Abstract
Despite the ubiquity of coinfection, we know little of the effects of intra-specific genetic variability on coinfection by distinct parasite species. Here we test the hypothesis that parasite multiplication depends on the combination of parasite genotypes that coinfect the host (that is Genotype.parasite × Genotype.parasite interaction). To that aim, we infected tomato leaves with the ecto-parasitic mites Tetranychus urticae and Tetranychus evansi. We tested all possible combinations between four T. urticae and two T. evansi populations sampled on different hosts or localities. There was no universal (that is genotype-independent) effect of coinfection on mite multiplication; in many cases the two species had no effect on each other. However, several combinations of T. evansi and T. urticae populations led to elevated T. evansi numbers. Similarly, T. urticae reproduction largely depended on the interaction between T. urticae and T. evansi populations. This evidence for genotype-by-genotype interaction between coinfecting parasites indicates that the effect of coinfection on parasite epidemiology and evolution may vary in space according to the genetic composition of local parasite populations; it further suggests the possibility of coevolution between parasites species that share the same hosts.
Similar content being viewed by others
Introduction
Understanding parasite epidemiology and evolution is a challenging task due to the large number of factors involved. One reason for this complexity is that parasites often share their hosts with other species with which they can interact (Pedersen and Fenton, 2007; Rigaud et al., 2010). Coinfection by several parasites of distinct species can have both positive or negative effects on parasite fitness (Pedersen and Fenton, 2007). For example, HIV infection, and its associated immuno-suppression, facilitates infection by other infectious diseases such as tuberculosis (Narain et al., 1992). Alternatively, infection by Schistosoma parasites has been suggested to protect hosts from malarial infection (Briand et al., 2005). The environment can also modulate the effect that two parasites have on each other. For instance, the outcome of the competition between mosquito parasites can depend on the amount of food available to the host and infectious doses (Fellous and Koella, 2009). Unfortunately, logistic and conceptual limitations often restrict empirical approaches to simple designs unable to reflect the diversity of relationships that may occur between parasites. Indeed, most studies on coinfection by different parasite species neglect their intra-specific genetic variability (Pedersen and Fenton, 2007; Rigaud et al., 2010). This source of variability is however very important as parasite populations are often genetically heterogeneous in space (for example Boubou et al., 2011; Criscione and Blouin, 2004; Prugnolle et al., 2005) leading to different epidemiologies in different contexts (for example Fellous et al., 2012) and because genetic variability is essential to evolution and coevolution.
In some cases, parasite genetic factors have been shown to determine whether a parasite suffers from, benefits or is unaffected by the presence of a second parasite. Indeed, parasite fitness can depend on the genetics of the focal parasite (Hodgson et al., 2004; Seppälä et al., 2009), as well as that of the parasite it shares its host with (Bell et al., 2006). However, the combined, simultaneous effect of both parasite genetics—that is Genotype.parasite × Genotype.parasite interactions (further referred to as Gp.Gp interactions)—on their success has received scant attention. We know however that Gp.Gp interactions can be influential as experiments with two fish parasites of the Diplostomum genus found that whether D. pseudospathaceum succeeds, or fails, at infecting depends on an interaction between its genotype and that of its competitor (Seppälä et al., 2012). In the present study, we focus on parasite multiplication once taxa have encountered on the same host. These approaches are complementary for two reasons. First, parasite fitness depends on infection success (Seppälä et al., 2012) but also on multiplication within (and therefore transmission from) the host, all these stages being important. Second, theoretical models of parasite dynamics that consider parasite–parasite relationships usually distinguish between superinfection (that is when only one of several competing parasites survives in the host) from coinfection (that is when several parasites coexist in the host) (Adler and Mosquera, 2002; Alizon et al., 2013). Among other things, coinfection would produce more virulent infection than superinfection; superinfection is predicted to favour greater diversity in parasite population than coinfection (Alizon, 2013).
To investigate the effect of Gp.Gp interactions between parasites we studied the coinfection between two phytophagous, ectoparasitic mites, Tetranychus evansi and Tetranychus urticae. Plant immune response to these mites include proteinase inhibitors that are reguated by jasmonic acid and salicylic acid pathways. T. evansi can manipulate host immune system and down-regulate the production of proteinase inhibitors (Sarmento et al., 2011a). By contrast, T. urticae has not been shown to inihibit plant defences, but some genotypes are less prone to activating them than others (Kant et al., 2008). These two species have also been shown to influence each other’s growth rate in captivity (Sarmento et al., 2011b) and distribution in the wild (Ferragut et al., 2013). We experimentally established competition between the two species placing several adult mites of each species on the same tomato leaf, a host plant where natural coinfection occurs in the field (Alain Migeon, personal communication; Simon Fellous, personal observation). We recorded offspring production and competition avoidance behaviour over one cycle of reproduction. Mite genotypes were manipulated by using four T. urticae populations collected on different host plant species: tomato (Solanum lycopersicum), citrus tree (Citrus aurantium) and rose bay (Nerium oleander) and two T. evansi populations representing the two major phylotypes encountered in Europe (Boubou et al., 2011) both sampled on tomato (Solanum lycopersicum). Choice of assay population was not trivial as coevolution and local adaptation can deplete genetic diversity and conceal potent Gp.Gp interactions (Heath and Nuismer, 2014). We hence chose to contrast allopatric populations of mites that were sampled on different host species and localities and we knew were genetically distinct. The life-histories of the four T. urticae populations we employed were investigated in a previous study (Figure 1 in Fellous et al., 2014), revealing reproducible phenotypic—and therefore genetically based—differences among them.
Materials & methods
Biological system
The two-spotted spider mite, T. urticae, is a cosmopolitan and highly polyphagous mite found on more than a thousand plant species (Migeon and Dorkeld, 2006). This mite feeds by piercing leaf parenchyma cells with its stylet and sucking out cell content (Tomczyk and Kropczynska, 1985). The species has an arrhenotokous reproductive system: diploid females produce haploid males with unfertilised eggs (Helle and Bolland, 1967). A life cycle can be completed in as little as 10 days on suitable hosts and at optimal temperature (c. 23 °C). After mated-females colonise new host plants, mites undergo several reproductive cycles until reaching carrying capacity or plant death (Helle and Sabelis, 1985).
The biology of T. evansi is in many ways similar to that of T. urticae. However it has a narrower host range as it is mostly found on solanaceous plants. It has a recent history of invasion: two major phylotypes are present in Europe (Boubou et al., 2011). The first phylotype invaded Africa and later on Europe, whereas the second one seemingly arrived directly from South America.
All mites were maintained before the experiment in our standard rearing conditions. T. urticae were fed detached leaves of bean, Phaseolus vulgaris, while T. evansi received Solanum nigrum leaves. These leaves were placed on water-soaked cotton wool in plastic boxes in a controlled-temperature chamber at 23 °C with 16 h of light per day.
Experimental populations
Our aim being to study the effects of intra-specific genetic variability on mite development, we selected experimental populations that we were certain would exhibit genetic differences based on the available literature and our experience. We thus assayed populations that originated from distinct regions and host species. Recent theory shows that restricting analyses to sympatric populations increases the risk of false negatives, that is of not finding G × G patterns even though they influence evolution (Heath and Nuismer, 2014). This derives from the strong depleting effects that adaptation and coevolution can have on standing genetic variation. The most common genotypes (that is those that would be most likely to be sampled) may thus not reflect existing genetic diversity. In other words, statistical interactions between genotypes from the same locality may largely underestimate the spread of functional genotype-by-genotype interactions at play in the field. However, G × G patterns can only lead to coevolution if assayed genotypes do encounter. In other words, for functional interactions between genotypes to be more than laboratory artifacts and influence natural processes, chosen genotypes must be representative of natural genetic diversity (see details in Heath and Nuismer, 2014).
Our choice of study populations followed the reasonning described in the previous paragraph. For T. urticae we used four populations from three different host plants (Solanum lycopersicum, Nerium oleander and Citrus aurantium) and three south-European localities. Published data and our own on-going population genetics work shows that, despite mite specialisation to the host plant (Fellous et al., 2014), there is substantial gene flow between T. urticae populations encountered on these plants and localities (Magalhães et al., 2007). For T. evansi we used each of the two European phylotypes (Boubou et al., 2011), which geographic distributions converge in southern Europe.
T. urticae populations were found c. 6 months earlier by a small number of individuals collected in the field and experienced strong bottlenecks during their first generations in the lab (Fellous et al., 2014). Two populations derived from individuals collected on tomato, S. lycopersicum: Population A was sampled in southern France and population B in Crete. Another population derived from individuals collected on N. olander, and the last one from individuals collected on C. aurantium, both in Spain. These are further referred to as T. urticae populations C and D, respectively. Genetic (or heritable epigenetic) differences between these populations are supported by their significantly different performance on tomato and other plants (Fellous et al., 2014). We used two T. evansi populations, both collected on tomato, from each of the two major phylotypes encountered in Europe (Boubou et al., 2011). The T. evansi populations, further referred to as one and two, correspond to phylotypes one and two, respectively, as described in (Boubou et al., 2011). Population one was sampled at Beausoleil (south-east of France) and population two near Sainte-Marie-la-Mer (south-west of France).
Experimental design
All combinations of T. urticae and T. evansi populations were tested with a full-factorial design (for coinfection treatments). We also included single infections of each species as controls treatments. There were thus eight coinfection treatments and six single infection treatments.
Replicates were organised in five blocks, each containing every treatment, and were run one week apart. Mites used for each block were produced independently by having c. 50 females of each population lay eggs on a tomato leaf during 24 h.
An experimental unit (that is, one replicate) was made of two tomato leaflets (S. lycopersicum, cultivar Heinz) placed on a bed of cotton wool soaked with water. For simplicity, we refer to these leaflets as leaves thorough the manuscript. Leaves partly overlapped (c. 2% of surface) to allow mite movement between them. Three adult, 3–7 days old females and one adult male per species (that is, the parental generation) were placed on the inoculated leaf at the beginning of the experiment (Figure 1). Coinfection treatments thus received twice more mites than single infection treatments, so as to study the effect of parasite co-occurrence without changing the initial density of the focal species. Adults were left on the leaves for five days, when we removed them after recording their survival and position (on the inoculated or non-inoculated leaves; Figure 1). This brief oviposition period synchronised the development of mite offspring and therefore avoided generation overlaps. It was an essential point in our protocol as the morphological differences between T. urticae and T. evansi become conspicuous at the adult stage only. Counts of adult offspring, and their position, occurred 18 days post-inoculation, after one reproductive cycle (Figure 1).
We studied dispersal by depositing, at the beginning of experiment, the individuals on a single tomato leaf (that is inoculated) in contact with a second leaf (that is non-inoculated) on the same cotonwool bed. Dispersal was estimated, after 5 and 18 days, as the proportion of individuals that were present on the non-inoculated leaf.
The experiment was conducted in a controlled-temperature chamber at 23 °C with 16 h of light per day.
Statistical analyses
We used generalised linear models (GLM) to assess the effects of T. urticae and T. evansi genotypes on mite multiplication. To analyse the numbers of live individuals 18 days post-inoculation (that is, the response variable) we employed models with Poisson distributions and log links. We first tested whether mite numbers were affected by the presence of the other mite species independent of its genotype (that is, general effects of coinfection). Initial models contained two factors describing the population of the focal mite species, a factor describing the absence or presence of the second mite species and their interaction. Later, we investigated interactions between mite genotypes with models containing the two factors describing their genotypes and their interaction. Mite genotypes were treated as fixed factors because they did not represent random selections of mite populations but were selected for their a-priori differences. Block was included in early models but not in the final ones as it was never significant. We used least-squares contrasts with likelihood-ratios tests for post-hoc tests. Analyses were carried out with the statistical package JMP 10.0.2.
Note that since we manually infected hosts with adult parasites, and therefore by-passed the first step of natural infections, we focused on the quantitative multiplication of mites not taking into account behaviours associated host plant choice.
For brevity, analyses of survival at the parental population (that is, after 5 days) and mite movement to the non-inoculated leaf after 18 days are reported in Supplementary Materials (Supplementary Tables S1 and S3,Supplementary Figures S1 and S2).
Results
Parasite survival and reproduction
After 18 days, that is after approximately one reproductive cycle, the presence of T. urticae (independent of its population) had a marginally non-significant effect on T. evansi numbers (X2=3.78, df=1, P=0.052). T. urticae presence indeed tended to facilitate the development of T. evansi (Figure 2). There was however no general effect (that is population-independent) of T. evansi presence on T. urticae numbers (X2=0.0001, df=1, P=0.99).
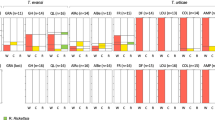
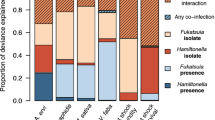
Instead, effects of the two mite species on each other turned out to be population specific, as revealed by the significant interactions between T. urticae and T. evansi populations (Table 1,Figures 2 and 3). While T. evansi population one benefited from the presence of all T. urticae populations (contrast With T. urticae population A, B, C and D vs Without T. urticae: X2=4.45, df=1, P=0.035) (Figure 2), T. evansi population two only benefited from the presence of T. urticae population C (contrast With T. urticae population C vs Without T. urticae: X2=37.8, df=1, P=0.009) (Figure 2). The interaction between T. urticae and T. evansi populations remained marginally non-significant (P=0.055) when only using the T. urticae populations sampled on tomato (Supplementary Table S2).
Similarly, the identity of the T. evansi population had a significant effect on T. urticae numbers (see significant interaction between T. urticae and T. evansi populations in Table 1, Figure 3). For example, in the case of T. urticae population B, T. evansi population two led to greater mite numbers than with T. evansi population one (contrast With T. evansi population two vs With T. evansi population one: X2=6.27, df=1, P=0.012). Interestingly, these same two T. evansi populations tended to have opposite effects on T. urticae population D, as these were more numerous in the presence of T. evansi population one than with population two (contrast With T. evansi population one vs With T. evansi population two: X2=3.5, df=1, P=0.061). The interaction between T. urticae and T. evansi populations was not significant anymore when only using the T. urticae populations sampled on tomato (Supplementary Table S2). This is not surprising as T. urticae population C, which was sampled on Nerium oleander, was excluded.
Within host movements and competition avoidance
In our experimental set-up, parental mites were deposited on one of two adjacent leaves (that is the inoculated leaf). There were very little movements between the inoculated and non-inoculated leaves during the 5 days before parents were removed, hence preventing us from testing the hypothesis that competitor avoidance was controlled by interactions between T. evansi and T. urticae genotypes. However, we noticed that the only three instances where T. evansi females were found on the non-inoculated leaf were in the presence of T. urticae population C (Supplementary Figure S3). A Fisher’s exact test revealed that T. urticae population C did increase T. evansi movement from the inoculated to the non-inoculated leaf (P=0.008). There was nonetheless no effect of T. urticae population identity on T. urticae movements after 5 days: analyses of mite positions after 18 days provided no support for interactions between mite populations and are reported as Supplementary materials.
Discussion
This experiment revealed that when two parasites of different species coinfect the same host their multiplication can be determined by interactions between their genotypes (that is Gp.Gp interaction). More precisely, we found that, during the first reproductive cycle of an infection (that is 18 days), numbers of ecto-parasitic mites T. urticae and T. evansi either increased or remained unaffected by the presence of the other species, and that this depended on the combination of mite populations in presence.
Epidemiological and evolutionary implications
Genotype-by-genotype interactions are important for ecological dynamics as they foster different demographics in localities populated by different genotypes. Since parasite populations are often genetically structured (that is, all genotypes are not encountered everywhere) (for example Boubou et al., 2011; Criscione and Blouin, 2004; Prugnolle et al., 2005), Gp.Gp interactions suggest that the effect each parasite has on the others should differ among locations. In some cases, one species may tend to exclude the other while favouring its spread in other locations—as when T. evansi population two encountered T. urticae population C in our experiment.
For coevolution between two organism to occur, a number of conditions must be filled. The first one is that their fitness depends on functional genotype-by-genotype interactions (Health and Nuismer, 2014; Thompson, 2005). These refer to situations where the phenotype of an organism is determined by the interaction between its genome and that of the other organism it may coevolve with. Functional genotype-by-genotype interactions can be characterised at the molecular level, but also evidenced by statistical genotype-by-genotype interactions, as we do here (Heath and Nuismer, 2014). It is also important that, for each species, focal genotypes may not be restricted to isolated localities, a condition that would be fulfilled by our choice of mite populations (see Methods). Our results therefore suggest possible coevolution between parasites that share the same host. This type of coevolution would be analogous to better-known host-parasite coevolution—which is based on Genotype.host × Genotype.parasite interactions (Agrawal and Lively, 2002), and coevolution between mutualists of different trophic levels (for example Health et al., 2012; Thompson, 2005). In addition, coevolution between parasite species would necessitate that the two species co-occur frequently on the same hosts so as to substantially affect the selective pressures they are under. As such, it may be faster during epidemic phases or when hosts exhibit low resistance to infection, hence creating so-called coevolutionary hot-spots (Thompson, 2005).
The composition of the host population also exerts strong selective pressures: parasites, such as T. urticae, readily adapt to specific host species or genotypes (Carletto et al., 2009; Fry, 1990; Legros and Koella, 2010; Magalhães et al., 2007). The T. urticae populations we used in our experiment were sampled on three different host plant species. Therefore, the different phenotypes of our T. urticae populations that were expressed in situation of coinfection may result from their specialisation to different hosts. It would explain why T. urticae population C stood out as eliciting both dispersal movements of T. evansi (Supplementary Figure S3) and significantly increasing T. evansi multiplication (Figure 2). When parasites’ genotype-specific effects are due to adaptation to different hosts, local composition of host community would control coinfection outcomes through parasite specialisation and local-adaptation.
The effect of coinfection on the evolution of parasite virulence has been extensively studied theoretically (Mideo et al., 2008; Alizon, 2013). In most models predicted evolutionary trajectory relies on the effects coinfecting parasites have on each other. However, no model to our knowledge takes Gp.Gp interactions into account. Indeed, they all assume that coinfection is either facilitative, neutral or detrimental; not that it varies according to the players involved. The integration of such variability would be an important challenge associated with reconciling theory on virulence evolution with empirical data (Alizon et al., 2013).
Underlying processes
The mechanism behind the interactions between T. evansi and T. urticae are unknown, identifying them would go beyond the aim of the present work. Since the mites mostly exhibited facilitative effects, classical means of competition—such as resource limitations and spiteful interactions—do not apply. Similarly, it was reported that T. evansi can alter T. urticae population growth by depositing silk-web on the leaf surface (Sarmento et al., 2011b). But web production was negligible in our experiment. We estimated mite performance by counting the number of offspring produced on plants exposed for a short to adult mites. This would accurately reflect reproductive output rate (that is number of offspring per unit of time). But other components of fitness such as life-time reproductive success (which depends on lifespan) and offspring condition were not taken into account.
Genotype-by-genotype interactions are best know between hosts and parasites (Lambrechts et al., 2006). They can have significant effects on epidemiology (Fellous et al., 2012) and underly host-parasite coevolution (Betts et al., 2014). Genotype.host × Genotype.parasite interactions often result from genotype-specific effects of parasites on host immunity (Lambrechts et al., 2013). A similar mechanism is therefore likely for Gp.Gp interactions. Some T. urticae populations activate plant immunity, while others manage to not activate it (Alba et al., 2015; Kant et al., 2008). It was also reported that T. evansi can inhibit plant defences (Sarmento et al., 2011a). We conducted our experiment on detached leaves rather than whole plants, some immune components may thus have been altered (Schmelz et al., 2001). But published data shows that local (that is non-systemic) defences are potent against T. evansi and T. urticae (Sarmento et al., 2011a). Therefore, differences among mite populations and species in their ability to activate and resist plant defences likely contribute to population-specific interactions between these parasites (Paterson, 2013). In fact, this may even be a general mechanism for Gp.Gp interactions as indirect interactions between parasites are often mediated by the host immune system (Frank, 2002). However, the role of host immunity for Gp.Gp interactions was recently tested with Diplostomum parasites of fish—which infection success rely on Gp.Gp interactions (Seppälä et al., 2012)—finding no evidence for genotype-specific priming of the host immune system (Rellstab et al., 2013).
Competitive interactions between T. evansi and T. urticae would have led to a reduction in T. urticae abundance and effective host range in southern Europe after T. evansi invasion (Ferragut et al., 2013). On tomato, competition was investigated by Sarmento and collaborators (Sarmento et al., 2011b). In their experiment T. evansi reduced T. urticae numbers within one generation. This discrepancy with our results may derive from genetic differences among the mite populations assayed as these authors only tested one population per species. Similarly, the host plant cultivar used by Sarmento (Santa Clara I-5300, CA, USA) is different from ours. Cultivars of the same plant species often vary broadly in terms of defences against parasites and pests (Krips et al., 1998; Xuéreb and Thiéry, 2006). The putative importance of plant immunity (and mite counter-defences) for Gp.Gp interactions suggests that host genetic identity may have been an important factor. Prior experience may have played a role too: we produced the parental generation of T. urticae on bean rather than tomato as in Sarmento et al. (2011b). Mite ability to exploit certain hosts is indeed known to be influenced by the host species that previous generations infected (Magalhães et al., 2011). Finally, Sarmento et al. (2011b) carried-out their experiment on whole tomato plants, while we used detached leaves, as is common for T. urticae phenotyping (for example Agrawal et al., 2002; Magalhães et al., 2011). All these elements illustrate how using lab data to make quantitative prediction on parasite dynamics may be deceptive. It is clear Gp.Gp interactions occur, but their importance to field dynamics remains to be elucidated.
Conclusions
Our experiment revealed that the multiplication of two coinfecting plant ectoparasites relied on Genotype.parasite × Genotype.parasite interactions. Indeed, the combination of parasite populations present determined mite reproduction and whether coinfection was beneficial or neutral. It follows that the local genetic composition of parasite populations may determine the reciprocal influence two coinfecting parasites have on each other and cascade on their epidemiologies. Ultimately this might lead to coevolution between parasites sharing the same host. Besides, the genetic identity of co-occurring parasites being determined by local selective pressures (for example nature of available hosts) coinfection outcome may vary with local host community composition.
Data archiving
Data available from the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.fg271.
References
Adler FR, Mosquera J . (2002) Super- and coinfection: filling the range. In: Dieckmann U, Metz JAJ, Sabelis MW, Sigmund K (eds), Adaptive Dynamics of Infectious Diseases: In Pursuit of Virulence Management. Cambridge University Press: Cambridge, MA, USA.
Agrawal A, Lively CM . (2002). Infection genetics: gene-for-gene versus matching-alleles models and all points in between. Evol Ecol Res 4: 79–90.
Agrawal AA, Vala F, Sabelis MW . (2002). Induction of preference and performance after acclimation to novel hosts in a phytophagous spider mite: adaptive plasticity? Am Nat 159: 553–565.
Alba JM, Schimmel BCJ, Glas JJ, Ataide LMS, Pappas ML, Villarroel CA et al. (2015). Spider mites suppress tomato defenses downstream of jasmonate and salicylate independently of hormonal crosstalk. New Phytol 205: 828–840.
Alizon S . (2013). Co-infection and super-infection models in evolutionary epidemiology. Int Focus 3: 20130031.
Alizon S, de Roode JC, Michalakis Y . (2013). Multiple infections and the evolution of virulence. Ecol Lett 16: 556–567.
Bell AS, de Roode JC, Sim D, Read AF . (2006). Within-host competition in genetically diverse malaria infections: parasite virulence and competitive success. Evolution 60: 1358–1371.
Betts A, Kaltz O, Hochberg ME . (2014). Contrasted coevolutionary dynamics between a bacterial pathogen and its bacteriophages. Proc Natl Acad Sci USA 111: 11109–11114.
Boubou A, Migeon A, Roderick G, Navajas M . (2011). Recent emergence and worldwide spread of the red tomato spider mite, Tetranychus evansi: genetic variation and multiple cryptic invasions. Biol Inv 13: 81–92.
Briand V, Watier L, Le Hesran J-Y, Garcia A, Cot M . (2005). Coinfection With Plasmodium falciparum And Schistosoma haematobium: Protective effect of schistosomiasis on malaria in senegalese children? Am J Trop Med Hyg 72: 702–707.
Carletto J, Lombaert E, Chavigny P, Brévault T, Lapchin L, Vanlerberghe-Masutti F . (2009). Ecological specialization of the aphid Aphis gossypii Glover on cultivated host plants. Mol Ecol 18: 2198–2212.
Criscione CD, Blouin MS . (2004). Life cycles shape parasite evolution: comparative population genetics of salmon trematodes. Evolution 58: 198–202.
Fellous S, Angot G, Orsucci M, Migeon A, Auger P, Olivieri I et al. (2014). Combining experimental evolution and field population assays to study the evolution of host range breadth. J Evol Biol 27: 911–919.
Fellous S, Duncan AB, Quillery E, Vale PF, Kaltz O . (2012). Genetic influence on disease spread following arrival of infected carriers. Ecol Lett 15: 186–192.
Fellous S, Koella JC . (2009). Infectious dose affects the outcome of the within-host competition between parasites. Am Nat 173: E177–E184.
Ferragut F, Garzón-Luque E, Pekas A . (2013). The invasive spider mite Tetranychus evansi (Acari: Tetranychidae) alters community composition and host-plant use of native relatives. Exp Appl Acar 60: 321–341.
Frank SA . (2002) Immunology and Evolution of Infectious Disease. Princeton University Press: Princeton.
Fry JD . (1990). Trade-offs in fitness on different hosts: evidence from a selection experiment with a phytophagous mite. Am Nat 136: 569–580.
Heath KD, Burke PV, Stinchcombe JR . (2012). Coevolutionary genetic variation in the legume-rhizobium transcriptome. Mol Ecol 21: 4735–4747.
Heath KD, Nuismer S . (2014). Connecting functional and statistical definitions of genotype by genotype interactions in coevolutionary studies. Front Genet 5: 77.
Helle W, Bolland HR . (1967). Karyotypes and sex-determination in spider mites (Tetranychidae). Genetica 38: 43–53.
Helle W, Sabelis MW . (1985) Spider Mites: Their Biology, Natural Enemies and Control. Elsevier: Amsterdam.
Hodgson DJ, Hitchman RB, Vanbergen AJ, Hails RS, Possee RD, Cory JS . (2004). Host ecology determines the relative fitness of virus genotypes in mixed-genotype nucleopolyhedrovirus infections. J Evol Biol 17: 1018–1025.
Kant MR, Sabelis MW, Haring MA, Schuurink RC . (2008). Intraspecific variation in a generalist herbivore accounts for differential induction and impact of host plant defences. Proc R Soc Lond B 275: 443–452.
Krips OE, Witul A, Willems PEL, Dicke M . (1998). Intrinsic rate of population increase of the spider mite Tetranychus urticae on the ornamental crop gerbera: intraspecific variation in host plant and herbivore. Ent Exp Appl 89: 159–168.
Lambrechts L, Fellous S, Koella JC . (2006). Coevolutionary interactions between host and parasite genotypes. Trends Parasitol 22: 12–16.
Lambrechts L, Quillery E, Novell V, Richardson JH, Jarman RG, Scott TW et al. (2013). Specificity of resistance to dengue virus isolates is associated with genotypes of the mosquito antiviral gene Dicer-2. Proc R Soc Lond B 280: 1751.
Legros M, Koella J . (2010). Experimental evolution of specialization by a microsporidian parasite. BMC Evol Biol 10: 159.
Magalhães S, Blanchet E, Egas M, Olivieri I . (2011). Environmental effects on the detection of adaptation. J Evol Biol 24: 2653–2662.
Magalhães S, Forbes M, Skoracka A, Osakabe MR, Chevillon C, McCoy KD . (2007). Host race formation in the Acari. Exp Appl Acar 42: 225–238.
Mideo N, Alizon S, Day T . (2008). Linking within-and between-host dynamics in the evolutionary epidemiology of infectious diseases. Trends Ecol Evol 23: 511–517.
Migeon A, Dorkeld F . (2006–2016). Spider mites web: a comprehensive database for the Tetranychidae. INRA. Available at https://www1.montpellier.inra.fr/CBGP/spmweb/.
Narain JP, Raviglione MC, Kochi A . (1992). HIV-associated tuberculosis in developing countries: epidemiology and strategies for prevention. Tuber Lung Dis 73: 311–321.
Paterson S . (2013). The immunology and ecology of co-infection. Mol Ecol 22: 2603–2604.
Pedersen AB, Fenton A . (2007). Emphasizing the ecology in parasite community ecology. Trends Ecol Evol 22: 133–139.
Prugnolle F, Theron A, Pointier JP, Jabbour-Zahab R, Jarne P, Durand P et al. (2005). Dispersal in a parasitic worm and its two hosts: consequence for local adaptation. Evolution 59: 296–303.
Rellstab C, Karvonen A, Louhi K-R, Jokela J . (2013). Genotype-Specific vs Cross-Reactive Host Immunity against a Macroparasite. PLOS One 8: e78427.
Rigaud T, Perrot-Minnot M-J, Brown MJF . (2010). Parasite and host assemblages: embracing the reality will improve our knowledge of parasite transmission and virulence. Proc R Soc Lond B 277: 3693–3702.
Sarmento RA, Lemos F, Bleeker PM, Schuurink RC, Pallini A, Oliveira MGA et al. (2011)a. A herbivore that manipulates plant defence. Ecol Lett 14: 229–236.
Sarmento RA, Lemos F, Dias CR, Kikuchi WT, Rodrigues JCP, Pallini A et al. (2011b). A herbivorous mite down-regulates plant defence and produces web to exclude competitors. PLOS One 6: e23757.
Schmelz EA, Alborn HT, Tumlinson JH . (2001). The influence of intact-plant and excised-leaf bioassay designs on volicitin-and jasmonic acid-induced sesquiterpene volatile release in Zea mays. Planta 214: 171–179.
Seppälä O, Karvonen A, Rellstab C, Louhi K-R, Jokela J . (2012). Reciprocal interaction matrix reveals complex genetic and dose-dependent specificity among coinfecting parasites. Am Nat 180: 306–315.
Seppälä O, Karvonen A, Tellervo Valtonen E, Jokela J . (2009). Interactions among co-infecting parasite species: a mechanism maintaining genetic variation in parasites? Proc R Soc Lond B 276: 691–697.
Thompson JN . (2005) The Geographic Mosaic of Coevolution. The University of Chicago Press: Chicago.
Tomczyk A, Kropczynska D . (1985) Effects on the host plant. In: Helle W, Sabelis MW (eds), Spider mites, their biology, natural enemies and control. Elsevier: Amsterdam Vol. 1A, pp 317–329.
Xuéreb A, Thiéry D . (2006). Does natural larval parasitism of Lobesia botrana (Lepidoptera: Tortricidae) vary between years, generation, density of the host and vine cultivar? Bull Entomol Res 96: 105–110.
Acknowledgements
We are grateful to G Angot, P Auger, A Migeon and A Tsagakarakou for helping in the lab and mite collection in the field. We also thank O Kaltz and A Duncan for comments on the manuscript. This project operated as cofounding for the GI-046 grant from Genome Canada and the Ontario Genomics Institute and the GL2-01-035 grant from the Ontario Research Fund–Global Leadership in Genomics and Life Sciences. Funding was provided by the French Agence Nationale de la Recherche (ANR 2010 BLAN 1715 02).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Orsucci, M., Navajas, M. & Fellous, S. Genotype-specific interactions between parasitic arthropods. Heredity 118, 260–265 (2017). https://doi.org/10.1038/hdy.2016.90
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.90
This article is cited by
-
Arbuscular mycorrhizal fungi and belowground herbivores interact to determine plant productivity and performance of subsequently feeding conspecifics by changing plant metabolites
Journal of Pest Science (2023)
-
Ecology and evolution of facilitation among symbionts
Nature Communications (2018)
-
Identification of spider-mite species and their endosymbionts using multiplex PCR
Experimental and Applied Acarology (2018)