SAR86 must scavenge retinol or a structurally related pigment. Retinol uptake pathways are unknown, as are the concentrations in seawater.
Carbon metabolism in SAR86
All four genomes contain the core components for aerobic respiration and lack the proteins required for carbon fixation via the reverse tricarboxylic acid (TCA) cycle, the reductive CoA pathway and the 2-hydroxypropionate cycle (Figure 4). The genomes do not contain nitrate reductase, nitrite reductase, sulfite reductase or the cytochromes typically involved in anaerobic metabolism like c3 and b1. Thus, SAR86 appears to be an aerobic heterotroph with the potential for phototrophic ATP production via proteorhodopsin. SAR86A, B and D contain a full complement of the TCA cycle proteins except for citric acid synthase. Instead, genes coding for 2-methylcitrate synthase, methylcitrate lyase and methylcitrate dehydrogenase are present. 2-Methylcitrate synthase can catalyze citrate synthesis from acetyl-CoA and methylcitrate synthesis from propionyl-CoA, which are derived from even and odd length fatty acids, respectively. Thus, SAR86 likely uses a dual TCA/methylCA cycle (Figure 4). An elegant parallel is the overabundance of lipases and the enzymes required to catalyze the beta-oxidation of fatty acids (Table 1), providing NADH, acetyl-CoA and propionyl-CoA.
The SAR86A, B and D genomes contain a complete Emden–Meyerhof–Parnas glycolysis pathway. SAR86B contains a complete pentose phosphate pathway, but SAR86A lacks the oxidative arm of the pathway (Figure 5). As with the amino-acid synthesis pathways, this does not appear to be an artifact, as the corresponding proteins in SAR86B occur in a genomic locale syntenic to that of SAR86A (Figure 5). In addition to the pentose phosphate genes, this genomic region codes for critical steps in glycolysis and glucose uptake. Across 73 oceanic metagenomes, the shared genes involved in glycolysis and glucose uptake are always more abundant than those coding for the oxidative arm of the pentose phosphate pathway, implying that substantial portions of natural SAR86 populations lack the oxidative pentose phosphate pathway (Figure 5). This would result in one less metabolic source of NADH production. An analogous scenario is observed in the marine cyanobacteria, where gain and loss of proteins within one genomic location results in dramatically different nitrogen assimilation capabilities among different lineages (Scanlan et al., 2009). This genomic region contains an abundant non-coding RNA identified in marine metatranscriptomic libraries that previously lacked genomic context (Shi et al., 2009) (Figure 5, Supplementary Figure S5). This implies that carbon assimilation in SAR86 is controlled by a rapidly responsive RNA-based regulation. The expanded sugar utilization metabolism in SAR86 contrasts sharply to SAR11, where some strains lack glycolysis altogether and others contain a modified Entner–Duodroff pathway (Schwalbach et al., 2010).
SAR86 genome and population diversity of carbon assimilation. (a) Alignments of orthologs regions of the SAR86A and B genomes are shown to scale. Pink lines join the orthologs from each genome. The small green ‘genes’ and connecting green lines indicate the position of the non-coding RNA found in metatranscriptomic datasets. An alignment of this small RNA is shown in Supplementary Figure S5. (b) The abundances of the genes found in both genomes (glycolysis or glucose uptake) or just the SAR86B genome across 73 GOS metagenomes. The shared genes are clearly conserved, yet many genomes in natural populations lack the oxidative arm of the pentose phosphate pathway or the genes involved in pyruvate metabolism found in only SAR86B. The units on the axes are the number of reciprocal best BLAST hits to either 107 core genes (x axis) or the genes shown in the left panel (y axis), followed by normalization for the number of genes in that category.
In addition to sugars and lipids, all SAR86 may be able depolymerize polysaccharides with glycoside and glycosyl hydrolases and degrade peptidoglycan into D-amino acids and D-sugars using a set of murein lytic hydrolases, D,D-carboxypeptidases and D,L amidases (Figure 4). The conversion to L-amino acids could be catalyzed by the two D-amino-acid racemases. Relative to SAR86A, SAR86B also has an additional D-sugar racemase, an β-agarase and several extra glycosyl/glycoside hydrolases, consistent with its genomic expansion in sugar utilization. SAR86 appears to have an incomplete gluconeogenesis pathway; all genomes lack fructose-1,6-bisphosphatase but contain 6-phosphofructokinase, which would prevent de-phosphorylation of 1,6-P-fructose into 6-P-fructose but allow the reverse reaction.
A focus on pmf-dependent transport across the outer membrane
Whereas the SAR11 genomes contain numerous substrate-binding protein ABC-type transporters for nutrient uptake across the cytoplasmic membrane, SAR86 contains only two (Table 1), specifically for oligo-peptides, which is consistent with amino acid auxotrophy, and ferric iron. All of the SAR86 genomes contain multiple major facilitator superfamily type transporters for simple metabolites, ammonium and phosphate. When considered with respect to genome size, the SAR86 genomes contain a highly disproportionate number of putative tonB-dependent outer membrane receptors (TBDR) relative to other bacteria (Table 1, Supplementary Figure S6). TBDRs are outer membrane receptors that catalyze high affinity transport of compounds larger than 600 Da across the outer membrane, including Fe-, Cu- and Ni-chelates, vitamin B12, N-acetyl-glucosamine, and carbohydrates (Blanvillian et al., 2008; Schauer et al., 2008).
TBDR-dependent transport requires a pmf across the cytoplasmic membrane. As noted by Morris et al. (2010), proteorhodopsin may provide a pmf for TBDR uptake. SAR86 has other mechanisms for generating a pmf, including respiration or NADH dehydrogenation (Figure 4). All four genomes contain multiple V-type pyrophosphatases, which generate a pmf through the breakdown of cytoplasmic pyrophosphates (Figure 4), and are important during shifts between starvation-to-nutrient-replete and dark-to-light conditions in the non-purple sulfur bacteria (Garcia-Contreras et al., 2004). Respiration is likely the dominant pathway for generating a pmf, with proteorhodopsin, NADH dehydrogenation and pyrophosphate breakdown providing ancillary support.
Although the vast majority of TBDRs are uncharacterized and phylogenetic analyses are uninformative about substrate specificity due to poor bootstrap support (Schauer et al., 2008), genome neighborhood analysis can provide insight. Many of the SAR86 TBDR genomic regions contain lipid-degrading enzymes, a phenomenon not previously observed in any organism, much less marine bacteria (Supplementary Figure S7). For example, one genomic locus includes a patatin (a phopholipase), a protein in the rhodanese/beta-lactamase family, and a flavin-dependent oxidoreductase. While the patatin might break a bond found in phospholipids, the other two enzymes do not. Instead, this enzymatic combination is potentially capable of breaking key bonds in sulfoquinovosyldiacylglycerol, a sulfolipid used by marine cyanobacteria (Van Mooy et al., 2009). After cytoplasmic uptake, the sulfoquinovose polar group can be degraded by the TCA cycle, recovering more ATP and NADH (Roy et al., 2003). A combination of esterase family proteins and a choline/carnitine/betaine transportor could potentially catalyze fatty acid removal and subsequent cytoplasmic uptake of polar head groups from betaine polar lipids. In the hyper-oligotrophic South Pacific, combined sulfoquinovosyldiacylglycerol and betaine polar lipids concentrations are typically >1.5 nM, each molecule containing >30 acyl bonds, making them highly energy rich (Van Mooy and Fredricks, 2010). Notably, the tonB loci in each genome contain different combinations of catabolic enzymes; diversification of carbon compound transport may drive the high phylogenetic diversity of SAR86.
Antibiotics gain access to the periplasm of gram-negative bacteria via non-specific TBDR uptake (Schauer et al., 2008), thus the presence of a disproportionate number of β-lactamases and a macrolide efflux system is not surprising (Table 1). All of the genomes also contain at least one cytochrome P450 that may be used for xenobiotic degradation. In contrast to all other abundant cultivated marine microbes, each genome encodes putative nitroreductases and nitropropane dioxygenases (Table 1), proteins generally associated with bacteria found in soils contaminated with industrial chemicals like trinitrotoluene (dynamite). Nitropropane dioxygenases degrade nitro-aromatic compounds, producing nitrate and nitrite in the process (Nishino et al., 2010). Nitroreductases degrade the same compounds but generate nitrite and ammonium. There is no evidence that SAR86 can subsequently assimilate the nitrate or nitrite. The presence of these enzymes within four genomes of a highly abundant bacterial clade suggests that nitro-aromatics are a biologically relevant and heretofore overlooked portion of the dissolved organic matter in the surface ocean. Potentially, the addition of nitroaromatics would also select for SAR86 over the more abundant SAR11 during cultivation efforts.
Carbon source specialization in metabolically streamlined planktonic bacteria
The ultimate source of dissolved organic carbon in the open ocean is planktonic biomass, which, in terms of carbon, is 65±9% protein, 19±4% carbohydrate and 16±6% lipid (Hedges et al., 2002). This material is released upon death, either by viral lysis, apoptosis or predation, creating dissolved and particulate organic carbon pools. Several abundant marine genera with TBDR-rich genomes have been implicated in the colonization and degradation of marine particles (Bauer et al., 2006; Thomas et al., 2008), fueling further production of dissolved organic carbon (Azam and Long, 2001). However, the SAR86 genomes lack the genes required for pili or flagellin formation, chemotaxis and motility, EPS production and other pathways known to mediate particle adhesion. This, along with its prevalence in metagenomes from the 0.1–0.8 μm size fraction, implies that SAR86 is predominantly free living (planktonic).
The two most abundant identifiable genomes of heterotrophic planktonic bacteria within the GOS dataset are SAR11 and SAR86 (Table 2). Despite a cosmopolitan distribution, neither of these organisms is a generalist; each exhibits genome and metabolic streamlining that precludes using all carbon compounds. Consistent with genomic predictions (Giovannoni et al., 2005; Schwalbach et al., 2010), cultivated strains of SAR11 can grow on dicarboxylic acids and simple peptides, a pool of organic carbon derived from upwards of 85% of cellular carbon biomass. SAR86 appears to specialize in the transport and degradation of lipids and polysaccharides. By focusing on different compounds, SAR11 and SAR86 might compete only sparingly for dissolved organic carbon. This also provides a tangible link between the crude biochemical composition of the dominant phytoplankton and the associated bacterial community, specifically that the relative abundance of SAR11 and SAR86 is controlled by the stoichiometry of protein, carbohydrate and lipid in plankton.
Accession codes
References
Azam F, Long RA . (2001). Oceanography: sea snow microcosms. Nature 414: 495–498.
Bauer M, Kube M, Teeling H, Richter M, Lombardot T, Allers E et al. (2006). Whole genome analysis of the marine Bacteroidetes ‘Gramella forsetii’ reveals adaptations to degradation of polymeric organic matter. Environ Microbiol 8: 2201–2213.
Beja O, Aravind L, Koonin EV, Suzuki M, Hadd A, Nguyen LP et al. (2000). Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science 289: 1902–1906.
Blanvillian S, Meyer D, Boulanger A, Lautier M, Guynet C, Denance N et al. (2008). Plant carbohydrate scavenging through Ton-B dependent receptors: a feature shared by phytopathogenic and aquatic bacteria. PloS One 2: e224.
Britschgi TB, Giovannoni SJ . (1991). Phylogenetic analysis of a natural bacterioplankton population by rRNA gene cloning and sequencing. Appl Environ Microbiol 57: 1707–1713.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Davidsen T, Beck E, Ganapathy A, Montgomery R, Zafar N, Yang Q et al. (2010). The comprehensive microbial resource. Nucleic Acids Res 38: d340–d345.
Dean FB, Hosono S, Fang L, Wu X, Faruqi AF, Bray-Ward P et al. (2002). Comprehensive human genome amplification using multiple displacement amplification. Proc Natl Acad Sci USA 99: 5261–5266.
Dean FB, Nelson, Giesler TL, Lasken RS . (2001). Rapid amplification of plasmid and phage DNA using Phi 29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res 11: 1095–1099.
DeLong EF, Beja O . (2010). The light-driven proton pump proteorhodopsin enhances bacterial survival during tough times. PLoS Biol 9: e1000359.
Dufresne A, Garczarek L, Partensky F . (2005). Accelerated evolution associated with genome reduction in a free-living prokaryote. Genome Biol 6: R14.
Dupont CL, Moffett JW, Ahner BA . (2006). Distributions of dissolved and particulate thiols in the sub-artic north Pacific. Deep Sea Res I 57: 553–566.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Emanuelsson O, Brunak S, vonHeijne G, Nielsen H . (2007). Locating proteins in the cell using TargetP, SignalP, and related tools. Nat Protoc 2: 953–971.
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE et al. (2010). The Pfam protein families database. Nucleic Acids Res 38: D211–D222.
Garcia-Contreras R, Celis H, Romero I . (2004). Importance of Rhodospirillum rubrum H-pyrophosphatase under low energy conditions. J Bacteriol 186: 6651–6655.
Giovannoni SJ, Tripps HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Gonzalez JM, Simo R, Massana R, Covert JS, Casamayor EO, Pedro-Alio C et al. (2000). Bacterial community structure associated with a dimethylsulfoniopropionate-producing North Atlantic algal bloom. Appl Environ Microbiol 66: 4237–4246.
Haft DH, Selengut JD, White GF . (2003). The TIGRFAMs database of protein families. Nucleic Acids Res 31: 371–373.
Hasona A, Crowley PJ, Levesque CM, Mair RW, Cvitkovitch DG, Bleiweis AS et al. (2005). Streptococcal viability and diminished stress tolerance in mutants lacking the signal recognition particle pathway or YidC2. Proc Natl Acad Sci USA 102: 17466–17471.
Hedges JI, Baldock JA, Gelinas Y, Lee C, Peterson ML, Wakeham SG . (2002). The biochemical and elemental compositions of marine plankton: a NMR perspective. Mar Chem 78: 47–63.
Hess M, Sczyrba A, Egan R, Kim T-W, Chokhawala H, Schroth G et al. (2011). Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331: 467.
Howard EC, Henriksen JR, Buchan A, Reisch CR, Burgmann H, Welsh R et al. (2006). Bacterial taxa that limit sulfur flux from the ocean. Science 314: 649–652.
Larsson P, Oyston PCF, Chain P, Chu MC, Duffield M, Fuxelius H et al. (2005). The complete genome sequence of Francisella tularensis, the causative agent of tularemia. Nat Genet 37: 153–159.
Lasken RS . (2007). Single cell genomic sequencing using Multiple Displacement Amplification. Curr Opin Microbiol 10: 1–7.
Malmstrom RR, Straza TRA, Cottrell MT, Kirchmann DL . (2007). Diversity, abundance, and biomass production of bacterial groups in the western Arctic Ocean. Aquat Microb Ecol 47: 45–55.
Mary I, Cummings DG, Biegala IC, Burkill PH, Archer SD, Zubkov MV . (2006). Seasonal dynamics of bacterioplankton community structure at a coastal station in the western English Channel. Aquat Microb Ecol 42: 119–126.
McCarren J, DeLong EF . (2007). Proteorhodopsin photosystem gene clusters exhibit co-evolutionary trends and shared ancestry among diverse marine microbial phyla. Environ Microbiol 9: 846–858.
Moriya Y, Itoh M, Okuda S, Yoshizawa A, Kanehisa M . (2007). KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35: W182–W185.
Morris RM, Nunn BL, Frazer C, Goodlett DR, Ting YS, Rocap G . (2010). Comparative metaproteomics reveals ocean-scale shifts in microbial nutrient utilization and energy transduction. ISME J 4: 673–685.
Morris RM, Vergin KL, Cho J-C, Rappe MS, Carlson CA, Giovannoni SJ . (2005). Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time-series study site. Limnol Oceanogr 50: 1687–1696.
Mullins TD, Britschigi TB, Krest RL, Giovannoni SJ . (1995). Genetic comparisons reveal the same unknown bacterial lineages in Atlantic and Pacific bacterioplankton communities. Limnol Oceanography 40: 148–158.
Nawrocki EP, Kolbe DL, Eddy SR . (2009). Infernal 1.0: inference of RNA alignments. Bioinformatics 25: 1335–1337.
Nishino SF, Shin KA, Payne RB, Spain JC . (2010). Growth of bacteria on 3-nitropropionic acid as a sole source of carbon, nitrogen, and energy. Appl Environ Microbiol 76: 3590–3598.
Raghunathan A, Ferguson HR, Bornarrth CJ, Driscoll M, Lasken RS . (2005). Genomic DNA amplification from a single bacterium. Appl Environ Microbiol 71: 3342–3347.
Roy AB, Hewlins MJE, Ellis AJ, Harwood JL, White GF . (2003). Glycolytic breakdown of sulfoquinovose in bacteria: a missing link in the sulfur cycle. Appl Environ Microbiol 69: 6434–6441.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II global ocean sampling expedition: Northwest Atlantic through Eastern Tropical Pacific. PloS Biol 5: 398–431.
Rusch DB, Martiny AC, Dupont CL, Halpern AL, Venter JC . (2010). Characterization of Prochlorococcus clades from iron-depleted oceanic regions. Proc Natl Acad Sci 107: 16184–16189.
Sabehi G, Beja O, Suzuki MT, Preston CM, DeLong EF . (2004). Different SAR86 groups harbour divergent proteorhodopsins. Environ Microbiol 6: 903–910.
Sabehi G, Loy A, Jung K, Partha R, Spudich JL, Isaacson T et al. (2005). New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Biol 3: e273.
Scanlan DJ, Ostrowski M, Mazard S, Dufrense A, Garczarek L, Hess WR et al. (2009). Ecological genomics of marine picocyanobacteria. Microbiol Mol Biol Rev 73: 249–299.
Schattenhofer M, Fuchs BM, Amann R, Zubkov MV, Tarren GA, Pernthaler J . (2009). Latitudinal distribution of prokaryotic picoplankton populations in the Atlantic Ocean. Environ Microbiol 11: 2078–2093.
Schauer K, Rodionov DA, de Reuse H . (2008). New substrates for ton-B-dependent transport: do we only see the ‘tip of the iceberg’? Trends Biochem Sci 33: 330–338.
Schmidt TM, DeLong EF, Pace NR . (1991). Analysis of a marine picoplankton community by 16S rRNA gene cloning and sequencing. J Bacteriol 173: 4371–4378.
Schwalbach MS, Tripps HJ, Steindler L, Smith DP, Giovannoni SJ . (2010). The presence of the glycolysis operon in SAR11 genomes is positively correlated with ocean productivity. Environ Microbiol 12: 490–500.
Seshadri R, Kravitz SA, Smarr L, Gilna P, Frazier M . (2007). CAMERA: A community resource for metagenomics. PloS Biol 5: e75.
Shi Y, Tyson GW, DeLong EF . (2009). Metatranscriptomics reveal unique microbial small RNAs in the ocean's water column. Nature 459: 266–269.
Stamatakis A . (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.
Steinder L, Schwalbach MS, Smith DP, Chan F, Giovannoni SJ . (2011). Energy starved Candidatus Pelagibacter ubique substitutes light-mediated ATP production for endogenous carbon respiration. PLoS One 6: e19725.
Teeling H, Meyerdierks A, Bauer M, Amann R, Glockner FO . (2004). Application of tetranucleotide frequencies for the assignment of genomic fragments. Environ Microbiol 6: 938–947.
Thomas T, Evan FF, Schleheck D, Mai-Prochnow A, Burke C, Penesyan A et al. (2008). Analysis of the Pseudoalteromonas tunicata genome reveals properties of a surface-associated life style in the marine environment. PLoS One 3: e3252.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA et al. (2009). Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J 3: 1148–1163.
Tripp HJ, Kitner JB, Schwalbach MS, Dacey JWH, Wilheim LJ, Giovannoni SJ . (2008). SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452: 741–744.
Van Mooy BAS, Fredricks HF . (2010). Bacterial and eukaryotic intact polar lipids in the eastern subtropical South Pacific: water column distribution, planktonic sources, and fatty acid composition. Geochim Cosmochim Acta 74: 6499–6516.
Van Mooy BAS, Fredricks HF, Pedler BE, Dyhrman ST, Karl DM, Lomas MW et al. (2009). Phytoplankton in the ocean use non-phosphorus lipids in response to phosphorus scarcity. Nature 458: 69–72.
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA et al. (2004). Environmental genome shotgun sequencing of the Sargasso Sea. Science 304: 66–74.
Webb ME, Marquet A, Mendel RR, Rebeille F, Smith AG . (2007). Elucidating biosynthetic pathways for vitamins and cofactors. Nat Prod Rep 24: 988–1008.
Williams KP, Gillespie JJ, Sobral BWS, Nordberg EK, Snyder EE, Shallom JM et al. (2010). Phylogeny of gammaproteobacteria. J Bacteriol 192: 2305–2314.
Woyke T, Xie G, Copeland A, Gonzalez JM, Han C, Kiss H et al. (2009). Assembling the marine metagenome, one cell at a time. PloS One 4: e5299.
Wu M, Eisen JA . (2008). A simple, fast, and accurate method of phylogenomic inference. Genome Biol 9: R151.
Yooseph S, Nealson K, Rusch DB, McCrow JP, Dupont CL, Kim M et al. (2010). Genomic and functional characterization of planktonic marine prokaryotes. Nature 468: 60–66.
Acknowledgements
We acknowledge funding from the Gordon and Betty Moore Foundation (grant award agreement #521) and the US Department of Energy, Office of Science, Office of Biological and Environmental Research (DE-FC02-02ER63453). MN, JY-G, ML and RSL were funded by a grant from the Alfred P Sloan Foundation. A NASA Astrobiology Institute Directors Discretionary Fund supported CLD.
Author contributions
RF and JCV were responsible for the project design and management. DBR and ALH assembled the SAR86 metagenomes from the GOS metagenomes. AR, RV, SY and CLD performed the phylogenetic analyses and JDS and DH provided genome completeness estimates. MN, JY-G, M-JL and RSL isolated and sequenced the SAR86 single-cell genomes. CLD and DBR annotated all the genomes and analyzed the combined data sets. CLD wrote the paper with contributions from DBR, SY, M-JL and KN, and subsequent final review by all authors.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Dupont, C., Rusch, D., Yooseph, S. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J 6, 1186–1199 (2012). https://doi.org/10.1038/ismej.2011.189
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.189
Keywords
This article is cited by
-
VBCG: 20 validated bacterial core genes for phylogenomic analysis with high fidelity and resolution
Microbiome (2023)
-
Metagenome-assembled genomes reveal greatly expanded taxonomic and functional diversification of the abundant marine Roseobacter RCA cluster
Microbiome (2023)
-
A shotgun approach to explore the bacterial diversity and a brief insight into the glycoside hydrolases of Samiti lake located in the Eastern Himalayas
Journal of Genetic Engineering and Biotechnology (2022)
-
Constructing metagenome-assembled genomes for almost all components in a real bacterial consortium for binning benchmarking
BMC Genomics (2022)
-
Availability of vitamin B12 and its lower ligand intermediate α-ribazole impact prokaryotic and protist communities in oceanic systems
The ISME Journal (2022)