Abstract
Reproduction and dispersal are key aspects of species life history that influence spatial genetic structure in populations. Several ant species in the genus Cataglyphis have evolved a unique breeding system in which new reproductives (that is, queens and males) are produced asexually by parthenogenesis; in contrast, non-reproductives (that is, workers) are produced via sexual reproduction by mates from distinct genetic lineages. We investigated how these two coexisting reproductive methods affect population-level spatial genetic structure using the ant Cataglyphis mauritanica as a model. We obtained genotypes for queens and their male mates from 338 colonies, and we found that the two lineages present in the study population occurred with equal frequency. Furthermore, analysis of spatial genetic structure revealed strong sex-biased dispersal. Because queens were produced by parthenogenesis and because they dispersed over short distances, there was an extreme level of spatial structuring: a mosaic of patches composed of clonal queens was formed. Males, on the other hand, dispersed over several hundred metres and, thus, across patches, ensuring successful interlineage mating.
Similar content being viewed by others
Introduction
Reproduction and dispersal have long been recognised as key components of species life history that influence genetic variability and structure in populations (Chesser, 1991; Ross, 2001; Clobert et al., 2012). Dispersal mediates gene flow both within and among populations and affects the spatial distribution of genotypes. For example, limited dispersal can cause relatedness among individuals to decrease with increasing geographical distance (that is, an isolation-by-distance (IBD) pattern; Vekemans and Hardy, 2004). Furthermore, breeding systems also directly affect genetic diversity and structure (Ross, 2001; Charlesworth et al., 2009; Pearse and Anderson, 2009). In general, multiple mating increases genetic variability among offspring. Moreover, mating between relatives tends to decrease heterozygosity over generations. Another critical factor shaping population genetic structure is the mode of reproduction, namely sexual or asexual (Halkett et al., 2005; Vallejo-Marín et al., 2010). In diploid organisms, strict asexuality is assumed to impact population genetic structure in two ways (Balloux et al., 2003; Halkett et al., 2005; Hartfield et al., 2016). First, it increases heterozygosity over generations due to the accumulation of mutations and the lack of genetic recombination (that is, Meselson effect). Second, beneficial mutations are expected to decrease genetic diversity within populations, as lineages with a new beneficial mutation will displace other lineages. However, asexual reproduction does not necessarily mean strict clonality. In most asexually reproducing organisms, it has been shown that occasional events of gene conversion and recombination during parthenogenesis may occur, thus reducing heterozygosity over time (Schurko et al., 2009; Schwander, 2016). All in all, the vast diversity of reproductive strategies, from obligate sexuality to strict asexuality, should translate into a variety of patterns of population genetic structure.
Ants display a vast array of reproductive systems, including sexual reproduction, parthenogenesis (that is, arrhenotoky and thelytoky), androgenesis and hybridogenesis (Helms Cahan and Keller, 2003; Rabeling and Kronauer, 2013; Schwander and Oldroyd, 2016; Trible and Kronauer, 2017). In hybridogenetic species, two divergent genetic lineages coexist within populations. Males and females from different lineages interbreed to produce the sterile worker caste. As a consequence, each colony consists of individuals with strikingly different genetic backgrounds: female and male reproductives represent alternative lineages, whereas workers are hybrids of these lineages. Thus, the two lineages must coexist in each population for the system to be maintained (Leniaud et al., 2012; Norman et al., 2016; Romiguier et al., 2016). Whereas the phylogeographic origin and consequences of social hybridogenesis have been widely studied (Sirviö et al., 2011; Darras et al., 2014a), the impact of dependent-lineage reproductive systems on local spatial genetic structure remains largely unknown.
Recent genetic studies have revealed a unique form of social hybridogenesis in Cataglyphis desert ants (Leniaud et al., 2012; Eyer et al., 2013; Darras et al., 2014a,2014b). Two genetically distinct lineages coexist and interbreed (Figure 1). Workers are produced by sexual reproduction between mates originating from different genetic backgrounds and are thus all interlineage hybrids. In contrast, female reproductives are produced by asexual reproduction, namely thelytokous parthenogenesis. Males are produced by arrhenotokous parthenogenesis and are haploid, as is usually the case in Hymenoptera. As a consequence of worker sterility, only maternal genes are perpetuated across generations in each lineage.
Social hybridogenesis in Cataglyphis. Two genetic lineages coexist in all populations (lineages 1 and 2 are represented in black and white, respectively). Queens from one lineage typically mate with males from the other lineage. Workers are produced by sexual reproduction and are all interlineage hybrids, whereas male and female reproductives are produced via arrhenotokous and thelytokous parthenogenesis, respectively. Lineage labels indicate the queens’ lineage of origin.
In this study, we investigated how population-level spatial genetic structure was influenced by dependent-lineage reproduction, parthenogenetic production of new queens and sex-biased dispersal using the polygynous (multiple-queen colonies) ant species Cataglyphis mauritanica as a model (Eyer et al., 2013). In this species, queens mate with males that fly in from other colonies; the queens then disperse on foot with nestmate workers to found a new colony nearby (that is, colony budding; Knaden and Wehner, 2006; Peeters and Aron, in press). Hybridogenesis and limited dispersal of queens is expected to impact population genetic structure in three ways: (i) both lineages must coexist in populations for worker production; at equilibrium, a one-to-one ratio of the two lineages should be maintained by negative frequency-dependent selection (Yamauchi and Yamamura, 2006, ii) reduced dispersal of parthenogenetically produced queens should generate strong micro-geographical genetic structure (that is, patches of colonies headed by same-lineage queens) and (iii) males being the dispersing sex, they must disperse far enough to ensure interlineage mating. To test these assumptions, we extensively sampled C. mauritanica colonies in a large population in Morocco. First, we confirmed the existence of two interbreeding lineages within the population and assessed lineage frequency by analysing the genotypes of females and their mates. Second, using spatial principal component analyses (sPCAs), we characterised spatial patterns of genetic variation in female and male reproductives, both within and among lineages. Third, we estimated male dispersal distances by assigning males to their putative natal colonies via genotype comparisons.
Materials and methods
Sampling
Cataglyphis mauritanica is found in semiarid habitats in northern Africa. We characterised the genetic structure of a population spread across 65 ha in Ifrane National Park (33°33′22′′ North; 5°14′15′′ West), which is located in the Atlas Mountains of Morocco. Previous studies in the same population showed that colonies typically contained hundreds to a few thousands workers (range: 200–3000), multiple queens (range: 2–50) that were most often singly mated, and that reproduction proceeded via social hybridogenesis (Eyer et al., 2013; Eyer 2014). In May 2015, we sampled 338 colonies just prior to the emergence of new reproductives. One hundred twenty-four colonies were completely excavated: all queens and workers were collected. For the remaining 214 colonies, 10 workers were sampled at nest entrances. All individuals were stored in 98% ethanol until genetic analyses could be carried out.
From the 124 excavated colonies, a total of 836 queens were collected (mean queen number per colony±s.d.: 12.4±11.8). To determine the genotypes of their mates (hereafter called the males), the queens were dissected to obtain the contents of their spermathecae. Each spermatheca was removed and placed in 5 μl of Ringer’s solution. The spermathecal wall was ruptured using forceps, and any seminal fluid was collected using a micropipette, taking extra care to avoid contamination with female tissue. The samples were then stored at −20 °C until genetic analyses could be carried out. We found that 744 of the 836 queens had sperm in their spermathecae; the others were unmated.
Genetic analyses
DNA was extracted from adult ants using the Chelex 100 method (see Darras et al., 2014a for details). To isolate sperm DNA, each sample of seminal fluid was first mixed into a solution containing 20 μl of 5% Chelex 100 (Bio-Rad, Hercules, CA, USA) and 3 μl of proteinase K (16 μg μl−1). Second, it was incubated, first for 30 min at 56 °C and then for 1.5 h at 85 °C. After a 3 min centrifugation at 12 000 r.p.m., 20 μl of the supernatant was transferred into a 1.5 ml tube.
Workers, queens and sperm were genotyped at nine microsatellite loci initially developed for use in C. cursor (Cc11 and Cc54; Pearcy et al., 2004) and C. hispanica (Ch01, Ch06, Ch08, Ch11, Ch12, Ch19 and Ch22; Darras et al., 2014a). All the markers were amplified in nine-plex PCR reactions carried out using a QIAGEN Type-it Microsatellite PCR Kit (10- μl reactions as per the manufacturer’s instructions; annealing temperature of 59 °C). The PCR products were genotyped using an automated Applied Biosystems ABI 3730 Sequencer (Applied Biosystems, Foster City, CA, USA). None of the haploid males were missing alleles at the chosen microsatellite markers (n=sperm from 602 males), which meant we detected no null alleles in our data set. The total and effective numbers of alleles per locus, as well as observed and expected heterozygosity are given in Supplementary Table S1.
In our study population, nestmate queens typically have been found to have identical multilocus genotypes because they are produced via parthenogenesis (Eyer et al., 2013). We confirmed this pattern by genotyping all the queens from two large colonies (n=47 and 48). Subsequently, we assessed the queen genotype for a given colony by sampling a single queen (n=124). Male genotypes were determined using the contents of the queens’ spermathecae. When an identical male genotype was found in the spermathecae of several nestmate queens, presumably because a single male had multiple mates, the genotype was included only once in the analyses to avoid statistical bias arising from non-independence of genotypes.
For the 214 colonies from which only workers were sampled, queen and male genotypes were inferred based on the workers’ genotypes (mean number of workers genotyped per colony±s.d.: 6±1.7; n=1167). Matrilines and patrilines were easily reconstructed because (a) the nestmate queens shared the same diploid genotype (no more than two different maternal alleles were found in workers irrespective of queen number); (b) males are haploid (that is, two workers with different paternal alleles at one locus belong to different patrilines); and (c) all the workers were hybrids of two highly divergent lineages (in most cases, assigning a worker’s alleles to two specific parents was straightforward).
Statistical analysis
Genetic lineages
We first confirmed the existence of two genetic lineages in our study population. We assigned all the queens and males to given lineages using a genetic-distance-based principal coordinate analysis, which was implemented in GenAlEx v. 6.502 (Peakall and Smouse, 2012). This method can correctly differentiate genetic lineages in Cataglyphis species that employ dependent-lineage reproductive systems (Leniaud et al., 2012; Darras et al., 2014a).
Analysis of spatial genetic structure
To determine how queens and males from each lineage were spatially distributed within the population, sPCAs (Jombart et al., 2008) were performed using the genotype data and the adegenet package (Jombart, 2008) in R v. 3.2.5 (R Development Core Team, 2016). This multivariate method has been found to perform better than conventional PCAs when dealing with spatial patterns (Jombart et al., 2008). In addition to its ability to detect multiscale spatial genetic patterns, sPCA has the advantage of being able to exploit data that do not meet Hardy–Weinberg expectations or that exhibit linkage disequilibrium among loci, which is not the case for most other spatially explicit approaches for examining genetic variability (Jombart et al., 2008). In our study, the analysis was aimed at detecting significant spatial genetic structure in the queens and males. The sPCA approach explicitly takes into account the spatial information associated with genetic data to search for spatial patterns based on allele frequencies within populations. It successfully distinguishes between positive and negative spatial autocorrelation patterns, which indicate global and local structure, respectively (Jombart et al., 2008). A global pattern corresponds to geographically distinct groups of individuals within which individuals have more similar genotypes than expected by chance. A local structure pinpoints stronger genetic differences among neighbours than among random pairs of entities.
A first sPCA was run using the full queen data set to search for spatial segregation among colonies headed by queens from different lineages. Then, two sPCAs were performed separately—for queens of each lineage—to test whether significant global and/or local structure was present. Finally, two sPCAs were carried out using data on males of each lineage to test for spatial patterns. For each sPCA, 9999 permutations were performed. When an sPCA revealed the presence of significant spatial structure, a Ward cluster analysis was carried out to detect the groups of queens/males that displayed high within-group similarity; the R packages cluster (Maechler et al., 2005) and gclust (Hurley, 2012) were used. Three independent criteria proposed by Borcard et al. (2011) were employed to create optimal clusters based on the resulting dendrograms. Genetic differentiation among clusters was then estimated by calculating FST values using SPAGeDi software v. 1.5 (Hardy and Vekemans, 2002).
To further investigate the strength of spatial genetic structuring for each sex within and among lineages, we determined relatedness by calculating the mean kinship coefficient, Fij (Loiselle et al., 1995), using SPAGeDi software. Fij expresses the level of kinship between individuals i and j relative to the mean kinship between randomly sampled individuals from the study population, which is assumed to be zero (Hardy, 2003). First, mean Fij values were determined for a set of distance intervals (maximum distance in each interval in metres: 30, 60, 100, 150, 200, 300, 400, 500, 700 and 900). Second, the relationship between these values and the natural logarithm of the geographical distance between i and j, ln(dij), was determined using linear regression; indeed, Fij is expected to decrease in a largely linear manner with ln(dij) when genetic structure is the result of limited gene dispersal (Vekemans and Hardy, 2004), which is what is expected for C. mauritanica queens. The degree of spatial genetic structuring in each sex and lineage was compared using the Sp statistic, where Sp=–b/(1−F(N)); b is the slope of the regression of Fij on ln(dij), and F(N) is the mean Fij between individuals from neighbouring colonies (Vekemans and Hardy, 2004). F(N) is estimated by calculating the mean Fij between individuals belonging to colonies separated by <30 m (that is, the first distance interval). Approximate standard errors (SE) were obtained by applying a jackknife procedure across loci. To test whether a non-random spatial genetic structure was present within each sex and lineage, colony locations were randomised 9999 times to obtain the distribution of b under the null hypothesis that there was no spatial structure.
Estimation of male dispersal distances
As C. mauritanica queens are produced by parthenogenesis, they are highly genetically similar and it is impractical to estimate their dispersal distances. As for C. mauritanica males, their dispersal distances cannot be determined from classical IBD models and genetic neighbourhood sizes (Rousset, 2000) because they do not contribute to the next generation of dispersers. Therefore, we took an alternative approach. First, we inferred the males’ potential natal colonies by comparing the genotype of each male with those of all the queens. Only multilocus genotypes that matched up perfectly at all loci were kept in the analyses. Second, we calculated the Euclidian distance between each male’s putative natal colony and the colony headed by the female with whom he mated (that is, as ascertained via sperm or worker genotypes). When males had several possible mothers, we calculated the mean Euclidian dispersal distance. Our estimates were exclusively based on effective dispersal events since they relied on a male having successfully bred. Because many queens had the same multilocus genotype, each male had multiple potential mothers (Results section), which meant that dispersal range may have been overestimated. However, as genetically similar queens were spatially aggregated and formed highly differentiated clusters within genetic lineages, we consider that our estimates of male dispersal distance were reliable.
Results
Genetic analyses indicated that 602 of the 744 mated queens were singly mated. In 48 colonies, we found nestmate queens carrying genetically similar sperm, suggesting they had mated with the same male or brother males (n=290 across all colonies). In these cases, we only counted the male genotype once. Ultimately, we observed a total of 407 male genotypes in the spermathecae. For the 214 colonies from which only workers were sampled, we were able to unambiguously reconstruct the queen’s genotype for 203 colonies; we reconstructed 628 male genotypes. Consequently, overall, we genotyped 1167 workers, 327 queens (124 queens: directly genotyped; 203 queens: genotype inferred from workers), and 1035 males (407 males: genotype ascertained from spermathecal contents; 628 males: genotype inferred from workers).
Genetic lineages
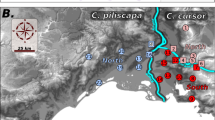
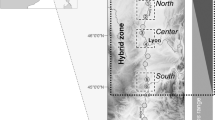
As expected, the principal coordinate analysis showed that queen and male genotypes formed two clusters that could be distinguished along the first coordinate, which accounted for 23% of the total genetic variance (see Supplementary Figure S1). Both sexes were found in each cluster. The two genetic lineages formed highly divergent gene pools, with all nine markers showing different allele distributions. On average, per locus, 77% of alleles (range: 50–100%) were representative of the lineage. Not a single queen or male displayed an intermediate genotype, suggesting none was the product of interlineage hybridisation or introgression. Queens from lineage 1 were found in 168 colonies (51.4%), whereas queens from lineage 2 were found in 159 colonies (48.6%). The two lineages were equally common (χ2 test: X21=0.25; P=0.62). The spatial distribution of the two lineages is illustrated in Figure 2.
Among the 602 singly mated queens, 80% had mated with a partner from the other lineage. However, a remarkable percentage of queens had mated with males of their own lineage (19 and 21% for lineages 1 and 2, respectively). All the workers (n=1167) were hybrids of the two lineages, indicating that they arose from interlineage mating.
Analysis of spatial genetic structure
Spatial genetic structure of the queens
Table 1 gives the main results of the sPCA performed on the genotypes of the queens and the males. The analysis found that the queens were clearly spatially aggregated based on lineage (PGlobal<0.001). This spatial pattern described 38% of the total genetic variance (first axis (Table 1)); the remaining 62% represented within-lineage variance, which led us to analyse each lineage separately.
For each lineage, three sPCA axes that corresponded to positively correlated allele distributions were selected based on their eigenvalues (PGlobal<0.001 for both lineages). Together, these three axes described 60 and 58% of the total genetic variance among lineage-1 and lineage-2 queens, respectively. No negative autocorrelations were found among neighbouring queens (lineage 1: PLocal=0.93; lineage 2: PLocal=0.73). The spatial pattern that emerged from the overlap of the three axes revealed the presence of five groups within lineage 1 and of four to five groups within lineage 2 (Figure 3). Ward clustering analyses, based on the three axes with the greatest explanatory power for each lineage, indicated that the optimal trees had five branches, confirming the existence of five distinct genetic groups of queens within each lineage (Figure 3). These groups corresponded to spatially distinct patches of genetically similar queens; within these patches, colonies were generally <300 m apart. The mean between-group FST value was high and significantly different from zero (FST±SE=0.40±0.06 and 0.36±0.08 for lineages 1 and 2, respectively; permutation tests: P<0.001 for both), indicating a high degree of genetic differentiation among clusters of the same lineage.
Spatial genetic structure of C. mauritanica queens and males from each lineage. Each dot represents a colony. Graphical results of the spatial principal component analysis (sPCA) are shown for queens (a and b) and males (e and f). The first three components of the sPCA are summarised by translating scores of queens’ or males’ genotypes on each sPCA axis into a channel of colour (red, green and blue). Dots with similar colours represent genetically related males or genetically related queens within each lineage. For the sake of clarity, a single male genotype per colony has been chosen at random. In both lineages 1 and 2, spatially distinct groups of queens sharing a similar genetic profile are visible (a and b). In contrast, males display no clear genetic spatial pattern (e and f). Results of the Ward clustering analyses performed on the three sPCA axes explaining the most variance are also shown for queens (c and d). Genetically distinct clusters of queens belonging to (c) lineage 1 and (d) lineage 2 are depicted with different colours and shapes. All three branching criteria supported the existence of five clusters within each lineage. Grey, wooded areas; white, grasslands.
Consistent with the sPCA results, the relatedness among queens from the same lineage decreased with geographical distance. The Sp statistics were significantly positive for both lineages (Sp±SE=0.22±0.025 and 0.15±0.015 for lineages 1 and 2, respectively; P<0.001 for both; Figure 4).
Thus, the strongly spatially structured genetics of queens in the population shows that queens dispersed over short distances, which is consistent with colony foundation by budding.
Spatial genetic structure of the males
The allele frequencies of the males were analysed using sPCA; the tests were lineage specific (Figure 3). Neither global nor local structure was found in either lineage (Table 1). However, within each lineage, the relatedness among males decreased with geographical distance, although to a lesser extent than in queens (Figure 4). Indeed, spatial structuring was 6 and 12 times less pronounced in males than in queens for lineages 1 and 2, respectively (lineage 1: Sp±SE=0.035±0.005 and lineage 2: 0.012±0.003). That said, the randomisation test revealed significant spatial genetic structure in males (P<0.001 for both lineages), indicating that male dispersal was somewhat limited.
Male dispersal distances
For each lineage, 73% of males (n=425 for lineage 1 and 329 for lineage 2; Supplementary Table S2) had a genotype that fully matched that of at least one queen. For these males, the mean number of potential natal colonies±s.d. was 12.3±8.8 and 13.3±8.3 for lineages 1 and 2, respectively. Based on the sPCA and clustering results, the putative mothers of each male belonged to 1.0±0.0 and 1.4±0.6 clusters for lineages 1 and 2, respectively. This finding indicates that a unique natal cluster could be identified for most males. The median dispersal distance was 344 m for lineage-1 males (first–third quartile range: 228–472 m) and 274 m for lineage-2 males (first–third quartile range: 162–406 m; Figure 5).
Cumulative frequency distributions of male dispersal distances. The solid and dashed lines represent the distributions of distances between male mates and their potential natal colonies for lineages 1 and 2, respectively. The vertical lines indicate the medians: 344 m for lineage-1 males and 274 m for lineage-2 males.
Discussion
Our results confirm the existence of a dependent-lineage reproductive system in our C. mauritanica study population. All the workers were hybrids of the two genetic lineages. Not a single worker had been produced by parthenogenesis or by sexual reproduction between partners of the same lineage. Therefore, sustained interactions between the two lineages, as well as their coevolution, appear to be necessary to maintain worker production. If one lineage were to become more common than the other, queens could end up mating primarily with males of the same lineage and then fail to acquire compatible sperm for worker production. As a consequence, negative frequency-dependent selection should maintain a one-to-one ratio of the two lineages at equilibrium (Yamauchi and Yamamura, 2006). Consistent with this prediction, we found that the two lineages occurred at nearly the same frequency in the population (51.4 and 48.6% of colonies belonged to lineages 1 and 2, respectively).
Our study underscores how both the production of new queens by thelytokous parthenogenesis and colony budding can strongly limit gene flow and result in an extremely high level of spatial genetic structuring within populations. Indeed, the sPCA results showed that colonies headed by queens belonging to the same lineage formed well-differentiated clusters. Within each lineage, there was additional spatial genetic structure: colonies headed by genetically similar queens were aggregated in spatially distinct groups. Accordingly, we found a significant decrease in relatedness among queens as geographical distance increased, which was indicative of a lineage-specific IBD pattern. To the best of our knowledge, only a single study to date has explored the spatial distribution of genetic variability in a hybridogenetic ant species. Using an IBD model, Suni and Gordon (2010) found that the limited dispersal of Pogonomyrmex queens resulted in lineages displaying moderate levels of genetic structure. However, unlike Cataglyphis queens, Pogonomyrmex queens only reproduce sexually and disperse by flight. Queens mate with males of their own lineage as well as with males of the alternative lineage; purebred females develop into queens, and interlineage hybrids become workers (Helms Cahan and Keller, 2003). These breeding and dispersal strategies likely account for the relatively weaker spatial structure observed in Pogonomyrmex than in Cataglyphis.
Consistent with the greater dispersal abilities of males compared to queens, males displayed a lower level of spatial genetic structure. While the sPCA results provided no evidence of genetic structure in males within each lineage, there was a slight but significant IBD pattern: relatedness among males decreased as geographical distance increased. This discrepancy might stem from differences in detection power between the two statistical approaches. The significant IBD pattern indicates that male dispersal is somewhat limited at the scale of the population (which extends over 1000 m). In line with this, 73% of males had a genotype that matched that of at least one queen within the population. Since the queens formed highly differentiated genetic clusters, we were able to identify the natal cluster of these males, confirming that they came from the study population.
We also found that male dispersal distances ranged from 0 to 930 m (maximum observable distance between same-lineage colonies: 1031 m); on average, lineage 1 and lineage 2 males dispersed over 354.2 m and 299 m, respectively (Supplementary Table S2). These distances appear to be great enough to ensure interlineage mating since the mean distance between nearest-neighbour colonies of different lineages was 95.8±83 m (range: 2.8–318.3 m). Although males are the dispersing sex and can fly relatively long distances, we found that ~20% of mating events were intralineage for both lineages. Two complementary explanations may account for this result. First, our estimates indicate that 10% of males travelled <100 m from their natal nests, which increased the probability that they would encounter queens of the same lineage (Figure 5). Second, dispersal over longer distances does not guarantee interlineage mating because males can still land in a patch of queens of the same lineage. Taking into account mean male dispersal distances in this study population, a lineage-1 male had a 48% probability of ending up in a colony of his own lineage; this figure was 54% for a lineage-2 male. Thus, if mating events were governed by chance, one would expect that ~50% of encounters would be intralineage. The difference between this latter estimate and the observed percentage (that is, 20%) could be explained by two non-exclusive hypotheses. First, intralineage mating could be limited by prezygotic reproductive constraints, such as semiochemical-based recognition and discrimination (Oppelt et al., 2008). However, the fact that 20% of mating events still occurred between partners of the same lineage would indicate that barriers to assortative mating are only partially efficient. Second, it could be that intralineage mating does occur at a higher rate but is associated with lower fitness; for instance, queens may die prematurely because they cannot produce workers (that is, postzygotic isolation). However, if such were the case, postzygotic isolation could be somewhat relaxed in multiple-queen colonies of C. mauritanica because queens that engage in intralineage mating could nonetheless benefit from the workers produced by sister queens that engage in interlineage mating. Indeed, a previous study in the hybridogenetic species C. hispanica showed that intralineage mating occurred in populations with multiple-queen colonies, while interlineage mating was the rule in populations with single-queen colonies (Darras et al., 2014a). Interestingly, intralineage matings raise the potential for sexual conflicts. On the one hand, mating between same-lineage partners could enhance male fitness because it results in the production of purebred diploid offspring (likewise thelytokous parthenogenesis) that develop into new queens (Darras et al., 2014b). On the other hand, queens mated with males from the same lineage cannot produce worker offspring meaning they are unable of founding new colonies on their own. In addition, sexual reproduction has a fitness cost for queens because it dilutes by half the genes transferred to their reproductive daughters. Altogether, this should select mechanisms promoting intralineage mating in males, while impeding it in females.
Sex-biased dispersal is assumed to be widespread in social Hymenoptera (Johnstone et al., 2012). In species where individuals cannot be directly tracked (for example, because they engage in nuptial flights), estimates of sex-biased dispersal mostly rely on comparisons of maternally inherited mtDNA and biparentally inherited nuclear microsatellites. However, these two markers differ in their mutation rates, which results in variation in heterozygosity that can potentially bias estimates of dispersal differences between the sexes (Jost, 2008). Here, we used assignment analyses, and the results provide independent evidence that supports sex-biased dispersal. They showed that C. mauritanica males dispersed over 300 m, on average, which is 10 times further than queens, who tend to found new colonies ~30 m away from their natal nests (Knaden and Wehner, 2006). Thus far, only one study has directly quantified male dispersal distances in a natural population of ants. In an elegant examination of dispersal and inbreeding in the wood ant (Formica exsecta), Vitikainen et al. (2015) found clear evidence of sex-biased dispersal: the median dispersal distance of males (140 m) was twice that of queens (60 m).
In conclusion, we have confirmed that two equally common genetic lineages exist in our study population of Cataglyphis mauritanica and that queens and males belonging to these lineages interbreed. The parthenogenetic production and short-distance dispersal of queens resulted in strong spatial genetic structure both within and among lineages. In contrast, males dispersed over several hundred metres and thus travelled to non-natal patches, which helped contribute to interlineage mating. Thanks to their diverse breeding strategies, social insects can potentially provide novel insights into how singular reproductive systems shape the spatial genetic structure of populations.
Data archiving
Genotype data have been submitted to the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.7ht5j.
References
Balloux F, Lehmann L, de Meeûs T (2003). The population genetics of clonal and partially clonal diploids. Genetics 164: 1635–1644.
Borcard D, Gillet F, Legendre P (2011)Numerical Ecology with R. Springer New York: New York, NY, USA.
Charlesworth D, Charlesworth D, Willis JH, Willis JH (2009). The genetics of inbreeding depression. Nat Rev Genet 10: 783–796.
Chesser RK (1991). Influence of gene flow and breeding tactics on gene diversity within populations. Genetics 129: 573–583.
Clobert J, Baguette M, Benton TG (eds) (2012)Dispersal Ecology and Evolution. Oxford University Press: Oxford, UK.
Darras H, Leniaud L, Aron S (2014a). Large-scale distribution of hybridogenetic lineages in a Spanish desert ant. Proc R Soc Lond B 281: 20132396.
Darras H, Kuhn A, Aron S (2014b). Genetic determination of female castes in a hybridogenetic desert ant. J Evol Biol 27: 2265–2271.
Eyer PA (2014). Mode de reproduction et diversité génétique chez les fourmis du genre Cataglyphis. Phil. Thesis, Université Libre de Bruxelles.
Eyer PA, Leniaud L, Darras H, Aron S (2013). Hybridogenesis through thelytokous parthenogenesis in two Cataglyphis desert ants. Mol Ecol 22: 947–955.
Halkett F, Simon JC, Balloux F (2005). Tackling the population genetics of clonal and partially clonal organisms. Trends Ecol Evol 20: 194–201.
Hardy OJ (2003). Estimation of pairwise relatedness between individuals and characterization of isolation‐by‐distance processes using dominant genetic markers. Mol Ecol 12: 1577–1588.
Hardy OJ, Vekemans X (2002). SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2: 618–620.
Hartfield M, Wright SI, Agrawal AF (2016). Coalescent times and patterns of genetic diversity in species with facultative sex: effects of gene conversion, population structure, and heterogeneity. Genetics 202: 297–312.
Helms Cahan S, Keller L (2003). Complex hybrid origin of genetic caste determination in harvester ants. Nature 424: 306–309.
Hurley C (2012). gclus: Clustering graphics, R package version 1.3.1. Available at: https://CRAN.R-project.org/package=gclus.
Johnstone RA, Cant MA, Field J (2012). Sex-biased dispersal, haplodiploidy and the evolution of helping in social insects. Proc Biol Sci 279: 787–793.
Jombart T (2008). adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24: 1403–1405.
Jombart T, Devillard S, Dufour A-B, Pontier D (2008). Revealing cryptic spatial patterns in genetic variability by a new multivariate method. Heredity (Edinb) 101: 92–103.
Jost L (2008). GST and its relatives do not measure differentiation. Mol Ecol 17: 4015–4026.
Knaden M, Wehner R (2006). Fundamental difference in life history traits of two species of Cataglyphis ants. Front Zool 3: 21.
Leniaud L, Darras H, Boulay R, Aron S (2012). Social hybridogenesis in the clonal ant Cataglyphis hispanica. Curr Biol 22: 1188–1193.
Loiselle BA, Sork VL, Nason J, Graham C (1995). Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82: 1420–1425.
Maechler M, Rousseeuw P, Struyf A, Hubert M, Hornik K (2005). Cluster analysis basics and extensions. R package version 1.12.1.
Norman V, Darras H, Tranter C, Aron S, Hughes WOH (2016). Cryptic lineages hybridize for worker production in the harvester ant Messor barbarus. Biol Lett 12: 20160542.
Oppelt A, Spitzenpfeil N, Kroiss J, Heinze J (2008). The significance of intercolonial variation of cuticular hydrocarbons for inbreeding avoidance in ant sexuals. Anim Behav 76: 1029–1034.
Peakall R, Smouse PE (2012). GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28: 2537–2539.
Pearcy M, Clémencet J, Chameron S, Aron S, Doums C (2004). Characterization of nuclear DNA microsatellite markers in the ant Cataglyphis cursor. Mol Ecol Notes 4: 642–644.
Pearse DE, Anderson EC (2009). Multiple paternity increases effective population size. Mol Ecol 18: 3124–3127.
R Development Core Team (2016)R: A Language and Environment for Statistical Computing. R foundation for Statistical Computing: Vienna, Austria.
Rabeling C, Kronauer DJC (2013). Thelytokous parthenogenesis in eusocial hymenoptera. Annu Rev Entomol 58: 273–292.
Romiguier J, Fournier A, Yek SH, Keller L (2016). Convergent evolution of social hybridogenesis in Messor harvester ants. Mol Ecol 26: 1108–1117.
Ross KG (2001). Molecular ecology of social behaviour: analyses of breeding systems and genetic structure. Mol Ecol 10: 265–284.
Rousset F (2000). Genetic differentiation between individuals. J Evol Biol 13: 58–62.
Schurko AM, Neiman M, Logsdon JM (2009). Signs of sex: what we know and how we know it. Trends Ecol Evol 24: 208–217.
Schwander T (2016). Evolution: the end of an ancient asexual scandal. Curr Biol 26: R233–R235.
Schwander T, Oldroyd BP (2016). Androgenesis: where males hijack eggs to clone themselves. Philos Trans R Soc Lond B Biol Sci 371: 20150534.
Sirviö A, Pamilo P, Johnson RA, Page RE, Gadau J (2011). Origin and evolution of the dependent lineages in the genetic caste determination system of Pogonomyrmex ants. Evolution 65: 869–884.
Suni SS, Gordon DM (2010). Fine-scale genetic structure and dispersal distance in the harvester ant Pogonomyrmex barbatus. Heredity (Edinb) 104: 168–173.
Trible W, Kronauer DJ (2017). Caste development and evolution in ants: it’s all about size. J Exp Biol 220: 53–62.
Vallejo-Marín M, Dorken ME, Barrett SCH (2010). The ecological and evolutionary consequences of clonality for plant mating. Annu Rev Ecol Evol Syst 41: 193–213.
Vekemans X, Hardy OJ (2004). New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13: 921–935.
Vitikainen EIK, Haag-Liautard C, Sundström L (2015). Natal dispersal, mating patterns, and inbreeding in the ant formica exsecta. Am Nat 186: 716–727.
Yamauchi A, Yamamura N (2006). Persistence conditions of symmetric social hybridogenesis in haplo-diploid hymenoptera. J Theor Biol 240: 24–31.
Acknowledgements
We thank O. Hardy and three reviewers for their critical evaluation of the manuscript. We are grateful to Morocco’s High Commission for Water, Forests and Combatting Desertification (HCEFLCD; Decision 12/2015) for granting us collection permits. We thank J Pearce-Duvet for her language editing services. This research was supported by PhD fellowships funded by the Belgian National Fund for Scientific Research (FRS-FNRS; FRIA fellowship to AK, and ASP fellowships to DB and HD) as well as by research grants from FRS-FNRS (2.4516.11; J.0063.14) and the Université Libre de Bruxelles (ARC 2010-2015 #5; to SA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website .
Supplementary information
Rights and permissions
About this article
Cite this article
Kuhn, A., Bauman, D., Darras, H. et al. Sex-biased dispersal creates spatial genetic structure in a parthenogenetic ant with a dependent-lineage reproductive system. Heredity 119, 207–213 (2017). https://doi.org/10.1038/hdy.2017.34
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.34
This article is cited by
-
Genetic diversity, paternal origin and pathogen resistance in Cataglyphis desert ants
Behavioral Ecology and Sociobiology (2023)
-
Evidence of spatial genetic structure in a snow leopard population from Gansu, China
Heredity (2021)
-
Population structure and sociogenetic organisation in a species with ergatoid queens, the desert ant Ocymyrmex robustior
Insectes Sociaux (2019)