Abstract
Striga hermonthica is an angiosperm parasite that causes substantial damage to a wide variety of cereal crop species, and to the livelihoods of subsistence farmers in sub-Saharan Africa. The broad host range of this parasite makes it a fascinating model for the study of host–parasite interactions, and suggests that effective long-term control strategies for the parasite will require an understanding of the potential for host range adaptation in parasite populations. We used a controlled experiment to test the extent to which the success or failure of S. hermonthica parasites to develop on a particular host cultivar (host resistance/compatibility) depends upon the identity of interacting host genotypes and parasite populations. We also tested the hypothesis that there is a genetic component to host range within individual S. hermonthica populations, using three rice cultivars with known, contrasting abilities to resist infection. The developmental success of S. hermonthica parasites growing on different rice-host cultivars (genotypes) depended significantly on a parasite population by host–genotype interaction. Genetic analysis using amplified fragment length polymorphism (AFLP) markers revealed that a small subset of AFLP markers showed ‘outlier’ genetic differentiation among sub-populations of S. hermonthica attached to different host cultivars. We suggest that, this indicates a genetic component to host range within populations of S. hermonthica, and that a detailed understanding of the genomic loci involved will be crucial in understanding host–parasite specificity and in breeding crop cultivars with broad spectrum resistance to S. hermonthica.
Similar content being viewed by others
Introduction
The extent to which populations of parasites are adapted to populations of their host species is thought to depend on the evolutionary potential (population size and generation time) of parasite and host populations and their respective among-population migration rates (Kaltz and Shykoff, 1998; Gandon, 2002). Higher evolutionary potential or migration rate in the parasite species (relative to the host) should lead to adaptation to their local sympatric hosts. This can occur because alleles that confer adaptation of parasite individuals to common host genotypes can increase in frequency rapidly, or spread between populations.
In many natural systems, the coevolutionary process may take place, potentially unhindered, in both host and parasite species. Some agricultural systems provide a different perspective, where artificial constraints are imposed on the host species’ evolution or migration. For example, annual plant species within the genus Striga are obligate root parasitic weeds whose hosts include major cereal crop plants of sub-Saharan Africa, including maize, sorghum, pearl and finger millet and upland rice. They are considered to be one of the major biological constraints to cereal production in sub-Saharan Africa causing yield losses that range from 40 to 100% (Ejeta, 2007). The evolutionary and demographic dynamics of the local host species (cereal crop) are largely controlled by man. On the other hand, the parasite population is relatively free to evolve. Striga species maintain large population sizes, and extensive and persistent seed banks, which can remain viable in the soil for up to 20 years (Parker and Riches, 1993). They are also extremely fecund producing between 50 000–200 000 seeds per flower spike, and are sometimes moved as seed from field to field on farm equipment or via contaminated crop seed sold at local markets (Berner et al., 1994). Given these characteristics, we would anticipate that Striga species would be adapted to their local crop hosts. Alternatively, the seed bank may provide some ‘adaptive memory’ of previous rotations of particular crop species or genotypes (cultivars). Such a reservoir of genetic diversity has important implications for the use of resistant crop cultivars as a strategy for controlling Striga as it makes it likely that Striga populations will be able to evolve novel virulence phenotypes (host specificities) that enable them to overcome the challenges imposed by novel host-resistance phenotypes.
The use of resistant crop cultivars is considered to be one of the most cost-effective strategies for control of Striga, however, their effective deployment is limited due to a lack of understanding of the underlying genetic and phenotypic basis of the adaptation of Striga populations to new host resistance phenotypes (Scholes and Press, 2008). Such knowledge would allow the responses of Striga parasites to selection imposed by resistant host crops to be anticipated, allowing host resistance phenotypes to be combined and deployed in an optimal manner in different agro-ecosystems. Breeding system is a further crucial factor in understanding the genetic and phenotypic basis of virulence in the genus Striga, which includes obligate outbreeding species such as Striga hermonthica (Safa et al., 1984) as well as naturally inbreeding species such as S. asiatica (Mohamed et al., 2001). Breeding systems are strongly linked to locally effective population sizes. Naturally inbreeding parasite species, whose populations have low effective sizes are expected to show a greater degree of local adaptation to their sympatric hosts than are outbreeding species that possess greater effective population sizes (Gandon and Nuismer, 2009). Locally effective population sizes will also influence the retention of variability for host range and virulence within individual populations.
Most studies of the molecular genetics of host–parasite specificity within the Striga–cereal host–parasite system have been carried out on S. gesnerioides, which is a naturally inbreeding species and infects a range of dicotyledenous hosts such as cowpea (for a review see Li et al., 2009). In this host–parasite interaction, there is clear evidence of host species–parasite population specificity. Recently, the first resistance gene conferring resistance to S. gesnerioides has been identified and cloned from cowpea and shown to exhibit a gene for gene interaction with a specific race (SG3) of the parasite (Li et al., 2009). The situation is much less clear for the outbreeding species S. hermonthica, in which greater genetic variation is likely to be maintained within populations from generation to generation (Bharatha et al., 1990). In many cases, S. hermonthica seeds collected from one cereal host can infect other cereal species, although there is evidence of some interspecies specificity, particularly with respect to the reciprocal infectivity of populations of S. hermonthica collected from sorghum and pearl millet (Vasudeva Rao and Musselman, 1987). Very few studies have investigated the existence of intraspecies variation in host range or specificity within populations of S. hermonthica, but different host species (and genotypes of a particular host species) show different levels of resistance to particular populations or ecotypes of S. hermonthica (Gurney et al., 2006; JD Scholes, personal communication).
Our aim in this study was to gain a more detailed understanding of the extent to which populations of the parasitic witchweed, S. hermonthica, can exploit a range of crop host genotypes, and to investigate the genetic component to host range within an individual population. We used a controlled experiment to investigate host range within and among three populations of parasitic witchweed (S. hermonthica) using three rice cultivars with known, contrasting abilities to resist infection as hosts (Gurney et al., 2006). We also used amplified fragment length polymorphism (AFLP) markers and a genome scan approach to test the hypothesis that the genetic diversity within a S. hermonthica population forms part of the ability of parasite populations to colonise different host genotypes and will contribute to the evolution of virulence or compatibility on new hosts. We show that, parasite success depends upon an interaction between rice-host cultivar (genotype) and parasite population. Our molecular analysis indicates that genetic diversity within individual populations of witchweed contributes to the range of host genotypes they can exploit.
Materials and methods
Plant material
Oryza sativa cultivars Nipponbare (subspecies japonica) and Kasalath (subspecies indica) were obtained from the Rice Genome Resource Centre, Tsukuba, Japan. IAC 165 (subspecies japonica) was provided by the International Rice Research Institute, Los Baños, Laguna, Philippines. In the experiments presented here, these rice cultivars acted as hosts for ecotypes (populations) of S. hermonthica parasites. The cultivars were chosen because they have previously shown wide variation in susceptibility to S. hermonthica. In addition, two of the cultivars, Nipponbare and Kasalath are parents of a Backcross Inbred Line population that has been used to identify quantitative trait loci underlying post-attachment resistance to S. hermonthica (Gurney et al., 2006; Swarbrick et al., 2009). The S. hermonthica seed populations used in this study were chosen as they were collected from different host species and from different regions of Africa thus maximising the potential for genetic variation between populations. They were collected from S. hermonthica plants parasitizing (a) maize (H5 hybrid) in Kibos, Kisumu, Western Kenya (seed collected in 1997), (b) rice in northern Ivory Coast (1997) and (c) millet in Gambia (1993). The S. hermonthica population collected from Kibos Kenya was selected for detailed analysis of genetic structure and diversity as it has been used to identify quantitative trait loci for post-attachment resistance in rice (for example, Gurney et al., 2006).
Inoculation, growth and maintenance of rice
S. hermonthica seeds were surface sterilised and preconditioned for 12 days exactly as described in Gurney et al., (2002a, 2002b). The rice cultivars were established in root observation chambers (rhizotrons) as described by Gurney et al. (2006) and maintained in a controlled environment growth room with an irradiance of 500 μmol m−2 s−1 at plant height, a 12 h photoperiod and a relative humidity of 60%. Rice plants were watered four times a day with a total of 200 ml of 40% Long Ashton nutrient solution, containing 1 mM ammonium nitrate (Hewitt, 1966). Fourteen days after sowing, rice plants were inoculated with 25 mg of preconditioned, pregerminated S. hermonthica seeds to ensure synchronous attachment to the host plant and thus to quantify resistance responses in the host (Gurney et al., 2006). Five replicate plants were established for each combination of rice cultivar and parasite population (45 rhizotrons in total).
Parasite development and host resistance
Twenty-one days after inoculation of the roots, the number of parasites attached to the roots of each host plant was scored using a binocular microscope (Prior Scientific Inc, Rockland, MA, USA). Each Striga plant attached to a rice-host root was classified by developmental stage as follows: S1, the parasite radicle had attached to the host root but leaf primordia had not emerged from the seed coat; S2, leaf primordia had emerged from the seed coat; S3, the parasite shoot had between one and three leaf pairs; S4, the parasite had four or more leaf pairs; S5, parasites that had been attached to the root and died (visibly necrotic by day 21). Host resistance was quantified as the number of parasites that attached to the roots but failed to develop by day 21 (sum of S1 and S5 developmental categories) and was expressed as a percentage of the total number of attached parasites (sum of all developmental categories).
Statistical analysis of parasite development and host resistance
Differences in rice-host resistance among populations of Striga and rice-host cultivars were assessed using quasibinomial generalised linear model with logit link function (the data were initially modelled using a binomial (logistic) generalised linear model, but the data were found to be overdispersed). Resistance was modelled via counts of post-attachment developmental failure and success, rather than the corresponding percentage resistance score. Our approach modelled count data rather than % or proportion data to enable the logistic model to take into account differences in sample size (numbers of seedlings counted) among rhizotron replicates (by weighting the data), and to enable appropriate modelling of the variance of mean resistance score, which is expected to vary across its potential range (0–100%). Parasite population, rice-host cultivar and their interaction were considered as explanatory variables. Thus, the fitted generalised linear model can be summarised as R∼Host+Parasite+Host:Parasite, where ‘R’ is the rice-host resistance score in the form of counts of attachment failures and successes of Striga seedlings (response variable), ‘Host’ is the rice-host cultivar (categorical predictor) and ‘Parasite’ is the S. hermonthica parasite population (categorical predictor). The colon indicates the inclusion of an interaction term. The significance of each variable was tested by comparing models either containing or not containing the variable of interest via a likelihood ratio test, with reference to the F-distribution. Non-significant variables were dropped from the model to yield the minimum adequate model.
Analysis of the genetic structure of the Kibos (Kenyan) population of S. hermonthica
S. hermonthica populations collected from different cereal host species were each able to parasitise our rice-host test cultivars. Our second goal in this study was to test the hypothesis that there is a genetic component to host range within individual populations of S. hermonthica. To test this hypothesis, we focussed our investigation on the S. hermonthica parasite population from Kibos (Kenya).
DNA extraction
Tissue from Striga plants (originating from the Kenyan population) was harvested from individuals (sub-populations) growing on IAC 165, Kasalath and Nipponbare rice hosts, between 25 and 40 days after inoculation with the parasite. Striga seedlings were selected at random within a subset of plants, except for the Nipponbare sub-population, where many host plants were grown to collect the 24 parasite plants that were sampled. We sampled and genotyped 99 individuals of S. hermonthica (Kibos, Kenya); 40 from the sub-rice-host cultivar IAC 165, 35 from Kasalath and 24 from Nipponbare. Approximately, 100 mg of leaf tissue from each parasite was placed into a grinding tube and stored frozen at −80 °C. To estimate the rate of genotyping error, a replicate set of leaf tissue for eight individual parasites selected at random from the entire set under study was collected. Total genomic DNA was isolated from the frozen leaf tissue using the method described by Whitlock et al. (2008a) with minor modifications. Frozen samples were homogenized with a 5 mm tungsten carbide/steel ball bearing inside a 2 ml grinding tube using a 8000 M Mixer/Mill (Glen Creston Ltd, Stanmore, UK). The chaotropic salt guanidine thiocyanate (5.5 M) was used to bind the genomic DNA inside the wells of a silica-containing microtitre filter plate (1.0 μm plus 0.65 μm glass fibre/hydrophilic membrane, Millipore, Billerica, MA, USA).
AFLP analysis
AFLP analysis followed the protocol of Whitlock et al. (2008a). In brief, genomic DNA was digested with the restriction endonucleases EcoRI (New England Biolab (UK), Hitchin, UK) and MseI (Roche Diagnostics, Burgess Hill, UK). Preselective PCR amplification reactions were carried out in a total volume of 10 μl using two primer pair combinations: E_A–M_C and E_T–M_C (Table 1). Six primer pair combinations were used for the selective PCR (Table 1). PCR products were separated and detected using an ABI3730 DNA sequencer (Applied Biosystems, Foster City, CA, USA) and sized using an internal ROX size standard (Applied Biosystems). The software GENEMAPPER V3.7 (Applied Biosystems) was used to interpret the resulting chromatograms. First, sets of fragment length classes (bins) of width 1 base pair were created for each primer combination in the interval 50–500 base pairs. Next GENEMAPPER was used to derive a table of peak heights describing the fluorescence maximum of any AFLP PCR product (peak) present in any of the predefined bins. Finally, the software AFLPSCORE version 1.4b (Whitlock et al., 2008b) was used to estimate genotyping error rate and to convert the peak-heights table into a table of presence–absence AFLP phenotypes. Error rates were measured as a simple percentage mismatch value and as allelic error rates (Hadfield et al., 2006; Whitlock et al., 2008b). In total (across all six AFLP primer combinations), 191 AFLP loci were scored (of which 183 were polymorphic in the study as a whole) for each individual of S. hermonthica. The mismatch error rate, weighted to take account of the differing numbers of loci arising from different primer combinations, was 5.02% (Table 2). A breakdown of the scoring parameters used, the error rates and the final number of loci retained for each selective primer combination is given in Table 2.
Measurement of genetic structure and diversity
Allele frequencies for each AFLP locus were calculated using the Bayesian estimator of Zhivotovsky (1999), assuming Hardy–Weinberg equilibrium (data from allozymes indicates that this is a reasonable assumption for S. hermonthica; Bharatha et al., 1990). We used previous distributions for the allele frequencies that were estimated from the data. Basic diversity indices, including gene diversity, were calculated using AFLPSURV (Lynch and Milligan, 1994; Vekemans et al., 2002). FST was calculated across the three sub-populations of S. hermonthica growing on different rice-host cultivars using Cockerham and Weir's β statistic (Cockerham and Weir, 1993). A permutation test was conducted to determine whether the observed overall level of population differentiation (FST) was significantly different from zero. This test involved 5000 random permutations of individuals among sub-populations to obtain a null distribution for FST. Genetic differentiation (pair-wise FST) was also calculated for all three possible pairs of sub-populations.
Outlier analyses
Two outlier analyses (Beaumont and Nichols, 1996; Luikart et al., 2003; Foll and Gaggiotti, 2008) were carried out to identify AFLP loci marking regions of the S. hermonthica genome associated with parasite host range. The first outlier analysis is described below, and was tailored to our experimental design. It aimed to identify candidate AFLP loci that exhibited an outlier pattern of genetic differentiation (that is, those under differential selection) among parasite sub-populations growing on different rice-host cultivars, while controlling for the effects of sampling and genotyping error. The second outlier analysis, for comparison with the first, was carried out using the software BAYESCAN (Foll and Gaggiotti, 2008). The reason we carried out the first outlier analysis was because (1) the genotyping error rate in our AFLP data was relatively high (mismatch rate=5.02%) and we wanted to understand the effect of this genotyping error on our results, and (2) the demographic history of our study was very simple; the sub-populations for which we collected genetic data are derived from the base (seed) population after only one generation of selection on rice cultivar hosts. We carried out the second outlier analysis (BAYESCAN; Foll and Gaggiotti, 2008) because it allows for the possibility that multiple lineages of S. hermonthica coexist in partial reproductive isolation within single spatial field ‘populations’. If these lineages were differentially selected on different rice-host cultivars in our experiment, then there could be background genetic differentiation among sub-populations throughout the parasite genome. To identify loci responding to selection correctly, BAYESCAN models this background differentiation. The second outlier analysis is therefore more conservative than the first, in modelling any possible background differentiation and estimating the probability that individual loci are under selection by host genotype given this background. In the BAYESCAN analysis, we took those loci with log10 (Bayes factor) >0.5 to show evidence of differential selection among rice-host cultivars (corresponding to ‘substantial’ evidence for selection on the Jeffrey's scale of evidence for Bayes factors).
The outlier analysis that we designed for our data assumed that (1) the rice cultivars exerted no selection with respect to the genotypes of S. hermonthica growing on them; (2) the Striga seeds represented a well-mixed random sample of seed from a single, very large and randomly mating population. These assumptions constituted a null hypothesis under which all 99 Striga individuals belong to a single population with a common set of allele frequencies for the 191 AFLP loci investigated in this study. We assumed, in addition, that (3) mutation had no effect on allele frequencies over the course of the experiment and that (4) there was no migration among sub-populations. Both of these latter assumptions are realistic given that the experiment took place within a single generation of S. hermonthica. Under the null hypothesis, and the assumptions above, any differences observed in the allele frequencies of Striga sub-populations growing on different rice cultivars arose from either the effects of sampling from the base population or from genotyping error. These two sources of variability (particularly sampling) could cause inflation in genetic differentiation (FST) among the three sub-populations. The alternative hypothesis was that differential selection via rice-host cultivar exerted an additional effect on allele frequencies, for individual S. hermonthica AFLP markers linked to loci contributing to success on different rice-host cultivars.
Simulated null distribution for genetic differentiation among parasite sub-populations (Outlier analysis 1)
We simulated a null distribution of FST for each AFLP locus to test whether genetic differentiation at each locus was great enough to attribute its size to selection by the rice-host plants, in addition to sampling and error variation. For each of the 191 AFLP loci, we calculated null allele frequency and its standard error for the total (Kibos, Kenya) population of S. hermonthica (three sub-populations growing on different rice-host cultivars, 99 individuals). These allele frequencies were set to be those of the base population of seed, which were known with error given by the standard error of allele frequency. Next, the R package MASTERBAYES was used to simulate replicates of the experiment (identical in size to the empirical data) by drawing AFLP genotypes for each locus within each individual from the base population (Hadfield et al., 2006; JD Hadfield, personal communication). Uncertainty in the base allele frequencies was expressed by adding an error quantity drawn from a normal distribution with standard deviation equal to the standard error of allele frequency in the base population. This uncertainty in base allele frequencies was added during every iteration of the simulation. Genotyping error was introduced using MASTERBAYES to create a set of observed genotypes that, for some observations, differed from the true genotypes, depending on the allelic (not mismatch) genotyping error rate we observed in our experiment (Table 2). The observed (c.f. true underlying) bi-allelic genotypes were converted to binary AFLP phenotypes. FST (Cockerham and Weir's β) was calculated for each AFLP locus in the simulated data, between pairs of sub-populations separately, using three sub-populations of identical size to those in our experiment (99 individuals in total). The simulation was repeated a further 9999 times. The median, 95th and 99th quantiles (the latter representing 5 and 1% confidence level) for each of the 191 loci were calculated from these data for each pair of sub-populations.
The genetic differentiation (FST; Cockerham and Weir, 1993) at each of the 191 AFLP loci was calculated using the observed experimental data, separately for each pair of rice-host sub-populations. We considered AFLP loci with genetic differentiation in excess of the 99th quantile of the simulated null distribution of FST to be differentiated beyond the expectations of the null model, that is, this S. hermonthica genotype was significantly associated with rice-host cultivar. Except where stated, all analyses were carried out using R (R Development Core Team, 2008).
AFLP genotypes and the Striga resistance scores (and associated metadata) have been deposited in Dryad (http://datadryad.org).
Results
Resistance of rice cultivars to S. hermonthica parasitisation
To determine the resistance of the three rice cultivars to the different populations of S. hermonthica, the number of parasites that attached to the roots but subsequently failed to develop further by 21 days was expressed as a percentage of the total number of attachments (resistance score; Figure 1). The failure of S. hermonthica plants to develop post attachment to their rice host (host plant resistance score) was explained by an interaction between rice-host cultivar and parasite population (F=9.67; P<0.0001). Figure 1 suggests that this interaction was driven by differences in post-attachment success of the different parasite populations on rice cultivar Nipponbare (that is, differential resistance of this cultivar to parasite populations). When the data for cultivar Nipponbare were excluded from the modelling of host resistance using a generalised linear model, there was neither a significant host cultivar–parasite genotype interaction (F=0.35, P=0.708), nor a significant main effect of parasite population (F=0.13, P=0.875). Only a significant effect of rice-host cultivar on parasite resistance remained (F=149.52, P<0.0001). The rice cultivar IAC 165 was very susceptible to all three populations of S. hermonthica; 72.3% of attached parasites developed successfully. Kasalath exhibited an intermediate resistance to the three S. hermonthica populations. In this case, 29.7% of attached parasites developed normally. Nipponbare exhibited differential resistance to the populations of S. hermonthica. This cultivar was very resistant to the S. hermonthica population from Kibos, Kenya (only 12.5% of parasites developed normally; Figure 1) and most susceptible to parasites collected from millet in Gambia (47.4% of parasites developed normally).
Resistance scores calculated for the rice cultivars IAC165, Kasalath and Nipponbare when infected with different S. hermonthica populations 21 days after inoculation. The Kenyan population of S. hermonthica was collected from individuals parasitizing maize (cultivar H511). The Ivory Coast and Gambian parasite populations were collected from rice and millet, respectively. Resistance scores represent the proportion of parasites that failed to develop once attached to the roots. Values are means±s.e. of five replicates.
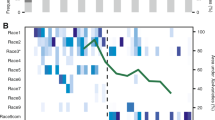
As a prelude to our genetic analysis of the Kenyan (Kibos) parasite population, we carried out an analysis of the development stage of the parasites from this population on the three rice cultivars 21 days after inoculation. This analysis indicated that the proportion of parasites that died or failed to develop on each host cultivar differed significantly (F=66.34, P<0.0001; Figure 2). The rice cultivar IAC 165 was the most susceptible; 71.6% of parasites that attached to the roots developed successfully (62.4% attached parasites per host plant at developmental stages S2–S4). The rice cultivar Nipponbare was the most resistant to infection by this S. hermonthica population. On average, 89% of parasites attached to each host root died very rapidly (that is, were necrotic) or failed to develop by day 21. Only a few parasites (normally fewer than 10 individuals per host plant) developed 2–3 pairs of leaves. Kasalath showed greater resistance to S. hermonthica than IAC 165 but less resistance than Nipponbare. On average, 53% of parasites that attached to the host roots died (Figure 2).
Development of S. hermonthica parasites (collected from Kibos, Kenya) on three different cultivars of rice-host plant. For each host cultivar, the graph shows the number of S. hermonthica plants attached to the host in each of five different developmental classes (S1, S2, S3, S4, S5) expressed as a percentage of the total number of plants that attached to the host: S1; the parasite radical had attached to the host root but leaf primordia had not emerged from the seed coat; S2, leaf primordia had emerged from the seed coat; S3, the parasite shoot had between one and three scale leaf pairs; S4, the parasite had four or more leaf pairs; S5, Parasites had attached but did not develop further and died. The total number of plants that attached to each rice-host cultivar is given in the panel to the left of the plot. Error bars are standard errors of the mean.
Genetic structure of the Kibos (Kenyan) population of S. hermonthica
Gene diversity of the Kibos population of S. hermonthica was 0.234, consistent with the species’ out-breeding mating system (Safa et al., 1984) and phenotypic diversity (for example, Jones and Safa, 1982). All three sub-populations of the Kibos population of S. hermonthica (that is, three sub-populations of parasites attached to different rice cultivar hosts) exhibited levels of genetic diversity similar to that observed in the total population (Table 3). The three sub-populations of S. hermonthica showed a small but significant level of genetic differentiation (FST=0.0131; P<0.001). When genetic differentiation of each pair of sub-populations was assessed separately, we found that Striga sub-populations growing on IAC-165 and Nipponbare rice hosts were the most genetically differentiated (FST=0.021; Table 3). These hosts also differed the most in their resistance phenotypes to the Kibos population of S. hermonthica (Figure 1). The lowest pair-wise genetic distance was observed when S. hermonthica sub-populations growing on the IAC-165 and Kasalath hosts were compared (FST=0.002). These hosts are more similar in their resistance phenotypes, and were the most susceptible to attack by parasites (Figure 1). An intermediate level of differentiation was observed between sub-populations growing on Kasalath and Nipponbare (FST=0.012).
Outlier analyses among rice-host sub-populations of S. hermonthica
We carried out two outlier analyses to identify AFLP loci that are putative markers for regions of the S. hermonthica genome controlling, or contributing to the differential success of the parasite on different rice-host cultivars (genotypes). First, we present the results of the outlier analysis that we tailored to our experimental design, including the genotyping error that we observed. We observed a total of 24 AFLP loci that were differentiated significantly relative to a simulated null model in which different rice-host cultivars do not exert selection on S. hermonthica genotype (Figure 3). We observed a range in genetic differentiation (FST) of 0.263–0.054 for these significantly differentiated loci (Figure 3), relative to a median background level of differentiation of <0.002 for any of the three sub-population pairs. We detected 19 loci as significant outliers (P<0.01) in the comparison of IAC-165 and Nipponbare hosted sub-populations, while only eight and three outliers were detected for the Kasalath–Nipponbare and IAC-165–Kasalath sub-population pairs, respectively (Figure 3). No loci were found to be significantly differentiated in all three sub-population pair-wise comparisons. However, five significantly differentiated loci were common to the IAC-165–Nipponbare and Kasalath–Nipponbare sub-population comparisons, while one differentiated locus was common to the IAC-165–Kasalath and IAC-165–Nipponbare sub-population comparison. There was a residual differentiation between the sub-populations (P<0.01) when the outlier loci were removed from the dataset, indicating that genetic differentiation at the subset of differentiated loci did not completely explain the differentiation observed on aggregate across all loci. In addition to the 24 loci whose genetic differentiation exceeded the 99th quantile of the null simulated distribution of FST, a further 38 loci showed levels of genetic differentiation that were in excess of the 95th quantile. The majority (82%) of these less-differentiated loci were identified in sub-population comparisons involving parasites from the Nipponbare rice-host sub-population. The second outlier analysis, carried out using the software BAYESCAN, indicated that only two outlier loci were under selection by rice-host cultivar (log10 Bayes Factor >0.5 for both loci; indicating ‘substantial’ evidence that these loci are under selection by rice-host cultivar). These loci were both detected in the IAC-165–Nipponbare S. hermonthica sub-population comparison, and they were also detected as significant outliers for the same sub-population comparison in the first outlier analysis.
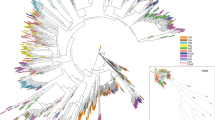
Observed genetic differentiation (FST) among pairs of S. hermonthica sub-populations for individual AFLP loci plotted against their null allele frequency. Lines indicate the quantiles of a simulated distribution of genetic differentiation under the null assumption that rice-host cultivars did not select for genetically differentiated sub-populations of S. hermonthica plants. Solid line, 99th quantile; dashed line, 95th quantile; fine dashed line, median. Loci whose genetic differentiation exceeds the 99th quantile have been shown using grey-filled triangular symbols.
Discussion
Here, we have used an experimental approach to investigate the host range (specificity) of three seed populations of S. hermonthica, using three rice-host cultivars that possess known contrasting parasite-resistance phenotypes. We also investigated the genetic component to host range within one of these S. hermonthica populations using AFLP molecular markers and an outlier analysis.
Variation for host range among parasite populations
All three of our parasite study populations showed some degree of success on different cultivars of rice-host plants (Figure 1). However, the success of parasites attached to rice hosts depended significantly on an interaction between rice cultivar and parasite population. This indicates a genetic component to host range and parasitic success among S. hermonthica populations, contingent on host genotype. The implications of this result are that there may be no simple route to breeding crops that are resistant to this parasite. Crop resistance developed against one population of S. hermonthica may not always be effective against others, thus emphasising the need to understand the genetic basis of host–parasite specificity.
Genetic variation for host range within parasite populations
S. hermonthica parasites from the Kibos population varied greatly and significantly in their developmental success after attaching to three rice-host cultivars (Figure 1). Relatively fewer S. hermonthica individuals were able to parasitise the most resistant cultivar of rice (Nipponbare) and the rate of development of these individuals was much slower compared with individuals on IAC 165 (Figure 2). These data are consistent with those reported in Gurney et al. (2006), where several quantitative trait loci underlying resistance in Nipponbare to this population of S. hermonthica were identified. Sub-populations of the Kibos population of S. hermonthica growing on the three rice-host cultivars were also significantly genetically differentiated from each other (Table 3). Together, these results indicate genetic variation for host range within the Kibos population. All three of the Kibos parasite sub-populations had originated, via one generation of selection on the different rice-host cultivars, from a single S. hermonthica seed population. Given this, the genetic differentiation we observed was not expected on the basis of preexisting differentiation. Furthermore, the genetic distance (FST) between pairs of S. hermonthica (Kibos) sub-populations corresponded to the relative difference in parasite resistance score of their rice-host cultivars. In other words, genetic differentiation was greatest between sub-populations of S. hermonthica growing on the most resistant (Nipponbare) and least resistant (IAC-165) cultivars of rice. We suggest that the most parsimonious explanation for our results is that the most resistant rice-host cultivar (Nipponbare) imposed relatively stronger selection on the genotype of establishing parasite seedlings than the other two rice-host cultivars. Our findings are in agreement with a similar study by Koyama (2000) in which sorghum cultivars differing in their resistance to S. hermonthica were grown in pots containing a S. hermonthica ecotype collected from a field in Alupe, Kenya. On the basis of isoenzyme and RAPD analyses, Koyama (2000) concluded that the parasite sub-populations on the resistant cultivars were significantly genetically differentiated from those on the susceptible cultivars.
Maintenance of variation for parasitic host range
When parasite populations have high evolutionary potential, parasitic virulence or migration rates, they are expected to become adapted to their sympatric hosts (Kaltz and Shykoff, 1998; Gandon and Michalakis, 2002). Some parasitic species show evidence of such adaptation (for example, Parker, 1985; Lively, 1989; Ebert, 1994), while in other cases, this has not been apparent (Morand et al., 1996; Mutikainen et al., 2000). In the Striga parasite–crop host system, evolution of the parasite population proceeds through short annual generation times in which virulence on the host plant can be very high. In addition, evolution in the host cereal crop is likely to be slow relative to that in the parasite. This is because crop adaptation is restricted by the farmer's choice of crop species and varieties, and farm management, including the practise of collecting and resowing crop seed, or moving it from other localities (Berner et al., 1994). Thus, we would expect to see local adaptation of populations of S. hermonthica to their sympatric host crop population (Gandon and Michalakis, 2002). Our results indicate that S. hermonthica populations retain a rather broader host range than the local adaptation argument would suggest. However, the rotation of crop cultivars and species that are each of relatively constant genetic background could provide an explanation for the maintenance of genetic variability for host range in S. hermonthica populations. This would act to retard tight adaptation of the parasite to individual crop varieties in favour of populations that are generalist (genetically diverse) with respect to their host plant genotype. Particular genotypes of the parasite would be favoured under certain crops or varieties to which they are better adapted, while other genotypes could be maintained in the population by the seed bank. This maintenance of genetic variation for host range within parasite populations could represent a potential (longer-term) cost to the farmer against any short-term yield advantage derived from crop rotation.
AFLP loci associated with host range
To determine which AFLP loci had responded to selection by rice-host cultivar, we conducted outlier analyses. The first of these analyses compared the observed value of genetic differentiation for each individual AFLP locus with a simulated distribution of values under the null hypothesis that rice cultivar had not exerted selection on parasite genotype. This analysis revealed a total of 24 AFLP loci (13% of the total) that were significantly differentiated in at least one parasite sub-population comparison. Six of these loci were significantly differentiated in more than one pair-wise comparison of S. hermonthica host-cultivar sub-populations. These six loci are candidate markers for regions of the S. hermonthica genome that may contain genes controlling parasite–host compatibility or host range. The total number of differentiated outlier loci (24) is perhaps surprising if all the loci represent separate genomic regions and if it reflects the true proportion of genes in the S. hermonthica genome responding to selection by host resistance genotype. Our second, more conservative outlier analysis supported the results of our first outlier analysis for two of the loci found to be differentiated in the IAC-165–Nipponbare S. hermonthica sub-population comparison. Although the two outlier analyses identified rather different numbers of AFLP loci responding to selection by host genotype, they both support the conclusion that S. hermonthica populations possess genetic variation for host range that is mediated by a subset of loci within the parasite genome. We suggest that, these outlier analyses bracket the probable true number of S. hermonthica genomic loci capable of responding to selection by rice-host cultivar (that is, controlling host range within parasite populations).
Our outlier analyses provide a first step towards the identification of loci controlling host range within individual populations of S. hermonthica. An understanding of the mechanisms of host compatibility and virulence in S. hermonthica, and their genetic basis will prove invaluable in understanding the response of this species to control measures based around resistant crop cultivars. Such knowledge may shed light on the specific types of parasite resistance that are likely to be most effective and long lasting, but will also reveal how S. hermonthica has come to be such a successful parasite species and such a damaging pest.
References
Beaumont MA, Nichols RA (1996). Evaluating loci for use in the genetic analysis of population structure. Proceed Royal Soc London B Biol Sci 263: 1619–1626.
Berner DK, Cardwell KF, Faturoti BO, Ikie FO, Williams OA (1994). Relative roles of wind, crop seeds, and cattle in dispersal of Striga spp. Plant Dis 78: 402–406.
Bharatha L, Werth CR, Musselman LJ (1990). A study of genetic diversity among host-specific populations of the witchweed Striga hermonthica (Scrophulariaceae) in Africa. Plant Syst Evol 172: 1–12.
Cockerham CC, Weir BS (1993). Estimation of gene flow from F-statistics. Evolution 47: 855–863.
Ebert D (1994). Virulence and local adaptation of a horizontally transmitted parasite. Science 265: 1084–1086.
Ejeta G (2007). The Striga scourge in Africa: a growing pandemic. In: Ejeta G, Gressel J (eds). Integrating New Technologies for Striga Control, Towards Ending the Witch-Hunt. World Scientific Publishing: Singapore. pp 3–15.
Foll M, Gaggiotti O (2008). A genome scan method to identify selected loci appropriate for both dominant and codominant markers: a bayesian perspective. Genetics 180: 977–995.
Gandon S (2002). Local adaptation and the geometry of host–parasite coevolution. Ecol Lett 5: 246–256.
Gandon S, Michalakis Y (2002). Local adaptation, evolutionary potential and host-parasite coevolution: interactions between migration, mutation, population size and generation time. J Evol Biol 15: 451–462.
Gandon S, Nuismer SL (2009). Interactions between genetic drift, gene flow, and selection mosaics drive parasite local adaptation. Am Nat 173: 212–224.
Gurney AL, Press MC, Scholes JD (2002a). Can wild relatives of sorghum provide new sources of resistance or tolerance against Striga species? Weed Res 42: 317–324.
Gurney AL, Taylor A, Mbwaga A, Scholes JD, Press MC (2002b). Do maize cultivars demonstrate tolerance to the parasitic weed Striga asiatica? Weed Res 42: 299–306.
Gurney AL, Slate J, Press MC, Scholes JD (2006). A novel form of resistance in rice to the angiosperm parasite Striga hermonthica. New Phytol 169: 199–208.
Hadfield JD, Richardson DS, Burke T (2006). Towards unbiassed parentage assignment: combining genetic, behavioural and spatial data in a Bayesian framework. Mol Ecol 15: 3715–3730.
Hewitt RJ (1966). Sand and Water Culture Methods Used in the Study of Plant Nutrition. Commonwealth Agricultural Bureaux: Wallingford, UK.
Jones BMG, Safa SB (1982). Variation in seed coat ornamentation in Striga hermonthica (Scrophulariaceae). Ann Bot 50: 629–634.
Kaltz O, Shykoff JA (1998). Local adaptation in host—parasite systems. Heredity 81: 361–370.
Koyama ML (2000). Genetic variability of Striga hermonthica and effect of resistant cultivars on Striga population dynamics. In: Haussman BIG, Hess DE, Koyama ML, Grivet L, Rattunde HFW and Geiger HH (eds). Breeding for Striga Resistance in Cereals. Margraf Verlag: Weikersheim, Germany, pp 247–260.
Li J, Lis KE, Timko MP (2009). Molecular genetics of race-specific resistance of cowpea to Striga gesnerioides. Pest Manage Sci 65: 520–527.
Lively CM (1989). Adaptation by a parasitic trematode to local populations of its snail host. Evolution 43: 1663–1671.
Luikart G, England PR, Tallmon D, Jordan S, Taberlet P (2003). The power and promise of population genomics: from genotyping to genome typing. Nat Rev Genet 4: 981–994.
Lynch M, Milligan BG (1994). Analysis of population genetic structure with RAPD markers. Mol Ecol 3: 91–99.
Mohamed KI, Musselman LJ, Riches CR (2001). The genus Striga (Scrophulariaceae) in Africa. Ann Missouri Botan Garden 88: 60–103.
Morand S, Manning SD, Woolhouse MEJ (1996). Parasite–host coevolution and geographic paterns of parasite infectivity and host suceptibility. Proceed R Soc London B Biol Sci 263: 119–128.
Mutikainen P, Salonen V, Puustinen S, Koskela T (2000). Local adaptation, resistance, and virulence in a hemiparastic plant-host plant interaction. Evolution 54: 433–440.
Parker C, Riches CR (1993). Parasitic Weeds of the World: Biology and Control. CAB International: Wallingford, UK.
Parker MA (1985). Local population differentiation for compatibility in an annual legume and its host-specific fungal pathogen. Evolution 39: 713–723.
R DCT (2008): R Foundation for Statistical Computing: Vienna, Austria.
Safa SB, Jones BMG, Musselman LJ (1984). Mechanisms favouring outbreeding in Striga hermonthica [Scrophulariaceae]. New Phytol 96: 299–305.
Scholes JD, Press MC (2008). Striga infestation of cereal crops- an unsolved problem in resource limited agriculture. Curr Opin Plant Biol 11: 1–7.
Swarbrick PJ, Scholes JD, Press MC, Slate J (2009). A major QTL for resistance of rice to the parasitic plant Striga hermonthica is not dependent on genetic background. Pest Manage Sci 65: 528–532.
Vasudeva Rao MJ, Musselman LJ (1987). Host specificity in Striga spp. and physiological strains. In: Musselman LJ (ed). Parasitic Weeds in Agriculture, Volume I Striga. CRC Press: Boca Raton, FL, pp 13–15.
Vekemans X, Beauwens T, Lemaire M, Roldan-Ruiz I (2002). Data from amplified fragment length polymorphism (AFLP) markers show indication of size homoplasy and of a relationship between the degree of homoplasy and fragment size. Mol Ecol 11: 139–151.
Whitlock R, Hipperson H, Mannarelli M, Burke T (2008a). A high-throughput protocol for extracting high-purity DNA from plants and animals. Mol Ecol Resourc 8: 736–741.
Whitlock R, Hipperson H, Mannarelli M, Butlin RK, Burke T (2008b). An objective, rapid and reproducible method for scoring AFLP peak-height data that minimizes genotyping error. Mol Ecol Resourc 8: 725–735.
Zhivotovsky LA (1999). Estimating population structure in diploids with multilocus dominant DNA markers. Mol Ecol 8: 907–913.
Acknowledgements
KH was funded by a scholarship from the Chinese government. We thank RK Butlin for comments on this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Huang, K., Whitlock, R., Press, M. et al. Variation for host range within and among populations of the parasitic plant Striga hermonthica. Heredity 108, 96–104 (2012). https://doi.org/10.1038/hdy.2011.52
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2011.52
Keywords
This article is cited by
-
High-throughput genotyping in onion reveals structure of genetic diversity and informative SNPs useful for molecular breeding
Molecular Breeding (2019)
-
Small-molecule antagonists of germination of the parasitic plant Striga hermonthica
Nature Chemical Biology (2016)
-
A candidate gene-based association study reveals SNPs significantly associated with bud burst in European beech (Fagus sylvatica L.)
Tree Genetics & Genomes (2015)