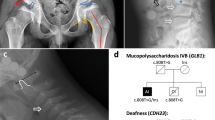
Abstract
GM2 gangliosides are a group of lysosomal lipid storage disorders that are due to mutations in HEXA, HEXB and GM2A. In our study, 10 patients with these diseases were enrolled, and Sanger sequencing was performed for the HEXA and HEXB genes. The results revealed one known splice site mutation (c.346+1G>A, IVS2+1G>A) and three novel mutations (a large deletion involving exons 6–10; one nucleotide deletion, c.622delG [p.D208Ifsx15]; and a missense mutation, c.919G>A [p.E307K]) in HEXA. In HEXB, one known mutation (c.1597C>T [p.R533C]) and one variant of uncertain significance (c.619A>G [p.I207V]) were identified. Five patients had c.1597C>T in HEXB, indicating a common mutation in south Iran. In this study, a unique large deletion in HEXA was identified as a homozygous state. To predict the cause of the large deletion in HEXA, RepeatMasker was used to investigate the Alu elements. In addition, to identify the breakpoint of this deletion, PCR was performed around these elements. Using Repeat masker, different Alu elements were identified across HEXA, mainly in intron 5 and intron 10 adjacent to the deleted exons. PCR around the Alu elements and Sanger sequencing revealed the start point of a large deletion in AluSz6 in the intron 6 and the end of its breakpoint 73 nucleotides downstream of AluJo in intron 10. Our study showed that HEXA is an Alu-rich gene that predisposes individuals to disease-associated large deletions due to these elements.
Similar content being viewed by others
Introduction
GM2 gangliosides are a group of lysosomal lipid storage disorders that resulted from disease-causing mutations in three autosomal recessive genes, including hexosaminidase A (HEXA), hexosaminidase subunit beta (HEXB) and GM2 ganglioside activator (GM2A). The HEXB gene produces a subunit of two related enzymes that include beta-hexosaminidase A and beta-hexosaminidase B. Both enzymes have two subunits. Beta-hexosaminidase A is made from one alpha subunit produced from the HEXA gene and one beta subunit (produced from the HEXB gene). However, beta-hexosaminidase B is only produced from two beta subunits encoded from the HEXB gene. Beta-hexosaminidase A and B, which are mainly found in lysosomes, break down sphingolipids, oligosaccharides and glycoproteins, and are critical in the central nervous system as they have very essential roles in breaking down the GM2 ganglioside in this system, preventing the accumulation of this product and its damage to the nervous system.1–3
According to the Human Gene Mutation Database (HGMD, http://www.hgmd.cf.ac.uk/ac/index.php), more than 182 disease-causing mutations of the HEXA gene have been identified in patients affected by Tay-Sachs disease (MIM #272800), including missense/nonsense, 96; splicing, 33; small deletions, 28; small insertions, 8; small indels, 3; and gross deletions, 1 (7.6 kb incl. promoter+ex. 1). Regarding the HEXB gene, to date, more than 107 pathogenic mutations in this gene have been reported in patents affected by Sandhoff disease (MIM #268800), including missense/nonsense, 39; splicing, 18; small deletion, 18; small insertion, 3; and gross deletion, 6. These mutations result in the accumulation of GM2 ganglioside in the central nervous system, damaging this systems and presenting the signs and symptoms identified in patients with the corresponding disorders. On the basis of the fully nonfunctional HEXA and HEXB proteins or reduced activity of these proteins resulting from mutations in their genes, these disorders are presented as a severe form found in infancy or a less severe type with a later onset. Tay-Sachs and Sandhoff diseases (which are clinically indistinguishable from each other) are autosomal recessive, progressive neurodegenerative disorders. Their classical infantile forms develop in infancy with developmental retardation, paralysis, dementia, blindness and death usually by 2 or 3 years of age.1,2,4–6 A study conducted by Smith et al.3 showed that the clinical findings in infantile-onset disease were generally as follow: hyperacusis with excessive startle response in 85% of individuals; increased myotatic reflexes in 85% of individuals (children with infantile-onset disease were not able to walk independently); a pattern of combined axial hypotonia with limb spasticity in 86% of individuals; visual deterioration in 89% of individuals; and seizures in 84% of individuals. Considering the importance of identifying genetic causes in our patients that are suggestive of Tay-Sachs and Sandhoff diseases, the aim of our study was to identify disease-causing mutations in these patients from the south of Iran and report their clinical findings.
Materials and methods
Ten unrelated patients suspected of GM2 ganglioside diseases from the south of Iran were recruited in the current study. All patients gave informed consent before undergoing genetic analysis of the HEXB and HEXA genes based on the requirements of the ethics committee in Comprehensive Medical Genetics Center, Shiraz University of Medical Sciences. Three milliliters of whole-blood samples from patients was collected in EDTA tubes and stored at −20 °C until use. Genomic DNA was prepared from peripheral blood lymphocytes using a DNA extraction kit (Yekta Tajhiz, Tehran, Iran) according to the manufacturer’s instructions. The genomic DNA concentration was measured by a NanoDrop (ND1000, NanoDrop Technologies, Wilmington, DE, USA) and stored at –20 °C until use.
All of the exon and exon–intron boundaries of the HEXA and HEXB genes were amplified by the PCR primers given in Table 1. These primer pairs were designed and evaluated according to their reference genomic sequences in ENSEMBLE (https://www.ensembl.org) and the use of NCBI BLAST(https://blast.ncbi.nlm.nih.gov), ENSEMBLE BLAST/BLAT (https://www.ensembl.org), UCSC (BLAT and In Silico PCR, https://genome.ucsc.edu) and Allele ID 7.5 (a bioinformatics software that helps design primers and probes). The total volume of the PCR reaction was 50 μl, which contained 1 μl of the forward and reverse primers (20 pmol/μl), 3 μl of DNA template (50–200 ng), 25 μl of TEMPase Hot Start 2x Master Mix Blue (Ampicon, A290806) and 20 μl of dH2O. PCR was carried out according to the Amplicon TEMPase Hot Start program. Ten microliters of PCR amplicons were visualized on a 2% agarose gel containing SYBR Safe (Invitrogen, Paisley, Scotland, UK). Then, the PCR products were sequenced using Sanger sequencing, and their results were analyzed using NCBI BLAST and Codon Code Aligner software (CodonCode Corporation, Centerville, MA, USA).
In addition, to predict the cause of a new large deletion in the HEXA gene due to Alu elements, RepeatMasker (http://www.repeatmasker.org) was used with the RMBlast search engine (a version that was compatible with the NCBI Blast tool suite). The Repeat masker program screens DNA sequences of interspersed repeats and low complexity DNA sequences. Sequence comparisons in RepeatMasker are carried out using one of the different search engines, including nhmmer, cross_match, ABBlast/WUBlast, RMBlast and Decypher. RepeatMasker uses information from curated libraries of repeats, Dfam and Repbase. We investigated any Alu repeats (especially Alu repeats flanking deleted exons) in the whole genomic structure of HEXA, covering a total of 34,695 bp, starting 1.4 kb upstream of exon 1 to 1.4 kb downstream of exon 14. Moreover, to identify the breakpoint of the identified large deletion around exons 6 and 10 in our patient, we performed a second PCR around the predicted Alu elements using the primers given in Table 1 (the two forward primers upstream of AluJo and AluSx in intron 5, and the reverse primers downstream of AluJo in intron 10). The PCR products were then used for Sanger sequencing. In addition, a coding sequence translator (http://www.fr33.net/translator.php) was used to predict the possible effect of this deletion on the amino-acid sequence of the HEXA protein.
To predict the functional effects and conservation status of a newly identified missense mutation in another patient, we also used different bioinformatics tools, including MutationTaster, MutationAssesor, Polyphen, Grantham, PhastCons, GERP, CADD, Fathmm and SIFT. The T-coffee multiple sequence alignment program (http://www.ebi.ac.uk/Tools/msa/tcoffee/) was applied to compare the amino-acid sequence of HEXA proteins across different species to obtain information about the conservation of amino acids at the site of mutation.
Results
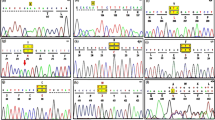
One known (NM_000520 (HEXA-201): c.346+1G>A, IVS2+1G>A)7 splice site and three novel HEXA mutations, including NM_000520 (HEXA-201): c.919G>A [p.E307K], the deletion of exons 6–10 in a homozygous state and NM_000520 (HEXA-201): c.622delG [p.D208Ifsx15], which are suspected as being homozygous, were identified. One variant of uncertain significance (NM_000521 (HEXB-201): c.619A>G [p.I207V])8 and one known mutation (NM_000521 (HEXB-201):c.1597C>T [p.R533C])9 in HEXB were found in a homozygous state (Table 2). From 10 patients, 5 cases only had the homozygous p.R533C mutation in HEXB, indicating its high frequency in our population. For the first time, a large homozygous deletion (deletion of exons 6–10) in HEXA was identified in our patient. Using the repeat masker tool, different Alu elements were found across the HEXA gene (Figure 1), including one Alu at the 5ʹ untranslated region, twelve in intron 1, one in intron 3, four in intron 5 flanking exon 6 (the first exon which was deleted in a block of five exon deletions), one in intron 7, one in intron 10 flanking exon 10 (the last exon which was deleted in the deleted block) and three Alus at the 3ʹ untranslated region. The exact breakpoint of the deletion around exons 6 and 10 were determined using different PCR products flanking the Alu elements in their corresponding intron-flanked deleted exons. In addition, the consequence of this deletion was predicted to be an in-frame deletion of 192 amino acids of the HEXA protein, leading to its abnormal function (Figures 2a–d and 3). In patients with this deletion, PCR did not show any band for exons 6–10, but the patient had PCR bands around Alu elements in introns 5 and 10 (1,000 bp for AluSx-AluJo and 1,500 bp for AluJo-AluJo), revealing the homozygous status. In her parents, all of the exons produced PCR bands and bands around Alu elements, indicating its heterozygous status (Figure 4). Using PCR around Alu elements (AluJo and AluSx in intron 5, and AluJo in intron 10) followed by Sanger sequencing, the results revealed that the breakpoints of the large deletion occurred in AluSz6 (its length was 312 nucleotides, and its breakpoint was in nucleotide 307 of this Alu) in the intron 6 and 73 nucleotides downstream of AluJo in intron 10 (this deletion also contained the whole AluJo sequence; Figure 2a, c). The names of the Alu elements were determined from the results of RepeatMasker and were extracted from Dfam and Repbase.
The breakpoint of the deletion and deleted region in the HEXA gene. (a) The red arrows show the breakpoint of the deletion (the region between these red arrows was completely deleted), blue arrows show the deleted region in the Sanger sequencing chromatogram and blue letters indicate the deleted nucleotides, including all of exons 6–10. Blue-highlighted nucleotides show the sequence of AluSz6, and yellow-highlighted nucleotides represent AluJo. (b) The homology between AluSz6 and AluJo is indicated as 76%, which is used for recombination between these two Alu elements. (c) Sequencing data confirmed that the breakpoint of the large deletion in HEXA was due to the Alu elements. (d) Normal and mutant HEXA proteins that resulted from the large deletion are compared, indicating the loss of 192 amino acids from the protein. In the normal sequence, the brown underline shows the deleted residues, and in the mutant sequence, the underline shows the breakpoints of the deletion and also the two joined amino acids after this deletion.
The similarity between AluSz6 and AluJo and its flanking sequence. There was high homology between these Alu elements, and the microhomology between 18 nucleotides of the AluSz6 and the sequence at the breakpoints helps to determine the occurrence of recombination between these sequences, leading to Alu-mediated deletion. Blue letters show homology between AluSz6 and AluJo. Red letters show microhomology at the breakpoints.
PCR products from patients with deletion of exons 6–10 in HEXA. (a) PCR for different exons of HEXA in patients for whom exons 6–10 were deleted as there were no PCR bands for them (here, only we show deletion of exons 6–8 from this deleted block). (b) PCR products using primers upstream of AluJo and AluSx in intron 5 and downstream of AluJo in intron 10. PCR revealed that the proband with the exon 6–10 deletion only had PCR bands produced from primers designed around the Alu elements (1,000 bp for AluSx-AluJo and 1,500 bp for AluJo-AluJo), revealing that the homozygous status due to the amplification of the reduced size of the exon 6–10 block resulted from this deletion. In our patient’s parents, not only were the bands amplified from the primers around the Alu elements detected but all of the exons of HEXA also produced PCR bands (here, only the PCR bands around the Alu elements are given for parents), indicating its heterozygous status. E, exon.
In addition to a novel large deletion, a new c.919G>A [p.E307K] missense mutation in HEXA was also identified as homozygous in one patient and was confirmed as being heterozygous using Sanger sequencing in her parents (Figure 5a). Moreover, different bioinformatics tools confirmed the pathogenicity of this mutation (Table 3). In addition, the T-coffee tool revealed that the glutamate residue at position 307 was highly conserved during evolution among different species (Figure 5b). Another new mutation, c.622delG [p.D208Ifsx15], was identified as being heterozygous in parents of the Case-MOE (Table 2 and Figure 5c), and since the death of patient was before the time of genetic testing, we only identified this deleterious mutation by performing Sanger sequencing of the whole HEXA and HEXB genes in both parents. Therefore, this case is suspected to be homozygous.
(a) Sanger data from the family with a novel c.919G>A (p.E307K) missense mutation. The reference sequence was obtained from the Ensembl database (https://www.ensembl.org/). Reference nucleotide and residues are highlighted in yellow. (b) Comparative alignment of the amino acids of HEXA across different species. The E307 residue is highly conserved during evolution. The conserved glutamic acid residue is shown in the rectangular box. Protein sequences were obtained from National Center for Biotechnology (NCBI). Symbols: (*)—identical amino acids; (:)—only similar amino acids. Pongo abelii: one of two species of orangutans; Macaca fascicularis: one species of macaque; Bos taurus (domestic cow): a species of oxen; Ovis aries: domestic sheep; Felis catus: domestic cat; Sus scrofa: wild boar or Eurasian wild pig; Rattus norvegicus: brown rat, also referred to as common rat, street rat and Norway rat; Mus musculus: house mouse; Gallus gallus (the red junglefowl): a type of domesticated fowl. (c) The Sanger chromatogram revealed a novel heterozygous c.622delG, p.D208Ifsx15 in both parents of the proband. Reference nucleotides are highlighted in yellow.
Discussion
The current study revealed the structure and distribution of a large deletion mutation consisting of exons 6–10 of HEXA in Tay-Sachs disease. The deletion was 4.6 kb long and spanned from 576 bp upstream of exon 6 to 475 bp downstream of exon 10. As shown in Figure 1, the Alu elements are located across the normal HEXA gene, which mainly flank the deleted region, demonstrating the involvement of these elements in large deletions in HEXA.
Alu elements are short interspersed elements that are moderately repetitive sequences and have a copy number of more than 1 million copies across the human genome (~11% of human genome). Alu elements are considered to be one of the most successful mobile elements.10,11 They are ~300 nucleotides in length and have a dimeric structure. The 3ʹ-end of an Alu element has a poly A-like structure, which plays a key role in its amplification mechanism.12–15 There are different subfamilies of Alu sequences that share similar nucleotides in each group, and the most common Alu subfamily in the human genome is the Alu S class (the most common Alu is AluSx).16 Alu sequences are responsible for the instability of the genome, which resulted from intrachromosomal and interchromosomal recombination events between these sequences.17,18 Deletions due to Alu repeats have been identified in different previously reported genes, such as the low-density lipoprotein receptor,19 NACHT, leucine-rich repeat, PYD-containing 720 and HEXB genes.21
It is worth noting that while most breakpoints of deletions occur in Alu elements, one of the deletion breakpoints in our study was observed 73 bp downstream of the AluJo element in intron 10 (AluJo and 73 bp after this element were deleted along with the deleted block). As shown in Figures 1 and 2, the deletion occurred due to recombination between AluSz6 and AluJo as the breakpoint occurred in the last nucleotides of AluSz6 and 73 bp downstream of AluJo. Although there is an AluJo in intron 6, its direction is opposite of AluJo located in intron 10 (Figure 1, the same direction is needed to cause a deletion when recombination between Alu/Alu occurs). In addition, the similarity between AluSz6 and AluJo is 76% (Figure 2b), which is suitable for recombination as Alu/Alu recombination occurs when Alu elements are completely identical (homologous). In addition, in most of the observed events, Alu has a mismatch (homeologous) at the start of recombination.11,22–24
An explanation of why the breakpoints in intron 10 occurred 73 bp downstream from AluJo is that that some microhomology was observed at this breakpoint, such as with nucleotide adenine (shown as red letters in Figure 3). This microhomology, along with the homology between Alu, makes it possible to observe these Alu/Alu deletions. It has been shown that these deletions are due to pathways such as microhomology-mediated end joining, which occurs when there are 5–25 nucleotides of microhomology between two Alu elements.25,26 Therefore, as shown in Figure 3, microhomology between two Alu elements can be one of the main causes of the large deletion in the HEXA gene.
The c.919G>A missense mutation in exon 8 of HEXA was a novel mutation and resulted in a substitution from glutamate, a negatively charged, polar amino acid (at position 307) to lysine, a positively charged, polar amino acid (p.E307K). This amino acid is predicted to be essential for the proper functions of this protein (Table 3) and is conserved in the evolution of different species, such as Mus musculus, Gallus gallus, Ovis aries and many others (Figure 5b). Moreover, Sanger sequencing of all of exons and exon/intron boundaries of the HEXA and HEXB genes in the proband only identified this mutation in exon 8 of HEXA. On the basis of our results, this new variation is a real mutation and is predicted to have damaging effects on the protein.
Regarding c.622delG and p.D208Ifsx15, it is obvious that this mutation causes a frameshift deletion that led to a premature translation termination. It is worth noting that one of the reported transcripts of HEXA (ENST00000569410.5, H3BTD4, http://www.ensembl.org), which lacks all of the amino acids after position 373, caused nonsense-mediated decay. Therefore, all of these amino acids are vital for the normal function of the HEXA protein. Since the only identified damaging variation across HEXA and HEXB in both parents of the patient was this deletion and all of the phenotypes of the patient overlapped with the clinical abnormalities found in Tay-Sachs disease, this frameshift mutation can be reported to be the genetic cause of Tay-Sachs disease in this family.
One previously reported variant of uncertain significance (c.619A>G [p.I207V]) that has a high allele frequency (0.15148 for G) in the main database, ExAC (http://exac.broadinstitute.org), was identified as homozygous in one of our patients (Case-NEGAH, Table 2). There are conflicting interpretations of this variant since some studies have reported that the c.619A>G variant converts the Ile to Val (in position 207) in a highly conserved region of the HEXB protein, which is considered to be associated with alterations in the catalytic activity and activator protein binding8. However, there are several reports that indicate that this variant is benign27,28 or likely benign.29 Studies conducted by Redonnet-Vernhet et al.28 showed that the healthy mother of the patient who was affected by Sandhoff’s disease due to a mutation of p.R505Q was homozygous for p.I207V, indicating that this variant did not cause Sandhoff’s disease. Therefore, we suggest that the c.619A>G variant identified in our patient cannot be the genetic cause of Sandhoff’s disease and other genes with a clinical diagnosis that overlapped this disease may be involved in this case, and other genetic tests are needed, such as whole-exome sequencing.
Current studies have shown that HEXA is an Alu-rich gene and is predisposed to disease-associated large deletions. We also found different mutations in the HEXA and HEXB genes and that the most common mutation in HEXB was c.1597C>T, which suggests that screening the HEXB gene for this prevalent mutation is a cost-effective test in individuals with suspicion of having Sandhoff disease. Such studies can help understand the different possible mechanisms of genetic changes in different diseases and help to investigate large deletions in HEXA and HEXB in individuals affected by Sandhoff and Tay-Sachs diseases who do not have any point mutations or small deletions and insertions. In addition, with identification of all of the Alu elements across HEXA in this study, other researchers can look for any possible deletions more easily in special cases of Sandhoff and Tay-Sachs diseases. Until now, no drugs have been used to treat Sandhoff and Tay-Sachs diseases, and a better understanding of their exact pathomechanism in the future can shed light on therapeutic approaches for these diseases using components involved in their biological processes.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Gravel RA et al. The Metabolic and Molecular Bases of Inherited Disease. vol. 3, 8th edn. McGraw-Hill: New York, NY, USA, 2001, 3827–3876.
Chin E, Bean L, Coffee B, Hegde MR . Novel human pathological mutations. Gene symbol: HEXA. Disease: Tay-Sachs disease. Hum Genet 2009; 126: 329.
Smith NJ, Winstone AM, Stellitano L, Cox TM, Verity CM . GM2 gangliosidosis in a UK study of children with progressive neurodegeneration: 73 cases reviewed. Dev Med Child Neurol 2012; 54: 176–182.
Langlois S, Wilson RD, Genetics Committee of the Society of O, Gynaecologists of C, CCMG Prenatal Diagnosis. Carrier screening for genetic disorders in individuals of Ashkenazi Jewish descent. J Obstet Gynaecol Can 2006; 28: 324–343.
O'Dowd BF, Klavins MH, Willard HF, Gravel R, Lowden JA, Mahuran DJ et al. Molecular heterogeneity in the infantile and juvenile forms of Sandhoff disease (O-variant GM2 gangliosidosis). J Biol Chem 1986; 261: 12680–12685.
Neudorfer O, Pastores GM, Zeng BJ, Gianutsos J, Zaroff CM, Kolodny EH et al. Late-onset Tay-Sachs disease: phenotypic characterization and genotypic correlations in 21 affected patients. Genet Med 2005; 7: 119–123.
Akli S, Chelly J, Lacorte JM, Poenaru L, Kahn A . Seven novel Tay-Sachs mutations detected by chemical mismatch cleavage of PCR-amplified cDNA fragments. Genomics 1991; 11: 124–134.
Banerjee P, Siciliano L, Oliveri D, McCabe NR, Boyers MJ, Horwitz AL et al. Molecular basis of an adult form of beta-hexosaminidase B deficiency with motor neuron disease. Biochem Biophys Res Commun 1991; 181: 108–115.
Yoshizawa T, Kohno Y, Nissato S, Shoji S . Compound heterozygosity with two novel mutations in the HEXB gene produces adult Sandhoff disease presenting as a motor neuron disease phenotype. J Neurol Sci 2002; 195: 129–138.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J et al. Initial sequencing and analysis of the human genome. Nature 2001; 409: 860–921.
Batzer MA, Deininger PL . Alu repeats and human genomic diversity. Nat Rev Genet 2002; 3: 370–379.
Kazazian HH Jr . Mobile elements: drivers of genome evolution. Science 2004; 303: 1626–1632.
Hedges DJ, Batzer MA . From the margins of the genome: mobile elements shape primate evolution. Bioessays 2005; 27: 785–794.
Deininger PL, Moran JV, Batzer MA, Kazazian HH Jr . Mobile elements and mammalian genome evolution. Curr Opin Genet Dev 2003; 13: 651–658.
Jurka J, Smith T . A fundamental division in the Alu family of repeated sequences. Proc Natl Acad Sci USA 1988; 85: 4775–4778.
Price AL, Eskin E, Pevzner PA . Whole-genome analysis of Alu repeat elements reveals complex evolutionary history. Genome Res 2004; 14: 2245–2252.
Callinan PA, Wang J, Herke SW, Garber RK, Liang P, Batzer MA et al. Alu retrotransposition-mediated deletion. J Mol Biol 2005; 348: 791–800.
Deininger PL, Batzer MA . Alu repeats and human disease. Mol Genet Metab 1999; 67: 183–193.
Faiz F, Allcock RJ, Hooper AJ, van Bockxmeer FM . Detection of variations and identifying genomic breakpoints for large deletions in the LDLR by Ion Torrent semiconductor sequencing. Atherosclerosis 2013; 230: 249–255.
Reddy R,A, Nguyen NM, Sarrabay G, Rezaei M, Rivas MC, Kavasoglu A et al. The genomic architecture of NLRP7 is Alu rich and predisposes to disease-associated large deletions. Eur J Hum Genet 2016; 24: 1445–1452.
Neote K, McInnes B, Mahuran DJ, Gravel RA . Structure and distribution of an Alu-type deletion mutation in Sandhoff disease. J Clin Invest 1990; 86: 1524–1531.
Hedges DJ, Deininger PL . Inviting instability: transposable elements, double-strand breaks, and the maintenance of genome integrity. Mutat Res 2007; 616: 46–59.
Witherspoon DJ, Zhang Y, Xing J, Watkins WS, Ha H, Batzer MA et al. Mobile element scanning (ME-Scan) identifies thousands of novel Alu insertions in diverse human populations. Genome Res 2013; 23: 1170–1181.
Konkel MK, Walker JA, Batzer MA . LINEs and SINEs of primate evolution. Evol Anthropol 2010; 19: 236–249.
Elliott B, Richardson C, Jasin M . Chromosomal translocation mechanisms at intronic alu elements in mammalian cells. Mol Cell 2005; 17: 885–894.
McVey M, Lee SE . MMEJ repair of double-strand breaks (director’s cut): deleted sequences and alternative endings. Trends Genet 2008; 24: 529–538.
Zhang ZX, Wakamatsu N, Akerman BR, Mules EH, Thomas GH, Gravel RA et al. A second, large deletion in the HEXB gene in a patient with infantile Sandhoff disease. Hum Mol Genet 1995; 4: 777–780.
Redonnet-Vernhet I, Mahuran DJ, Salvayre R, Dubas F, Levade T . Significance of two point mutations present in each HEXB allele of patients with adult GM2 gangliosidosis (Sandhoff disease) homozygosity for the Ile207-->Val substitution is not associated with a clinical or biochemical phenotype. Biochim Biophys Acta 1996; 1317: 127–133.
Hara A, Uyama E, Uchino M, Shimmoto M, Utsumi K, Itoh K et al. Adult Sandhoff's disease: R505Q and I207V substitutions in the HEXB gene of the first Japanese case. J Neurol Sci 1998; 155: 86–91.
Acknowledgements
We thank the families of our patients for their participation in this research.
Author information
Authors and Affiliations
Contributions
MF: conceived and designed the experiments. HD: conceived, designed, performed all of the experiments related to the new large deletion and Alu elements as well as bioinformatics analysis for novel deletions and missense mutations, analyzed the data and wrote the paper. MA: designed all of the primers for HEXA and HEXB and performed PCR. SM: performed some of the experiments. FD: analyzed some of the Sanger Sequencing results. FK: extracted DNA from the blood samples.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Dastsooz, H., Alipour, M., Mohammadi, S. et al. Identification of mutations in HEXA and HEXB in Sandhoff and Tay-Sachs diseases: a new large deletion caused by Alu elements in HEXA. Hum Genome Var 5, 18003 (2018). https://doi.org/10.1038/hgv.2018.3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hgv.2018.3
This article is cited by
-
Biochemical and mutational analyses of HEXA in a cohort of Egyptian patients with infantile Tay-Sachs disease. Expansion of the mutation spectrum
Orphanet Journal of Rare Diseases (2023)
-
Clinical presentation and outcome in infantile Sandhoff disease: a case series of 25 patients from Iranian neurometabolic bioregistry with five novel mutations
Orphanet Journal of Rare Diseases (2018)