Abstract
In higher plants, the self-incompatibility mechanism is important for inhibition of self-fertilization and facilitation of out-crossing. In Brassicaceae, the self-incompatibility response is mediated by allele-specific interaction of the stigma-localized S-locus receptor kinase (SRK) with the pollen coat-localized ligand (SCR/SP11). All self-incompatible Brassicaceae plants analyzed have been found to have the SRK and SCR/SP11 genes in the S-locus region. Although Arabidopsis thaliana is self-compatible, transformation with functional SRK-SCR genes from self-incompatible Arabidopsis species confers the self-incompatibility phenotype to A. thaliana. The allele-specific interaction between SRK and SCR activates the downstream signaling cascade of self-incompatibility. Yeast two-hybrid analysis with a kinase domain of SRK as bait and genetic analysis suggested several candidate components of self-incompatibility signaling in Brassica. Recently, A. thaliana genes orthologous to the identified genes for Brassica self-incompatibility signaling were evaluated by using a self-incompatible transgenic A. thaliana plant and these orthologous genes were found not to be involved in self-incompatibility signaling in the transgenic A. thaliana. In this review, we describe common and different aspects of S-locus genomic regions and self-incompatibility signaling between Brassica and Arabidopsis.
Similar content being viewed by others
Introduction
Higher plants have a self-incompatibility mechanism for preventing self-fertilization and facilitating out-crossing. Self-incompatibility is considered to contribute to the maintenance of genetic diversity and avoidance of inbreeding depression. The Brassicaceae self-incompatibility system is well studied. Recognition specificity of this self-incompatibility system is determined by a diploid genotype of a parent plant. In self-pollination, pollen germination and pollen tube penetration of the cell wall of stigma papillar cells are inhibited.
Self-incompatibility is generally controlled by a single locus, the S locus. In Brassicaceae, the S-locus r eceptor k inase (SRK) and S -locus c ysteine r ich protein/S-locus p rotein 11 (SCR/SP11) genes, which encode highly polymorphic proteins as female and male determinants of recognition specificity, respectively, have been found at the S locus.1–3 Because these two genes are tightly linked with each other and inherited as a single Mendelian locus, a set of alleles of the S-locus genes is referred to as S haplotype.4 The SRK gene is expressed in the stigma papillar cells and encodes a plasma membrane-localized receptor kinase, which has a highly polymorphic extracellular receptor domain (S domain, hereafter) followed by a transmembrane domain and a serine/threonine kinase domain.1,5 Some variants of SRK exhibit more than 30% amino-acid sequence divergence in the S domain.6,7 SCR/SP11 (SCR hereafter) is expressed in anthers and its translational products are secreted to the pollen coat.8 SCR is a small peptide, ∼60 amino acids of mature form, and functions as the ligand for SRK.2,9,10 SCR is also highly polymorphic and less than 50% amino-acid sequence similarity is shared between S haplotypes.2,7,11–13 Although they have high sequence diversity, all SCR proteins appear to form a typical defensin-like 3D structure consisting of three β-sheets and one α-helix.14,15 In self-pollination, stigma-localized SRK interacts with SCR of the same S haplotype located on the pollen surface and activates a self-incompatibility signaling cascade, resulting in inhibition of self-pollen germination and tube penetration of the stigma papillar cell wall.
Arabidopsis thaliana, which is a model plant belonging to the family Brassicaceae, had not been used for studies of self-incompatibility mechanism because A. thaliana is a self-compatible species due to lack of functional SRK and/or SCR.16–21 However, transformation with functional SRK-SCR genes from self-incompatible Arabidopsis and closely related species, such as Arabidopsis lyrata, Arabidopsis halleri and Capsella gradiflora, confers self-incompatibility phenotype to A. thaliana,20,22–26 indicating that A. thaliana has the molecular components that are required for self-incompatibility signaling and can be used for studies of the Brassicaceae self-incompatibility mechanism.
The plant family Brassicaceae contains 338 genera and 3709 species, 308 of the 338 genera being assigned to 44 tribes.27,28 These tribes are grouped into three major linages.29–32 Arabidopsis and Brassica belong to lineage I and II, respectively,33 and these two genera were separated approximately 15 million years ago. Whole genome duplication or triplication has occurred only in the Brassica lineage but not in Arabidopsis since their separation. These observations suggest that Brassica and Arabidopsis would have different genetic backgrounds, although both self-incompatible plants of these two genera possess the SRK and SCR genes for recognition specificity of self-incompatibility. In this mini-review, we describe the molecular components functioning in SRK-mediated self-incompatibility signaling in Brassica and in planta evaluation results of these identified molecular components by using self-incompatible transgenic A. thaliana. We also discuss common and different aspects of self-incompatibility between Brassica and Arabidopsis.
The s-locus in Brassica and Arabidopsis
Although introduction of Arabidopsis SRK-SCR genes confers the self-incompatibility response to A. thaliana,20,22–26 construction of self-incompatible transgenic A. thaliana plants by introduction of the Brassica SRK-SCR gene pair has not succeeded.34 One possible explanation for this failure is that Brassica SRK and/or SCR are too greatly differentiated to function in Arabidopsis.
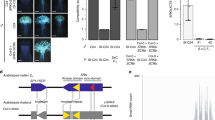
Molecular genetic studies have elucidated an interesting difference of the S-locus regions between Brassica and Arabidopsis. In Brassica, three genes are generally found in the S locus. In addition to the SRK and SCR genes, the S-l ocus g lycoprotein (SLG) gene is located at the S locus. The SLG gene encodes a stigma soluble glycoprotein showing high similarity to the S-domain of SRK. Like SRK, SLG is a highly polymorphic protein between S haplotypes. The role of SLG in self-incompatibility remains unclear. Because some S haplotypes lack the functional SLG gene at the S locus,35 the SLG gene is not considered to be an essential component in the self-incompatibility in Brassica. The S domain of SRK of a self-compatible Brassica rapa S-54 mutant has been found to be 100% identical to the S-54 SLG gene,36 suggesting that gene conversion between SRK S domain and SLG occurred, although this gene conversion caused the loss of the SRK function. This observation indicates one possible role of SLG in self-incompatibility, namely that the SLG gene contributes to production of a new SRK allele by gene conversion.
The SLG gene has not been found at the S locus of any A. lyrata S haplotypes. Instead of the SLG gene, the ARK3 gene, which is closely related to the SRK gene and contains the S domain, transmembrane domain and kinase domain, is located at the A. lyrata S locus. The ARK3 gene, as well as SRK and SCR, has been affected by positive selection.37 In addition, gene conversion between SRK and ARK3 was detected as observed in the Brassica S locus. This gene conversion occurred in the region of the kinase domain,37 and possibly functioned to promote SRK evolution to produce new substrate specificity.
Positive regulators in the self-incompatibility signaling pathway in Brassica
M-l ocus p rotein k inase (MLPK) has been identified in self-incompatible B. rapa plants as a positive regulator of Brassica self-incompatibility.38 MLPK is a protein kinase and a self-compatible mlpk mutant of B. rapa has a G194R mutation in its kinase domain.38 In vitro kinase assay has shown that MLPK has autophosphorylation activity, whereas MLPK protein of the self-compatible mlpk mutant has no activity.38 The MLPK gene produces two alternative transcripts (MLPKf1 and MLPKf2) from distinct transcription initiation sites.39 MLPKf1 and MLPKf2 produce proteins of 404 and 410 amino acids, respectively, and only their N-terminus sequences are different between them.39 MLPKf2 transcripts are more abundant than MLPKf1 transcripts in the stigma.39 Both MLPK forms have an N-myristoylation motif, which functions as a plasma membrane targeting motif.38,39 Bimolecular fluorescence complementation assay has indicated that MLPK interacts with SRK at the plasma membrane and that this interaction is independent of the SCR peptide, suggesting that MLPK might form hetero-oligomers with SRK on the plasma membrane.39 In addition, in vitro kinase assay has revealed that MLPK is phosphorylated by SRK, suggesting that MLPK could be a substrate of SRK.40 However, a complementation experiment with the MLPK gene to demonstrate the function of MLPK has not been performed. Further analysis is required to confirm that MLPK functions in Brassica self-incompatibility signaling.
By using yeast two-hybrid approach with the SRK kinase domain as bait, Arm repeat-containing protein (ARC1) has been identified as a positive regulator of the SRK-mediated signaling cascade.41 ARC1 is a plant U-Box E3 ubiquitin ligase, which functions to attach ubiquitin to target proteins. ARC1 is predominantly expressed in the stigma, and ARC1 is phosphorylated by SRK and MLPK in vitro.41,42 ARC1 has been observed in both cytosol and nuclei when expressed in tobacco BY-2 cells, and it was relocated to the ER-localized proteasomes when ARC1 and SRK910 were coexpressed.42,43 The knockdown of the ARC1 gene in a self-incompatible B. napus ‘W1’ line has been found to result in a partial breakdown of self-incompatibility phenotype,44 suggesting that the ARC1 gene is required for the Brassica self-incompatibility.
Further analysis by using yeast two-hybrid analysis with ARC1 as bait has identified Exo70A1 to be an interactor with ARC1.45 Exo70A1 was ubiquitinated by ARC1 in vitro.45 Exo70A1 is a subunit of the exocyst complex, and mutation of the A. thaliana orthologous gene affected fertility.45,46 Knockdown of EXO70A1 by RNAi in the stigma of self-compatible B. napus ‘Westar’ showed a reduced number of pollen grains on the stigma surface after pollination,45 and, in contrast, expression of Exo70A1 by SLR1 promoter, which is a stigma specific promoter,47 in the self-incompatible B. napus ‘W1’ line partially overcame self-incompatibility.45 In addition, co-expression of the SRK and ARC1 genes caused redistribution of Exo70A1 from cytosol to ER-associated proteasomes in tobacco BY-2 cells.45 In a current model, activated SRK (and MLPK) phosphorylates ARC1, and then the phosphorylated ARC1 ubiquitinates Exo70A1 for proteasome-mediated degradation, resulting in inhibition of pollen germination in self-pollinated stigmas of self-incompatible Brassica plants (Figure 1).
The other srk interactors in Brassica
By using yeast two-hybrid screening with the SRK kinase domain as bait, Thioredoxin H-Like proteins (THL1 and THL2) have been identified as interactors of SRK.48 THL1/2-SRK interaction was mediated by a cysteine residue at a transmembrane domain of SRK protein.49 In vitro analysis suggested that the addition of recombinant THL1/2 proteins inhibited autophosphorylation activity of SRK, and that this inhibition was suppressed by a pollen coat fraction of the same S haplotype.50 Suppression of THL1/2 gene expression in self-compatible B. napus ‘Westar’, which has functional SRK that is identical to B. oleracea SRK15,51 showed spontaneous inhibition of pollen germination and pollen tube elongation.51 These results suggest that THL1/2 proteins function as inhibitors of SRK-mediated signaling in Brassica plants.
In addition to the ARC1 and THL1/2 proteins, kinase-associated protein phosphatase (KAPP), sorting nexin 1 and calmodulin have been identified as SRK interactors.52 Yeast two-hybrid experiments have shown that KAPP interacted with the SRK kinase domain.52 In vitro experiments have revealed that SRK phosphorylated KAPP and KAPP dephosphorylated SRK,52 suggesting that KAPP might function in attenuation of SRK signaling. Calmodulin has been identified by yeast two-hybrid analysis with the kinase domain of a kinase-dead mutant of SRK as the bait, and interacted with SRK in a Ca2+-dependent manner.52 Sorting nexin 1 has also been also found to interact with the kinase-dead mutant.52 However, the necessity and role of these proteins in self-incompatibility remain unclear.
Self-incompatibility signaling pathway in Arabidopsis
Although A. thaliana is a self-compatible plant, self-incompatible transgenic A. thaliana plants have been successfully constructed by introducing the SRK-SCR genes of closely related species, such as A. lyrata, A. halleri and C. gradiflora, into A. thaliana.20,22–26 These results indicate that A. thaliana has all the molecular components required for self-incompatibility signaling other than SRK and/or SCR. Because of a highly efficient and easy transformation protocol of A. thaliana and many genetic resources, the transgenic A. thaliana plants enable in planta evaluation of the molecular components in the self-incompatibility mechanism that were identified in Brassica plants.
The A. thaliana APK1b gene (At2g28930) shows the highest similarity to the B. rapa MLPK gene. Like the MLPK gene, APK1b produced two transcripts from two distinct initiation sites.39,53 In addition, the B. rapa chromosomal region containing MLPK shows the highest synteny with an A. thaliana chromosomal region containing APK1b.53 The transgenic SRKb-SCRb A. thaliana carrying the T-DNA insertion mutation in APK1b (SALK_055314), which is a null mutation,39,54 showed a self-incompatibility response toward self pollen, indicating that the apk1b mutation did not affect the self-incompatibility response in transgenic SRKb-SCRb A. thaliana plants.53 Recently, it has been reported that an apk1b mutation in A. thaliana affected stomatal conductance.55 These reports suggest that APK1b does not function in the self-incompatibility signaling, but functions in another signaling cascade.
The B. rapa genome contains three putative orthologous genes of A. thaliana APK1b, i.e., MLPK (Bra000478), Bra035659 and Bra040929, because of an extra genome triplication in Brassica species. Although the B. rapa genomic region containing the MLPK gene shows the highest similarity to that containing APK1b, synteny analysis using the A. thaliana genomic region containing APK1b as a query revealed that the Brassica genomic region containing Bra035659 has the highest synteny among the three genomic regions of Brassica, indicating that Bra035659 is the orthologous gene of APK1b and A. thaliana does not contain an orthologous gene of Brassica MLPK. These observations may suggest that, if MLPK is required for the self-incompatibility signaling in Brassica plants, the MLPK gene would have appeared as a positive regulator of self-incompatibility signaling after species differentiation between Brassica and Arabidopsis.
A survey of the A. thaliana genome revealed that A. thaliana does not have an orthologous gene of the Brassica ARC1 gene.53,56 In contrast to A. thaliana, A. lyrata, which is a self-incompatible Arabidopsis species, has an ortholog of ARC1.56 Knockdown of the ARC1 gene in the A. lyrata stigmas has been reported to cause partial breakdown of the self-incompatibility response.56 In addition, the A. lyrata ARC1 and B. rapa ARC1 genes have been found to confer strong self-incompatibility phenotype to SRKb-SCRb transgenic A. thaliana Col-0,57 which shows a transient self-incompatibility phenotype, suggesting that ARC1 plays an important role in Arabidopsis self-incompatibility signaling. However, SRKb-SCRb transgenic A. thaliana C24 plants show a strong self-incompatibility response, although the ARC1 gene is not found in the A. thaliana C24 genome nor in the A. thaliana Col-0 genome.56
A. thaliana has an orthologous gene (At5g03540) of Brassica EXO70A1 encoding Exo70A1,53 which has been identified as a putative substrate of ARC1 in Brassica.45 Unlike self-incompatible B. napus ‘W1’ plants, overexpression of EXO70A1 in SRKb-SCRb transgenic A. thaliana was not found to affect the self-incompatibility response.53 This result suggests that Exo70A1 has no effect on self incompatibility in A. thaliana. In contrast to the result of Samuel et al.,45 Li et al.46 have recently reported that A. thaliana exo70a1 mutant did not show defects in pollen germination and pollen tube elongation, when hand-pollinated, supporting the conclusion of Kitashiba et al.53 However, to answer the discrepancy between the result of Samuel et al.45 and Li et al.,46 Safavian et al. have reported that the pollination defect in the A. thaliana exo70a1 mutant was observed at 40% humidity, but the defect was rescued at 80% humidity, suggesting that the pollination defect of the A. thaliana exo70a1 mutant was humidity-dependent.58 To conclude whether Exo70A1 functions in self-incompatibility, further studies are required on degradation of EXO70A1 in self-pollinated stigmas of self-incompatible plants and effect of high humidity, such as 80% humidity, on Brassica self-incompatibility response.
THL1/2 were identified as negative regulators of SRK-mediated self-incompatibility signaling.50,51 The effect of THL proteins on self-incompatibility was examined by using A. thaliana T-DNA insertion mutants of the AtTRX3 and AtTRX4 gene, which are orthologous genes of Brassica THL1 and THL2, respectively.59 Unlike the THL1/2 genes, the attrx3, attrx4 and attrx3attrx4 mutations did not affect the A. thaliana self-incompatibility response.59 The effect of thioredoxin H–SRK interaction on self-incompatibility was also examined by using a transgenic A. thaliana plant expressing an SRKb (C463W) mutant gene, which has a mutation at the Cys463 residue that corresponds to the Cys residue required for the thioredoxin H–SRK interaction.49 The phenotype of the transgenic A. thaliana plants expressing the SRKb (C463W) mutant gene was indistinguishable from that of SRKb transgenic A. thaliana plants under both self- and cross-pollination, indicating that the replacement of the Cys residue does not affect the self-incompatibility response in transgenic A. thaliana.59 In addition, the fact that the Cys residue is not conserved among some SRK haplotypes of not only Arabidopsis but also Brassica plants, such as A. lyrata SRKa and SRK3, A. halleri SRK3 and B. oleracea SRK68.59
Perspectives
After identification of the SRK gene in Brassica, subsequent research has presented several candidates as possible components of self-incompatibility signaling in Brassica. In vitro experimental results have provided an interesting model of a signaling cascade in self-incompatibility in Brassica (Figure 1). However, because of the difficulty of transformation and gene targeting disruption in self-incompatible Brassica plants, the constructed model for self-incompatibility has not been confirmed by in planta experiments using gene-knockout self-incompatible mutant plants.
Because the gene targeting deletion method of non-model organisms including Brassica has not been developed, it has been difficult to examine the necessity and roles of the identified candidates in self-incompatibility mechanism in Brassica. However, recently, new genome editing methods, such as TALEN and CRISPR/Cas,60 have been developed. The development of these methods should enable us to construct null mutants of non-model self-incompatible plants to examine the effects of the candidate genes on self-incompatibility signaling. In turn, molecular components that function in Arabidopsis self-incompatibility signaling have not been identified. Therefore, identification of the molecular components in Arabidopsis self-incompatibility is also required. In conclusion, in planta evaluation of candidate genes in the Brassica self-incompatibility signaling and the identification of molecular components of the Arabidopsis self-incompatibility signaling contribute to the knowledge of not only the molecular mechanism of self-incompatibility, but also evolutionary aspects of the self-incompatibility mechanism in Brassicaceae.
References
Stein JC, Howlett B, Boyes DC, Nasrallah ME, Nasrallah JB . Molecular cloning of a putative receptor protein kinase gene encoded at the self-incompatibility locus of Brassica oleracea. Proc Natl Acad Sci USA 1991; 88: 8816–8820.
Schopfer CR, Nasrallah ME, Nasrallah JB . The male determinant of self-incompatibility in Brassica. Science 1999; 286: 1697–1700.
Suzuki G, Kai N, Hirose T et al. Genomic organization of the S locus: Identification and characterization of genes in SLG/SRK region of S9 haplotype of Brassica campestris (syn. rapa). Genetics 1999; 153: 391–400.
Nasrallah JB, Nasrallah ME . Pollen-stigma signaling in the sporophytic self-incompatibility response. Plant Cell 1993; 5: 1325–1335.
Takasaki T, Hatakeyama K, Suzuki G, Watanabe M, Isogai A, Hinata K . The S receptor kinase determines self-incompatibility in Brassica stigma. Nature 2000; 403: 913–916.
Chen CH, Nasrallah JB . A new class of S sequences defined by a pollen recessive self-incompatibility allele of Brassica oleracea. Mol Gen Genet 1990; 222: 241–248.
Sato K, Nishio T, Kimura R et al. Coevolution of the S-locus genes SRK, SLG and SP11/SCR in Brassica oleracea and B. rapa. Genetics 2002; 162: 931–940.
Iwano M, Shiba H, Funato M, Shimosato H, Takayama S, Isogai A . Immunohistochemical studies on translocation of pollen S-haplotype determinant in self-incompatibility of Brassica rapa. Plant Cell Physiol 2003; 44: 428–436.
Kachroo A, Schopfer CR, Nasrallah ME, Nasrallah JB . Allele-specific receptor-ligand interactions in Brassica self-incompatibility. Science 2001; 293: 1824–1826.
Takayama S, Shimosato H, Shiba H et al. Direct ligand-receptor complex interaction controls Brassica self-incompatibility. Nature 2001; 413: 534–538.
Schopfer CR, Nasrallah JB . Self-incompatibility. Prospects for a novel putative peptide-signaling molecule. Plant Physiol 2000; 124: 935–940.
Watanabe M, Ito A, Takada Y et al. Highly divergent sequences of the pollen self-incompatibility (S) gene in class-I S haplotypes of Brassica campestris (syn.rapa) L. FEBS Lett 2000; 473: 139–144.
Okamoto S, Sato Y, Sakamoto K, Nishio T . Distribution of similar self-incompatibility (S) haplotypes in different genera, Raphanus and Brassica. Sex Plant Reprod 2004; 17: 33–39.
Mishima M, Takayama S, Sasaki KI et al. Structure of the male determinant factor for Brassica self-incompatibility. J Biol Chem 2003; 278: 36389–36395.
Chookajorn T, Kachroo A, Ripoll DR, Clark AG, Nasrallah JB . Specificity determinants and diversification of the Brassica self-incompatibility pollen ligand. Proc Natl Acad Sci USA 2004; 101: 911–917.
Kusaba M, Dwyer K, Hendershot J, Vrebalov J, Nasrallah JB, Nasrallah ME . Self-incompatibility in the genus Arabidopsis: characterization of the S locus in the outcrossing A. lyrata and its autogamous relative A. thaliana. Plant Cell 2001; 13: 627–643.
Sherman-Broyles S, Boggs NA, Farkas A et al. S locus genes and the evolution of self-fertility in Arabidopsis thaliana. Plant Cell 2007; 19: 94–106.
Tang C, Toomajian C, Sherman-Broyles S et al. The evolution of selfing in Arabidopsis thaliana. Science 2007; 317: 1070–1072.
Shimizu KK, Shimizu-Inatsugi R, Tsuchimatsu T, Purugganan MD . Independent origins of self-compatibility in Arabidopsis thaliana. Mol Ecol 2008; 17: 704–714.
Boggs NA, Nasrallah JB, Nasrallah ME . Independent S-locus mutations caused self-fertility in Arabidopsis thaliana. PLoS Genet 2009; 5: e1000426.
Dwyer KG, Berger MT, Ahmed R et al. Molecular characterization and evolution of self-incompatibility genes in Arabidopsis thaliana: the case of the Sc haplotype. Genetics 2013; 193: 985–994.
Nasrallah ME, Liu P, Nasrallah JB . Generation of self-incompatible Arabidopsis thaliana by transfer of two S locus genes from A. lyrata. Science 2002; 297: 247–249.
Nasrallah ME, Liu P, Sherman-Broyles S, Boggs NA, Nasrallah JB . Natural variation in expression of self-incompatibility in Arabidopsis thaliana: Implications for the evolution of selfing. Proc Natl Acad Sci USA 2004; 101: 16070–16074.
Liu P, Sherman-Broyles S, Nasrallah ME, Nasrallah JB . A cryptic modifier causing transient self-incompatibility in Arabidopsis thaliana. Curr Biol 2007; 17: 734–740.
Boggs NA, Dwyer KG, Shah P et al. Expression of distinct self-incompatibility specificities in Arabidopsis thaliana. Genetics 2009; 182: 1313–1321.
Tsuchimatsu T, Suwabe K, Shimizu-Inatsugi R et al. Evolution of self-compatibility in Arabidopsis by a mutation in the male specificity gene. Nature 2010; 464: 1342–1346.
Warwick SI, Francis A, Al-Shehbaz IA . Brassicaceae: species checklist and database on CD-Rom. Plant Syst Evol 2006; 259: 249–258.
Warwick SI, Mummenhoff K, Sauder CA, Koch MA, Al-Shehbaz IA . Closing the gaps: phylogenetic relationships in the Brassicaceae based on DNA sequence data of nuclear ribosomal ITS region. Plant Syst Evol 2010; 285: 209–232.
Al-Shehbaz IA, Beilstein MA, Kellogg EA . Systematics and phylogeny of the Brassicaceae (Cruciferae): an overview. Plant Syst Evol 2006; 259: 89–120.
Bailey CD, Koch MA, Mayer M et al. Toward a global phylogeny of the Brassicaceae. Mol Biol Evol 2006; 23: 2142–2160.
Beilstein MA, Al-Shehbaz IA, Kellogg EA . Brassicaceae phylogeney and trichome evolution. Am J Bot 2006; 93: 607–619.
Beilstein MA, Al-Shehbaz IA, Mathews S, Kellogg EA . Brassicaceae phylogeny inferred from phytochrome A and NDHF sequence data: tribes and trichomes revisited. Am J Bot 2008; 95: 1307–1327.
Franzke A, Lysak MA, Al-Shehbaz IA, Koch MA, Mummenhoff K . Cabbage family affairs: the evolutionary history of Brassicaceae. Trends Plant Sci 2011; 16: 108–116.
Bi YM, Brugière N, Cui Y, Goring DR, Rothstein SJ . Transformation of Arabidopsis with a Brassica SLG/SRK region and ARC1 gene is not sufficient to transfer the self-incompatibility phenotype. Mol Gen Genet 2000; 263: 648–654.
Suzuki T, Kusaba M, Matsushita M, Okazaki K, Nishio T . Characterization of Brassica S-haplotypes lacking S-locus glycoprotein. FEBS Lett 2000; 482: 102–108.
Fujimoto R, Sugimura T, Nishio T . Gene conversion from SLG to SRK resulting in self-compatibility in Brassica rapa. FEBS Lett 2006; 580: 425–430.
Guo YL, Zhao X, Lanz C, Weigel D . Evolution of the S-locus region in Arabidopsis relatives. Plant Physiol 2011; 157: 937–946.
Murase K, Shiba H, Iwano M et al. A membrane-anchored protein kinase involved in Brassica self-incompatibility signaling. Science 2004; 303: 1516–1519.
Kakita M, Murase K, Iwano M et al. Two distinct forms of M-locus protein kinase localize to the plasma membrane and interact directly with S-locus receptor kinase to transduce self-incompatibility signaling in Brassica rapa. Plant Cell 2007; 19: 3961–3973.
Kakita M, Shimosato H, Murase K, Isogai A, Takayama S . Direct interaction between S-locus receptor kinase and M-locus protein kinase involved in Brassica self-incompatibility signaling. Plant Biotechnology 2007; 24: 185–190.
Gu T, Mazzurco M, Sulaman W, Matias DD, Goring DR . Binding of an arm repeat protein to the kinase domain of the S-locus receptor kinase. Proc Natl Acad Sci USA 1998; 95: 382–387.
Samuel MA, Mudgil Y, Salt JN et al. Interactions between the S-domain receptor kinases and AtPUB-ARM E3 ubiquitin ligases suggest a conserved signaling pathway in Arabidopsis. Plant Physiol 2008; 147: 2084–2095.
Stone SL, Anderson EM, Mullen RT, Goring DR . ARC1 is an E3 ubiquitin ligase and promotes the ubiquitination of proteins during the rejection of self-incompatible Brassica pollen. Plant Cell 2003; 15: 885–898.
Stone SL, Arnoldo M, Goring DR . A breakdown of Brassica self-incompatibility in ARC1 antisense transgenic plants. Science 1999; 286: 1729–1731.
Samuel MA, Chong YT, Haasen KE, Aldea-Brydges MG, Stone SL, Goring DR . Cellular pathways regulating responses to compatible and self-incompatible pollen in Brassica and Arabidopsis stigmas intersect at Exo70A1, a putative component of the exocyst complex. Plant Cell 2009; 21: 2655–2671.
Li S, Chen M, Yu D et al. EXO70A1-mediated vesicle trafficking is critical for tracheary element development in Arabidopsis. Plant Cell 2013; 25: 1774–1786.
Franklin TM, Centre JI . SLR1 function is dispensable for both self-incompatible rejection and self-compatible pollination processes in Brassica. Sex Plant Reprod 1996; 9: 203–208.
Bower MS, Matias DD, Fernandes-Carvalho E et al. Two members of the thioredoxin-h family interact with the kinase domain of a Brassica S locus receptor kinase. Plant Cell 1996; 8: 1641–1650.
Mazzurco M, Sulaman W, Elina H, Cock JM, Goring DR . Further analysis of the interactions between the Brassica S receptor kinase and three interacting proteins (ARC1, THL1 and THL2) in the yeast two-hybrid system. Plant Mol Biol 2001; 45: 365–376.
Cabrillac D, Cock JM, Dumas C, Gaude T . The S-locus receptor kinase is inhibited by thioredoxins and activated by pollen coat proteins. Nature 2001; 410: 220–223.
Haffani YZ, Gaude T, Cock JM, Goring DR . Antisense suppression of thioredoxin h mRNA in Brassica napus cv. Westar pistils causes a low level constitutive pollen rejection response. Plant Mol Biol 2004; 55: 619–630.
Vanoosthuyse V, Tichtinsky G, Dumas C et al. Interaction of calmodulin, a sorting nexin and kinase-associated protein phosphatase with the Brassica oleracea S locus receptor kinase. Plant Physiol 2003; 133: 919–929.
Kitashiba H, Liu P, Nishio T, Nasrallah JB, Nasrallah ME . Functional test of Brassica self-incompatibility modifiers in Arabidopsis thaliana. Proc Natl Acad Sci USA 2011; 108: 18173–18178.
Rea AC, Liu P, Nasrallah JB . A transgenic self-incompatible Arabidopsis thaliana model for evolutionary and mechanistic studies of crucifer self-incompatibility. J Exp Bot 2010; 61: 1897–1906.
Elhaddad NS, Hunt L, Sloan J, Gray JE . Light-induced stomatal opening is affected by the guard cell protein kinase APK1b. PLoS One 2014; 9: e97161.
Indriolo E, Tharmapalan P, Wright SI, Goring DR . The ARC1 E3 ligase gene is frequently deleted in self-compatible Brassicaceae species and has a conserved role in Arabidopsis lyrata self-pollen rejection. Plant Cell 2012; 24: 4607–4620.
Indriolo E, Safavian D, Goring DR . The ARC1 E3 ligase promotes two different self-pollen avoidance traits in Arabidopsis. Plant Cell 2014; 26: 1525–1543.
Safavian D, Jamshed M, Sankaranarayanan S, Indriolo E, Samuel MA, Goring DR . High humidity partially rescues the Arabidopsis thaliana exo70A1 stigmatic defect for accepting compatible pollen. Plant Reprod 2014; 27: 121–127.
Yamamoto M, Nasrallah JB . In planta assessment of the role of thioredoxin h proteins in the regulation of S-locus receptor kinase signaling in transgenic Arabidopsis thaliana. Plant Physiol 2013; 163: 1387–1395.
Kim H, Kim JS . A guide to genome engineering with programmable nucleases. Nat Rev Genet 2014; 15: 321–334.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The other authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Yamamoto, M., Nishio, T. Commonalities and differences between Brassica and Arabidopsis self-incompatibility. Hortic Res 1, 14054 (2014). https://doi.org/10.1038/hortres.2014.54
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2014.54
This article is cited by
-
Ancestral self-compatibility facilitates the establishment of allopolyploids in Brassicaceae
Plant Reproduction (2023)
-
Downregulated expression of S2-RNase attenuates self-incompatibility in “Guiyou No. 1” pummelo
Horticulture Research (2021)
-
Impact of whole genome triplication on the evolutionary history and the functional dynamics of regulatory genes involved in Brassica self-incompatibility signalling pathway
Plant Reproduction (2020)
-
The poplar pangenome provides insights into the evolutionary history of the genus
Communications Biology (2019)
-
Progress on deciphering the molecular aspects of cell-to-cell communication in Brassica self-incompatibility response
3 Biotech (2018)