Abstract
Feral livestock may harbor genetic variation of commercial, scientific, historical or esthetic value. The origins and uniqueness of feral cattle on Chirikof Island, Alaska, are uncertain. The island is now part of the Alaska Maritime Wildlife Refuge and Federal wildlife managers want grazing to cease, presumably leading to demise of the cattle. Here we characterize the cattle of Chirikof Island relative to extant breeds and discern their origins. Our analyses support the inference that Yakut cattle from Russia arrived first on Chirikof Island, then ~120 years ago the first European taurine cattle were introduced to the island, and finally a large wave of Hereford cattle were introduced on average 40 years ago. In addition, this mixture of European and East-Asian cattle is unique compared with other North American breeds and we find evidence that natural selection in the relatively harsh environment of Chirikof Island has further impacted their genetic architecture. These results provide an objective basis for decisions regarding conservation of the Chirikof Island cattle.
Similar content being viewed by others
Introduction
Contemporary cattle in the harsh environment of Chirikof Island are largely isolated in the North Pacific Ocean, unmanaged, and thought to descend from many generations of feral stock (McKnight, 1964). Feral livestock, animal populations that were formerly domesticated but currently live independently of humans, may be sources of genetic variation with potential commercial, scientific, historical or esthetic value (van Vuren and Hedrick, 1989). Thus, the cattle on Chirikof Island may have genetic variants that are rare or absent in domesticated breeds that are used in commerce. Sources of this variation include founder effects, random drift and mutation followed by natural selection within the population to confer adaptation to the particular environmental conditions present on Chirikof Island.
Specific origins of the feral cattle population on Chirikof Island are ambiguous. Cattle farming was common in Yakutia (Sakha), Russian Siberia, throughout the period of Russian colonial possessions in the Americas (from 1733 to 1867) (Kotkin and Wolff, 1995). Local Siberian cattle were first imported to Alaska in the 1790s as part of an effort to establish an agricultural colony in Russian America, Alaska (Bancroft, 1886). Further, the desire for beef, milk and butter led to a very general importation of Siberian cattle from Petropaulovsk, Yakutia (Sakha Republic), to every post in Alaska (Elliott, 1887). In 1798, the Russians established an outpost on Chirikof Island (Long, 1975). Thus, it is plausible that cattle of Sakha origin were brought to isolated Chirikof Island in the Gulf of Alaska at this time. Cattle originating from the United States were reported to have first arrived on Chirikof Island in the mid-1880s (Long, 1975). In 1927, 400 beef cattle were tallied on Chirikof Island (USDA, 1929). According to reports in the popular press, enterprising cattle producers have sporadically added Hereford, Angus, Highland, Shorthorn and perhaps other breeds of cattle to the Chirikof Island population throughout the 1900s (Fields, 2000; D’Oro, 2003, 2005). Collectively, these events likely contribute to the complexity of the genetic structure of the cattle on Chirikof Island.
In 1980, the island became part of the Alaska Maritime Wildlife Refuge. In the 1980s, the US Fish and Wildlife Service began removing introduced species from various islands in the refuge, mainly foxes introduced by fur traders, but also cattle, although not from the isolated Chirikof Island at that time. The government granted grazing leases for Chirikof Island in the twentieth century. The last grazing lease for Chirikof Island expired in 2000 and a permit to remove the cattle expired in 2003. However, the cattle were not removed and currently remain on Chirikof Island. In late 2013, the US Fish and Wildlife Service restated its intent ‘to restore these islands and finally help them fulfill their congressionally mandated destiny as a wildlife refuge’ (Medeiros, 2013). Federal wildlife managers seek to remove the cattle from Chirikof Island to stop grazing and enhance habitat for birds (Press, 2013). It is widely presumed that this restoration will result in the extirpation of the feral Chirikof Island cattle. However, knowledge of the extent and nature of genetic diversity may aid in objective and rational decision making relative to potential conservation of this germplasm (FAO, 2004). Therefore, the objective of the present work was to more definitively quantify the origins, admixture and divergence of the Chirikof Island cattle relative to more readily accessible domestic cattle.
Materials and methods
We genotyped 10 Chirikof Island cattle using the Illumina (Illumina, Inc., San Diego, CA, USA) BovineSNP50 BeadChip (Matukumalli et al., 2009), and BovineSNP50 genotypes were also obtained for 40 Yakut and 22 Kalmyk cattle, both from Russian Siberia. DNA for genotyping was extracted by phenol−chloroform precipitation. These single-nucleotide polymorphism (SNP) data were integrated with the data generated from the worldwide sampling of domestic cattle conducted by Decker et al. (2014a, b). The data set contained 43,018 SNPs after removing 25 parentage SNPs that are duplicated on the BovineSNP50 BeadChip (see DATA DRYAD repository for the updated PLINK map file).
For the SNP data, autosomal SNPs and a single pseudo-autosomal X chromosome SNP were analyzed. SNP filtering was previously described (Decker et al., 2014b). Principal component analysis implemented in the smartpca program of EIGENSOFT (Patterson et al., 2006), ancestry graphs implemented in TreeMix (Pickrell and Pritchard, 2012) and ancestry models implemented in fastSTRUCTURE (Raj et al., 2014) were used to assess the relationship of Chirikof Island cattle to 136 breeds of domesticated bovids (137 breeds total). These breeds arose from three domesticated (sub)species: Bos javanicus, Bos taurus indicus and Bos taurus taurus. In smartpca, to account for the effects of linkage disequilibrium, for each SNP the residual of a regression on the previous two SNPs was input to the principal component analysis (see EIGENSOFT POPGEN README). FastSTRUCTURE, with values of K from 1 through 40, was used to evaluate inferred genomic components for K populations. Two metrics from fastSTRUCTURE were used to assess appropriate values of K for the population structure contained in the data set. The metric  is the value of K that maximizes the log marginal likelihood lower bound. The metric
is the value of K that maximizes the log marginal likelihood lower bound. The metric  is the minimum value of K that accounts for almost all of the ancestry in the data set.
is the minimum value of K that accounts for almost all of the ancestry in the data set.
We also used weighted linkage disequilibrium decay statistics as implemented in a version of ALDER v1.03 (Loh et al., 2013) modified to allow for >23 chromosomes, to estimate the number of generations since admixture and lower bounds of ancestry fractions from the reference populations. ALDER fits three linkage disequilibrium decay curves, with the first curve weighted by allele frequency divergences between the test population and the first reference population, the second curve weighted by allele frequency divergences between the test population and the second reference population, and the third curve weighted by allele frequency divergences between the two reference populations, and then compares estimates from these three curves for consistency. A linkage map based on 43 573 individuals from 632 parents, primarily defining half-sib families, and genotyped with the BovineSNP50 assay was constructed at the University of Missouri (unpublished) and used to define the genetic distances (cM) among the SNP loci.
We used TreeSelect (Bhatia et al., 2011) to identify selected loci for which the increased divergence occurred on the Chirikof Island branch. TreeSelect fits a three-population phylogeny and identifies loci where the divergence between the central node and the population at the tip is greater than expected due to random drift alone. Because TreeSelect works best on closely related populations (Bhatia et al., 2011), we used the Hereford and Yakut breeds as the two other populations in the TreeSelect analysis, based on their relationship to Chirikof Island cattle from other analyses.
Results and discussion
MacNeil et al. (2007) found the Chirikof Island cattle to be genetically variable using microsatellites and relatively unique when compared with the Angus, Charolais, Hereford, Highland, Limousin, Red Angus, Salers, Shorthorn, Simmental, Tarentaise and Texas Longhorn breeds present in the United States during the early 2000s. Since that time, Bos taurus cattle breeds from a wide range of geographical origins have been characterized using SNPs, and Decker et al. (2014a, b) assembled a genome-wide data set for 134 cattle breeds detailing the population structure of domesticated cattle worldwide. Here, we use these data along with additional BovineSNP50 genotypes of Yakut and Kalmyk cattle to accurately describe the ancestry and history of Chirikof Island cattle.
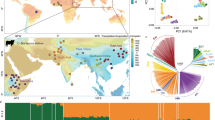
From the principal component analysis of the data, the first two principal components show the Chirikof Island cattle as clustering at the margin of European Bos taurus taurus, tending toward the Russian Yakut Bos taurus taurus cattle (Figure 1). The Chirikof cattle also cluster within the variation of American Criollo breeds, which have a history of admixture. The Chirikof Island cattle are positioned between the Hereford and Yakut cattle (Figure 1b), consistent with admixture between the two breeds (McVean, 2009).
(a) Principal component plot incorporating Chirikof Island cattle into the analysis of worldwide patterns of ancestry, divergence and admixture in domesticated cattle of Decker et al., (2014a, b). Samples from Asia are in green, Africa in black, Europe in blue, Americas in red, Australia in orange and Chirikof Island cattle in purple. (b) Expanded image of lower-left quadrant containing samples from Chirikof Island cattle. Chirikof Island cattle are in purple, Hereford in brown and Yakut in orange. (Samples in blue and red with principal component 1 values >−0.015 are Italian and American Criollo breeds).
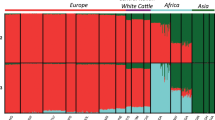
In the Bayesian clustering analysis of the SNP genotypes, the marginal likelihood was maximized at K=19 (Figure 2 and Supplementary Figure S1). These results suggest that Chirikof cattle range from ~49 to 59% ancestry, which is similar to Hereford, and from ~33 to 40% ancestry, which is similar to Yakut, with the remaining ancestry being a mixture of various breeds.
Bar plot showing the extent of admixture in Chirikof Island cattle at K=19 (marginal likelihood maximized) as derived from BovineSNP50 genotypes from 137 breeds. Only breeds that contributed to Chirikof Island cattle are plotted. For all 137 breeds see Supplementary Figure S1.
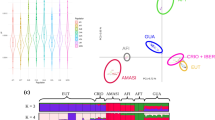
In TreeMix, allele frequency data are used to model genetic drift between populations using a Gaussian model. The residuals between the observed covariances between populations and the phylogenetic model covariances are compared. When residuals are large, populations are more closely related than depicted by the phylogeny. In these instances, TreeMix adds edges to the phylogeny (making it a network) to account for admixture between populations. These edges are added in an iterative manner until sufficient model fit is obtained. The phylogenetic network analysis agrees with the fastSTRUCTURE results, indicating that Chirikof cattle received 34.4% of their ancestry from a Yakut ancestor, with the remainder of their ancestry being most similar to Hereford (Figure 3).
(a) Phylogenetic network of the inferred relationships between 19 cattle breeds and the Chirikof Island cattle. The plotted network accounts for 98.3% of the covariance between populations. The color of admixture edges denotes the percent ancestry contributed by the donor to the admixed population; the edge from Yakut to Chirikof Island cattle accounts for 34.4% of Chirikof Island cattle ancestry. (b) Plot of the residuals between the phylogeny with no migration edges and the observed data. (c) Plot of the residuals between the phylogenetic network with two migration edges and the observed data.
Finally, we used weighted linkage disequilibrium decay curves to estimate the timing and amount of admixture in the Chirikof Island cattle using ALDER v1.03 (Loh et al., 2013). With Chirikof Island cattle as the test population, Hereford as reference population 1 and Yakut as reference population 2, we find significant evidence that Chirikof Island cattle are admixed from relatives of these reference populations (Z=11.06, P=1.9e−28). However, ALDER reported that the decay rates (number of generations since admixture) were not consistent in this analysis, but we infer that this result may be historically accurate. In a single-reference-population analysis using Hereford for which the minimum distance between SNPs was 2 cM in the University of Missouri linkage map (due to high background linkage disequilibrium), we estimated that the admixture occurred 8.19±1.58 (Z=5.17, P=1.2e−7) generations ago. The lower bound (not the point estimate) for the extent of Hereford ancestry in Chirikof Island cattle was estimated to be 21.0±1.4%. Conversely, with Yakut as the single reference and the minimum distance between SNPs at 2 cM, ALDER estimated that the admixture occurred 23.87±6.34 generations ago (Z=3.76, P=8.5e−5), and from this analysis Chirikof Island cattle have an estimated minimum of 9.1±1.5% Yakut-like cattle ancestry. These incongruent numbers of generations since admixture can easily be reconciled. Assuming a generation interval of 5 years (MacEachern et al., 2009; Decker et al., 2012), we can infer that Yakut-like cattle arrived first on Chirikof Island, and then ~120 years ago the first European taurine cattle were introduced to the island by US farmers, while finally Hereford cattle were introduced to the island about 40 years ago. These analyses provide a detailed description of the nature, proportion and timing of admixture of the Chirikof Island cattle.
The Chirikof Island cattle also may be an economically important resource if they harbor variants that have allowed them to adapt to the harsh environment of Chirikof Island. We performed an analysis seeking to identify selected loci, but our test using TreeSelect was underpowered (Supplementary Figures S2 and S3) due to the small sample size and granular estimates of allele frequencies. Nevertheless, we identified 12 SNPs in eight loci for which the change in allele frequency suggests that strong selection has acted upon these loci (Supplementary Table S1). For example, the most extremely diverged SNP from TreeSelect on chromosome 8 at 100.2 Mbp (Supplementary Table S1, P=0.0005) had an allele frequency of 0.413 in the central node of the phylogeny, but an allele frequency of 0.0125 in the Chirikof Island cattle. Genes in the near vicinity of these eight loci are involved in immune-defense response, embryonic development and cancer.
Chirikof Island is isolated in the western Gulf of Alaska, 61 km from the nearest land and 74 km southwest of the Kodiak archipelago (Withrow, 2015). Thus, the existence of cattle on Chirikof Island has undoubtedly resulted from immigration events mediated by people. Absence of a deepwater harbor results in immigration and emigration events being extremely difficult and thus most likely involving only small numbers of animals. The limited number of founders in these admixture events likely influenced the differentiation of Chirikof cattle (Mayr, 1954).
The results of our analyses indicate that the Chirikof Island cattle arise from at least three immigration events, and today they are a composite of British (mostly Hereford) and Siberian Yakut cattle. Despite historical records indicating the influence of other breeds of cattle, our results indicate that Chirikof cattle contain predominately Hereford and Yakut ancestry. This composite, while clearly Bos taurus taurus, appears to be at least as distinct as recognized breeds of cattle, and the mixture of British and Siberian cattle employed to form the Chirikof Island population is unique compared with other North American breeds. Further, the timing of these events from genetic data is in strong agreement with historical records. Natural selection in the relatively harsh environment of Chirikof Island has likely impacted their genetic architecture.
Data archiving
Data available from the Dryad Digital Repository: http://dx.doi.org/10.5061/dryad.cf03b.
References
Bancroft HH . (1886) History of Alaska, 1730–1885. Antiquarian Press Ltd.: New York, NY, USA.
Bhatia G, Patterson N, Pasaniuc B, Zaitlen N, Genovese G, Pollack S et al. (2011). Genome-wide comparison of African-ancestry populations from CARe and other cohorts reveals signals of natural selection. Am J Hum Genet 89: 368–381.
Decker JE, McKay SD, Rolf MM, Kim J, Molina Alcalá A, Sonstegard TS et al. (2014a). Data from: Worldwide Patterns of Ancestry, Divergence, and Admixture in Domesticated Cattle. Dryad Digit Repos; e-pub ahead of print 27 March 2014 doi:10.1371/journal.pgen.1004254.
Decker JE, McKay SD, Rolf MM, Kim J, Molina Alcalá A, Sonstegard TS et al. (2014b). Worldwide patterns of ancestry, divergence, and admixture in domesticated cattle. PLoS Genet 10: e1004254.
Decker JE, Vasco DA, McKay SD, McClure MC, Rolf MM, Kim J et al. (2012). A novel analytical method, birth date selection mapping, detects response of the Angus (Bos taurus) genome to selection on complex traits. BMC Genomics 13: 606.
d’Oro R . (2003) Alaskan Wild Cattle Herd Faces Expulsion. Associated Press. Available from http://espn.go.com/outdoors/conservation/news/2003/1120/1666710.html (accessed 8 August 2005).
d’Oro R . (2005) Chirikof Cattle Will Roam Alone as Dispute Simmers. Associated Press, Anchorage Daily News: Anchorage, AK, USA.
Elliott HW . (1887) Our Arctic Province: Alaska and the Seal Isands. C. Scribner’s Sons: New York, NY, USA.
FAO. (2004). Measurement of Domestic Animal Diversity. Proceedings of the Working Group on Genetic Resources for Food and Agriculture. (Commission on Genetic Resources for Food and Agriculture, 3rd Session); 31 March–2 April 2004 Rome, Italy.
Fields WM . (2000) Now It Can Be Told: Stories of Alaskan Pioneer Ranchers. Publication Consultants: Anchorage, AK, USA.
Kotkin S, Wolff D . (1995) Rediscovering Russia in Asia: Siberia and the Russian Far East. ME Sharpe: Armonk, NY, USA.
Loh P-RR, Lipson M, Patterson N, Moorjani P, Pickrell JK, Reich D et al. (2013). Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193: 1233–1254.
Long JS . (1975) McCord of Alaska: Statesman for the Last Frontier. Dillon/Liederbach: Cleveland, OH, USA.
MacEachern S, Hayes B, McEwan J, Goddard M . (2009). An examination of positive selection and changing effective population size in Angus and Holstein cattle populations (Bos taurus) using a high density SNP genotyping platform and the contribution of ancient polymorphism to genomic diversity in domestic cattle. BMC Genomics 10: 181.
MacNeil MD, Cronin MA, Blackburn HD, Richards CM, Lockwood DR, Alexander LJ . (2007). Genetic relationships between feral cattle from Chirikof Island, Alaska and other breeds. Anim Genet 38: 193–197.
Matukumalli LK, Lawley CT, Schnabel RD, Taylor JF, Allan MF, Heaton MP et al. (2009). Development and characterization of a high density SNP genotyping assay for cattle. PLoS One 4: e5350.
Mayr E (1954). Change of genetic environment and evolution. In: Huxley J, Hardy AC, Ford EB (eds). Evolution as a Process. Allen & Unwin: London, UK, pp 157–180.
McKnight TL . (1964) Feral Livestock in Anglo-America. University of California Press: Oakland, CA, USA.
McVean G . (2009). A genealogical interpretation of principal components analysis. PLoS Genet 5: e1000686.
Medeiros A . (2013) Wildlife Refuge Plans to Address Cattle Damage to Islands. US Fish and Wildlife Service: Anchorage, AK, USA.
Patterson NJ, Price AL, Reich D . (2006). Population structure and eigenanalysis. PLoS Genet 2: e190.
Pickrell JK, Pritchard JK . (2012). Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet 8: e1002967.
Press A . (2013) The Threat to Cattle Herds on Remote Alaskan Islands. Epoch Times: New York, NY, USA.
Raj A, Stephens M, Pritchard JK . (2014). fastSTRUCTURE: variational inference of population structure in large SNP datasets. Genetics 197: 573–589.
USDA. (1929) Survey of the Grazing Possibilities of the Aleutian Islands and Vicinity Report of the Alaska Agricultural Experiment Stations USDA: Washington, DC, USA.
van Vuren D, Hedrick PW . (1989). Genetic conservation in feral populations of livestock. Conserv Biol 3: 312–317.
Withrow JJ . (2015). Notes on the birds of Chirikof Island, Alaska. West Birds 46: 28–48.
Acknowledgements
JFT was supported by National Research Initiative competitive grant numbers 2011-68004-30214, 2011-68004-30367, 2013-68004-20364 and 2015-67015-23183 from the USDA National Institute of Food and Agriculture. JED was supported by Hatch Project MO-HAAS0027 and USDA NSRP-8 MO-MSAS0014 funds. Zoya Ivanova, Ruslan Popov, Innokentyi Ammosov and Mikhail Ozerov are acknowledged for collaboration in sampling of Yakut and Kalmyk breeds. JK was supported by the Academy of Finland grant number 286040.
Author contributions
MDM, JED and JFT conceived the study. JFT, JED and JK provided SNP genotypes. RDS provided the linkage map. AM conducted historical research on Russian settlements. JED and MDM analyzed the data. All authors contributed to interpretation of the results. JED, JFT and MDM drafted the manuscript with input from all authors.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Decker, J., Taylor, J., Kantanen, J. et al. Origins of cattle on Chirikof Island, Alaska, elucidated from genome-wide SNP genotypes. Heredity 116, 502–505 (2016). https://doi.org/10.1038/hdy.2016.7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2016.7
This article is cited by
-
Refining the genetic structure and relationships of European cattle breeds through meta-analysis of worldwide genomic SNP data, focusing on Italian cattle
Scientific Reports (2020)
-
Genome-wide assessment of genetic diversity and population structure insights into admixture and introgression in Chinese indigenous cattle
BMC Genetics (2018)
-
Whole-genome SNP analysis elucidates the genetic structure of Russian cattle and its relationship with Eurasian taurine breeds
Genetics Selection Evolution (2018)
-
Conservation status and historical relatedness of Italian cattle breeds
Genetics Selection Evolution (2018)
-
Species composition and environmental adaptation of indigenous Chinese cattle
Scientific Reports (2017)