Abstract
Eukaryotic protein sequences often contain amino-acid homopolymers that consist of a single amino acid repeated from several to dozens of times. Some of these are functional but others may persist largely because of high expansion rates due to DNA slippage. However, very long homopolymers with over a hundred repeats are very rare. We report an extraordinarily long homopolymer consisting of 306 tandem serine repeats from the single-celled eukaryote Dictyostelium discoideum, which also has a multicellular stage. The gene has a paralog with 132 repeats and orthologs, also with high serine repeat numbers, in various other Dictyostelid species. The conserved gene structure and protein sequences suggest that the homopolymer is functional. The high codon diversity and very poor alignment of serine codons in this gene between species similarly indicate functionality. This is because the serine homopolymer is conserved despite much DNA sequence change. A survey of other very long amino-acid homopolymers in eukaryotes shows that high codon diversity is the rule, suggesting that these too may be functional.
Similar content being viewed by others
Introduction
Amino-acid homopolymers are common and abundant in eukaryotic proteins (Haerty and Golding, 2010; Mularoni et al., 2010). Some of these repeat sequences are functional (Fondon and Garner, 2004; Huntley and Golding, 2006; Fondon et al., 2008; Haerty and Golding, 2010). Some exhibit dysfunctional effects in individuals that have expanded homopolymers, as in several human neurological diseases (Orr and Zoghbi, 2007) and developmental disorders (Utsch et al., 2002). However, there is considerable debate over whether such sequences are generally functional (Ellegren, 2004; Luo et al., 2012). Because of the high expansion rate due to DNA slippage, many homopolymers may persist despite an absence of selection for them (Schlötterer and Tautz, 1992).
Exceptionally long natural homopolymers are rare but might provide insight into the functionality question. We report on one of the longest known homopolymers in any eukaryote, 306 tandem iterations of the hydrophilic amino acid serine (poly-S306) in the social amoeba Dictyostelium discoideum, a soil-living eukaryote able to switch from solitary single cell growth to multicellular development upon starvation (Raper, 1984). Two factors suggest that the repeat is functional. First, it has homologs, although with shorter repeats, throughout the ancient Dictyostelids (Schaap et al., 2006). Second, these serine repeats have very high codon diversity, something that is also true of other long amino-acid homopolymers in eukaryotes. The high diversity of serine codons has previously been used to infer selection on serine repeat genes (Huntley and Golding, 2006).
The genome of the social amoeba D. discoideum is extremely rich in amino-acid homopolymers. These occur in 34% of the predicted proteins and make up 3.3% of all amino acids of the whole proteome (Eichinger et al., 2005). The most abundant are homopolymers of asparagine and glutamine, with 2091 gene models containing homopolymers with over 20 of one of these amino acids (Eichinger et al., 2005), but threonine and serine repeats are also common. A multispecies comparison suggests that this high abundance is largely attributable to its unusually high genomic AT content, which allows more simple triplet DNA repeats to be initiated by chance, providing more raw material for the normal repeat expansion process via replication slippage (Tian et al., 2011). Selection on the proteins bearing these repeats may not be strong because they tend to be coded by genes with low expression and high rates of substitutional change (Sucgang et al., 2011). An unusually large fraction of these proteins lack homologs in D. purpureum and when they do have homologs, the existence, type and location of repeats usually differs (Sucgang et al., 2011). Moreover, a set of 50 genes with long asparagine homopolymers had just as much variation in repeat number among individuals as a comparable set of noncoding triplet repeats. This is consistent with the protein-coding repeats being under no greater selection than the noncoding ones (Scala et al., 2012).
In this study, we focus on an extraordinarily long polyserine repeat, provide some evidence that it is functional and suggest that this may often be true of extremely long amino-acid homopolymers.
Materials and methods
Genome and transcriptome data
Expression sequence tags (ESTs), predicted gene models and proteins of D. discoideum (Eichinger et al., 2005) and D. purpureum (Sucgang et al., 2011) were retrieved from dictyBase (http://dictybase.org/). The predicted gene models and proteins of Polysphondylium pallidum and D. fasciculatum (Heidel et al., 2011) were retrieved from the database http://sacgb.fli-leibniz.de. The gene models of D. intermedium and P. violaceum were predicted using the program GENEID (Guigo et al., 1992; Blanco et al., 2007) and MAKER (Cantarel et al., 2008) based on the assemblies of 454-sequencing reads with 63 × and 19 × coverage of the genome, respectively. These species range across groups 1, 2 and 4 of Schaap’s categorization of social amoebae (Schaap et al., 2006). The RNA-seq transcriptome data of D. discoideum and D. purpureum were provided by Parikh et al. (2010).
Homopolymer identification
Amino-acid homopolymers with ⩾20 perfect tandem iterations were identified from the predicted proteins using the in-house Perl scripts.
Ortholog identification
Orthologs of genes with large serine repeats were identified using the program Inparanoid version 3.0 (Remm et al., 2001), a method based on the best reciprocal blast hits of protein sequences. A minimum cutoff for similarity of 50 bits and 50% overlap was used.
Codon diversity calculation
We calculated the codon diversity (D) of coding sequences of each homopolymer on the basis of the genetic diversity proposed by Hedrick (2005):

Here pi is the frequency of the ith codon and k is the overall number of codon types used by the homopolymer. This is the probability that two randomly drawn codons are different.
Database search of amino-acid homopolymers
To identify other long amino-acid homopolymers, we searched (using the blastp program) using 200-mer amino-acid repeats as queries against NCBI’s nr database (04-15-2011). We manually checked the matched protein sequences and screened out those with truncated coding sequences and homopolymers of fewer than 100 repeats. To assess the expression of genes with homopolymers we searched, using BLAST (megablast), the ESTs from NCBI’s EST database (04-15-2011) that match our gene sequences with homopolymers.
Gene ontology
To test whether the 80 serine proteins with serine repeats of at least 20 residues was overrepresented in gene ontology (GO) categories, we use a hypergeometric test in the program BiNGO (Maere et al., 2005) a plugin of the platform Cytoscape (Shannon et al., 2003).
Phosphorylation site prediction
We predicted phosphorylations site on the protein sequence using an artificial neural network method-based program NetPhos (Blom et al., 1999).
Results and discussion
The homopolymer poly-S306 was identified from a protein model, which we named LSR1 (long serine repeat), of D. discoideum. The 306 serine repeats occupy 82% of the entire protein sequence with only 65 non-serine amino acids at the N-terminus and 3 at the C-terminus. The repeat is exceptional not just in length, but also in several other characteristics.
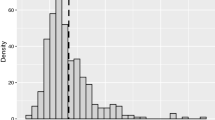
Except for a 70-tandem repeat of the codon TCA in the middle, this polyserine is encoded by a complex mixture of five synonymous codons rather than iteration of a single codon (Figure 1a). We measured its codon diversity (D), which is the probability that two randomly chosen codons are different. The D=0.58 of poly-S306 is in the top 9 among overall of 3391 amino-acid homopolymers with over 20 repeats in D. discoideum. Serine homopolymers generally have higher codon diversity than other amino-acid repeats in D. discoideum (Eichinger et al., 2005), but the poly-S306 ranks in the top 5 of 81 even among serine homopolymers of >20 repeats (Figure 2). The very high codon diversity is consistent with selection for point mutations that break up perfect repeats and reduce excessive replication slippage (Haerty and Golding, 2010). At the same time it is clear that, of all the point mutations that would reduce slippage, only serine coding ones have been favored; preservation of serines seems to be crucial.
Structure and expression pattern of genes containing the extraordinarily long amino-acid homopolymer. (a) Gene structure of the D. discoideum LSR1 (gene ID: DDB_G0271670) that includes 306 serine tandem repeats. The gray lines show the sequences of DNA, RNA and protein. The black bars mark the coding regions of DNA sequence, mRNA of the RNA sequence and tandem repeats of amino-acid serine. The feature of trinucleotide repeats in DNA sequence and ESTs coving RNA sequence are marked by thin gray bars. (b) Codon usage frequency. The x-axis marks the codon numbers. The y-axis marks the polyserines from Dictyostelids (DI=D. intermedium, DD=D. discoideum, PP=P. pallidum, PV=P. violaceum, DP=D. purpureum, DF=D. fasciculatum) and the codon diversity (numbers in parentheses). Bars with different fill effects show the frequency of each codon. (c) RNA-seq transcriptional profile. x-Axis marks the development stages (8–24 h) and the solitary stage (0 h). The y-axis marks the log RNA-seq read counts.
Distribution of codon diversities of the long homopolymers (⩾20 repeats) from D. discoideum. The D values (y-axis) are plotted against the length of homopolymers (x-axis) with a different color and symbol for each amino acid (n=3391). The two unusually long (>100 repeats) homopolymers poly-S306 and poly-S132 are identified.
The homopolymer poly-S306 has a paralog in D. discoideum (gene LSR2). Although its serine homopolymer is far shorter (132 serines), it is still the second longest homopolymer in the genome. It also has exceptional codon diversity, ranking third among all D. discoideum homopolymers longer than 20 amino acids, and second among the serine homopolymers (Figure 2).
These two genes have interspecific orthologs, with 51–196 iterations of serines, identified from five other species of Dictyostelids (D. intermedium, D. purpureum, D. fasciculatum, P. pallidum and P. violaceum). Unlike most homopolymers in D. discoideum (Sucgang et al., 2011), there is an ortholog in D. purpureum with serine repeats (196) in the same position of the protein. The proteomes of these two species are generally as divergent as those of mammals and bony fish (Sucgang et al., 2011), so this is a long-term conservation of the repeat sequence in this location. Moreover, apparent homologs also exist in even more distant Dictyostelids, P. pallidum and D. fasciculatum, although in the former the region after the serine repeat has greatly expanded and has homology to Ras guanine nucleotide exchange factors. The genes from other species have similar gene structure as LSR1 and LSR2 in D. discoideum (Supplementary Figure S1). The flanking amino-acid residues of the polyserines share high sequence similarities among species, especially at the longer N-terminus of the protein sequence (Supplementary Figure S2).
However, codons of the polyserines are nearly completely diverged between species (Supplementary Figure S3). As with poly-S306 in D. discoideum, the polyserines of its homologs are a diverse combination of the serine codons, with high codon diversities, D ranging from 0.41 to 0.79, among which ‘TCA’ is the most common codon (Figure 1b). The conservation of the serine repeats in the absence of much underlying DNA conservation strongly suggests that selection is maintaining the serines.
Despite conservation of gene structure and the serine repeat, the function of this gene is unclear. The ESTs from full-length cDNAs (Morio et al., 1998; Urushihara et al., 2004; Sucgang et al., 2011) indicate that the genes producing the exceptionally long polyserines in D. discoideum (LSR1 and LSR2 gene encoding poly-S306 and poly-S132, respectively) and D. purpureum (LSR gene encoding poly-S196) are transcribed (Figure 1a and Supplementary Figure S2). The RNA-seq transcriptional profiles show that the latter two are strongly upregulated in the multicellular stages in contrast to the solitary stage (Parikh et al., 2010), but that LSR1 with 306 serine repeats is expressed more consistently throughout the life cycle (Figure 1c). It may be that selection for the very long repeats in D. discoideum LSR1 is related to this shift in timing, with perhaps also a shift in function. The NetPhos program predicted a very high phosphorylation potential for many of the serine residues in the long repeat regions (Supplementary Figure S4). Finally, the 80 D. discoideum genes with 20 or more tandem serines did not show overrepresentation in any gene ontology categories. Although our bioinformatics/evolutionary approach indicates functionality, what the function is will need to be discovered through more mechanistic studies.
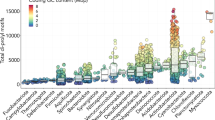
That this gene is special is also supported by a database search, which found only 35 homopolymers with over 100 iterations of a single amino acid in 17 eukaryotic organisms apart from those found in Dictyostelids (Supplementary Table S1). Most of them are from predicted proteins without functional annotation and only eight of them have data showing EST coverage of coding sequences. Strikingly, most of them also have high codon diversities. These are not usually as high as the Dictyostelium long serine repeats, but serine has six codons and therefore a maximum D of 0.83, whereas the maximum for four codons is D=0.75 and for two codons is D=0.5. Eighteen of the 31 Ds in Supplementary Table S1 are for two-codon amino acids (D, Q, Y, K), and many of these approach the maximum diversity possible. There is a possible bias in that extremely long perfect repeats might be missed in genome projects because of alignment difficulties. However, our results indicate that the high codon diversity of very long repeats is real and may indicate that these repeats may more often be functional than shorter, less diverse homopolymers. Even if their initial expansion was through slippage of perfect triplet repeats, their high codon diversity suggests that they have endured through millions of years. Functionality of the sequence would explain why, during that long time, the repeat sequence has not been lost via deletion or modified to include other amino acids.
References
Blanco E, Parra G, Guigo R . (2007). Using geneid to identify genes. Curr Protoc Bioinformatics Chapter 4: Unit 4 3.
Blom N, Gammeltoft S, Brunak S . (1999). Sequence and structure-based prediction of eukaryotic protein phosphorylation sites. J Mol Biol 294: 1351–1362.
Cantarel BL, Korf I, Robb SM, Parra G, Ross E, Moore B et al. (2008). Maker: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res 18: 188–196.
Eichinger L, Pachebat JA, Glockner G, Rajandream MA, Sucgang R, Berriman M et al. (2005). The genome of the social amoeba Dictyostelium discoideum. Nature 435: 43–57.
Ellegren H . (2004). Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5: 435–445.
Fondon JW 3rd, Garner HR . (2004). Molecular origins of rapid and continuous morphological evolution. Proc Natl Acad Sci USA 101: 18058–18063.
Fondon JW 3rd, Hammock EA, Hannan AJ, King DG . (2008). Simple sequence repeats: genetic modulators of brain function and behavior. Trends Neurosci 31: 328–334.
Guigo R, Knudsen S, Drake N, Smith T . (1992). Prediction of gene structure. J Mol Biol 226: 141–157.
Haerty W, Golding GB . (2010). Genome-wide evidence for selection acting on single amino acid repeats. Genome Res 20: 755–760.
Hedrick PW . (2005) Genetics of Populations 3rd edn Jones and Bartlett Publishers: Boston.
Heidel AJ, Lawal HM, Felder M, Schilde C, Helps NR, Tunggal B et al. (2011). Phylogeny-wide analysis of social amoeba genomes highlights ancient origins for complex intercellular communication. Genome Res 21: 1882–1891.
Huntley MA, Golding GB . (2006). Selection and slippage creating serine homopolymers. Mol Biol Evol 23: 2017–2025.
Luo H, Lin K, David A, Nijveen H, Leunissen JA . (2012). Prorepeat: an integrated repository for studying amino acid tandem repeats in proteins. Nucleic Acids Res 40: D394–D399.
Maere S, Heymans K, Kuiper M . (2005). Bingo: a cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21: 3448–3449.
Morio T, Urushihara H, Saito T, Ugawa Y, Mizuno H, Yoshida M et al. (1998). The Dictyostelium developmental cDNA project: generation and analysis of expressed sequence tags from the first-finger stage of development. DNA Res 5: 335–340.
Mularoni L, Ledda A, Toll-Riera M, Alba MM . (2010). Natural selection drives the accumulation of amino acid tandem repeats in human proteins. Genome Res 20: 745–754.
Orr HT, Zoghbi HY . (2007). Trinucleotide repeat disorders. Annu Rev Neurosci 30: 575–621.
Parikh A, Miranda ER, Katoh-Kurasawa M, Fuller D, Rot G, Zagar L et al. (2010). Conserved developmental transcriptomes in evolutionarily divergent species. Genome Biol 11: R35.
Raper KB . (1984) The Dictyostelids. Princeton Universtiy Press: Princeton.
Remm M, Storm CE, Sonnhammer EL . (2001). Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J Mol Biol 314: 1041–1052.
Scala C, Tian X, Mehdiabadi NJ, Smith MH, Saxer G, Stephens K et al. (2012). Amino acid repeats cause extraordinary coding sequence variation in the social amoeba Dictyostelium discoideum. PLoS One 7: e46150.
Schaap P, Winckler T, Nelson M, Alvarez-Curto E, Elgie B, Hagiwara H et al. (2006). Molecular phylogeny and evolution of morphology in the social amoebas. Science 314: 661–663.
Schlötterer C, Tautz D . (1992). Slippage synthesis of simple sequence DNA. Nucleic Acids Res 20: 211–215.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. (2003). Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498–2504.
Sucgang R, Kuo A, Tian X, Salerno W, Parikh A, Feasley CL et al. (2011). Comparative genomics of the social amoebae Dictyostelium discoideum and Dictyostelium purpureum. Genome Biol 12: R20.
Tian X, Strassmann JE, Queller DC . (2011). Genome nucleotide composition shapes variation in simple sequence repeats. Mol Biol Evol 28: 899–909.
Urushihara H, Morio T, Saito T, Kohara Y, Koriki E, Ochiai H et al. (2004). Analyses of cDNAs from growth and slug stages of Dictyostelium discoideum. Nucleic Acids Res 32: 1647–1653.
Utsch B, Becker K, Brock D, Lentze MJ, Bidlingmaier F, Ludwig M . (2002). A novel stable polyalanine [poly(a)] expansion in the hoxa13 gene associated with hand-foot-genital syndrome: proper function of poly(a)-harbouring transcription factors depends on a critical repeat length? Hum Genet 110: 488–494.
Acknowledgements
We thank Gad Shaulsky for data access. This material is based upon work supported by the National Science Foundation under grant no. DEB-1201671.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Rights and permissions
About this article
Cite this article
Tian, X., Strassmann, J. & Queller, D. A conserved extraordinarily long serine homopolymer in Dictyostelid amoebae. Heredity 112, 215–218 (2014). https://doi.org/10.1038/hdy.2013.96
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2013.96