Abstract
The neotropical fish, Hoplias malabaricus, is well known for its population-specific karyotypic diversity and the variation of its sex chromosomes. Seven karyomorphs (A to G) have been previously described with an XY, X1X2Y and XY1Y2 sex chromosome system found in karyomorphs B, D and G, respectively. We compared the chromosomal characteristics of karyomorphs C and D using C-banding, staining with CMA3 and DAPI, and by mapping the location of 18S rDNA, 5SHindIII-DNA and (TTAGGG)n repeat sequences. Our results show conserved karyotypes in both karyomorphs, a nascent XX/XY sex chromosome system in karyomorph C and the origin of neo-Y chromosome in karyomorph D. The X and Y chromosomes of karyomorph C differ only slightly because of the amplification of repetitive sequences on the X chromosome, resulting in a homomorphic condition in all females and a heteromorphic condition in all males examined. Our study showed that chromosomes X and 20 of karyomorph C have similar patterns to the X1 and X2 chromosomes of karyomorph D, and are probably homologous. We showed that the neo-Y chromosome of karyomorph D shares similar patterns to the chromosomes Y and 20 of karyomorph C, and probably evolved through tandem fusion between Ypter/20pter. An interstitial site of the satellite 5SHindIII-DNA on the neo-Y reinforces the hypothesized dicentric nature of this chromosome. Our study shows the initial steps in XY chromosome differentiation in H. malabaricus and, in a broader context, contributes to the understanding of the evolutionary pathway leading to a multiple X1X2Y sex chromosome system in fishes.
Similar content being viewed by others
Introduction
Cytologically differentiated sex chromosomes have been described only in a few neotropical freshwater fish species. Distinct heterochromatic patterns between the XY and ZW sex chromosomes were well documented in some cases. The W chromosome is always highly heterochromatinized in species of the genus Triportheus (Bertollo and Cavallaro, 1992; Artoni et al., 1998) and in some species of Leporinus, Semaprochilodus, Microlepdogaster and Parodon (Galetti and Foresti, 1986; Feldberg et al., 1987; Andreata et al., 1993; Moreira-Filho et al., 1993). The addition of heterochromatin also occurs either in the X chromosome of Hoplias malabaricus (Born and Bertollo, 2000) and Eigenmannia virescens (Almeida-Toledo et al., 2001), or in the Y chromosome of Pseudotocinclus tietensis (Andreata et al., 1992). Even in multiple sex chromosome systems, which are usually derived from chromosomal rearrangements, heterochromatin can have a function in sex chromosome differentiation, as documented for Eigenmannia sp. 2 (Almeida-Toledo et al., 2000).
In recent years, the presence of transposable elements and other repetitive DNA sequences has been extensively studied in fish (Martins, 2007), providing important information on the differentiation and evolution of sex chromosomes (Nanda et al., 1990, 2000, 2002; Stein et al., 2001; Volff et al., 2003; Parise-Maltempi et al., 2007). These DNA elements are able to modify the molecular composition of the sex chromosomes, and to reduce the rate of recombination between them (Liu et al., 2004), which represent crucial steps in the differentiation of sex chromosomes (Charlesworth et al., 2005).
The widespread neotropical Erythrinidae fish H. malabaricus shows a conspicuous karyotypic diversification, with seven identified karyomorphs (A to G). These karyomorphs are easily distinguishable regarding the diploid number, the shape and size of chromosomes, and the sex chromosome system, which collectively suggest the existence of cryptic species (Bertollo et al., 1986, 2000). Three distinct sex chromosome systems have been reported, the XX/XY in karyomorph B, the X1X1X2X2/X1X2Y in karyomorph D and the XX/XY1Y2 in karyomorph G (Bertollo et al., 2000).
Meiotic analysis, along with G-banding, C-banding and replication banding patterns, improved the identification of sex chromosomes in karyomorph D and evidenced that in males a small submetacentric chromosome, similar to X2, was fused onto the short arms of a larger submetacentric chromosome, similar to X1, giving rise to a large neo-Y chromosome (Bertollo et al., 1997). Although different synapsis configurations could be observed, complete pairing of the sex trivalent was frequently found in pachytene cells suggesting a heterosynaptic process (Bertollo and Mestriner, 1998).
Karyotypic analysis suggested that karyomorph D (2n=40 females/2n=39 males) is more closely related to karyomorph C (2n=40 in both sexes), and that it is probably a derivation from a C-like karyomorph (Bertollo et al., 2000; Cioffi et al., 2009). The use of DNA sequences as probes for fluorescent in situ hybridization (FISH) analyses has greatly contributed to the study of fish sex chromosomes, not only because this method provides additional information about the structure of these chromosomes, but also because it enables the comparison of genomes of different species (Nanda et al., 2000). In this sense, repetitive DNA probes might be useful to differentiate heteromorphic sex chromosomes by signaling the accumulation of repetitive DNA sequences on one or both homologous of an ancestral homomorphic condition (Kejnovsky et al., 2009). We searched for specific chromosomal markers in karyomorphs C and D to highlight the evolution of the X1X2Y sex system. The chromosomes of H. malabaricus karyomorphs C and D were subjected to a comparative analysis through differential staining methods (C-banding, 4′,6-diamidino-2-phenylindole (DAPI) and chromomycin A3 (CMA3) fluorochromes), and FISH mapping of the 18S rDNA, satellite 5SHindIII-DNA and telomeric repeats on the chromosomes. These approaches allowed us to identify an XX/XY sex chromosome system in an early stage of differentiation in karyomorph C and to recognize the evolutionary pathway leading to the multiple X1X2Y sex chromosome system in karyomorph D.
Materials and methods
Specimens, mitotic chromosome preparation, chromosome staining and karyotyping
Nine females and 11 males of H. malabaricus belonging to karyomorph C, and 7 females and 10 males belonging to karyomorph D were collected from the Bento Gomes River (Poconé, Mato Grosso State, Brazil) and the Monjolinho stream (UFSCar reservoir, São Paulo State, Brazil), respectively. The specimens were deposited in the fish collection of the Cytogenetic Laboratory, Departamento de Genética e Evolução, Universidade Federal de São Carlos. Mitotic chromosomes were obtained from cell suspensions of the anterior kidney using the conventional air-drying method (Bertollo et al., 1978). Chromosomes were sequentially stained with Giemsa and C-banded using barium hydroxide (Sumner, 1972) to detect C-positive heterochromatin. Fluorochrome staining with GC-specific CMA3 and AT-specific DAPI was carried out as described by Sola et al. (1992). Images were captured by the software CoolSNAP system, Image Pro Plus, 4.1 (Media Cybernetics, Silver Spring, MD, USA) coupled to an Olympus BX50 microscope (Olympus Corporation, Ishikawa, Japan). Approximately 30 metaphase spreads were analyzed per specimen to determine the diploid chromosome number and karyotype structure. The chromosomes were classified as either metacentric (m) or submetacentric (sm), according to their arm ratios (Levan et al., 1964).
Meiotic chromosome analysis
Meiotic chromosomes were obtained according to the method of Kligerman and Bloom (1977), as described in Bertollo and Mestriner (1998). In brief, this method consists of sectioning the testes into small fragments and subjecting them to a hypotonic treatment. After tissue fixation, testis fragments were treated with 50% solution of glacial acetic acid and minced until a homogeneous cell suspension was obtained. Drops of this suspension were deposited on a cleaned slide heated at 30 °C, with the aid of a fine-tipped Pasteur pipette, and immediately sucked back, forming a cell ring of about 1 cm in diameter on the slide. Chromosomes were stained with 5% Giemsa solution in phosphate buffer with a pH of 6.8 for 5–6 min. Finally, the slide was rinsed with running water and air dried.
FISH probes
Two tandemly arrayed DNA sequences isolated from the H. malabaricus genome were used as probes. The first probe contained a copy of the repetitive satellite 5SHindIII-DNA sequence with 360 bp composed of a 95-bp segment similar to the 5S rRNA gene, and a 265-bp segment similar to the NTS of the 5S rRNA gene (Martins et al., 2006). The second probe corresponded to a 1400 bp segment of the 18S rRNA gene obtained by PCR from nuclear DNA (Cioffi et al., 2009). The 5SHindIII-DNA probe was labeled with biotin-14-dATP and the 18S rDNA probe was labeled with DIG-11-dUTP, both by nick translation, and according to the manufacturer's recommendations (Bionick Labeling System, Roche, Mannheim, Germany). The telomeric DNA sequence (TTAGGG)n was also used as a probe in males of the karyomorph D. This probe was generated by PCR (PCR DIG-Probe Synthesis Kit; Roche) in the absence of template using (TTAGGG)5 and (CCCTAA)5 as primers (Ijdo et al., 1991).
Fluorescent in situ hybridization
FISH analysis was performed on mitotic and meiotic chromosome spreads (Pinkel et al., 1986). The metaphase chromosome slides were incubated with RNAse (40 μg ml−1) for 1.5 h at 37 °C. After denaturation of chromosomal DNA with 70% formamide/2 × SSC at 70 °C, spreads were incubated in 2 × SSC for 4 min at 70 °C. The hybridization solution (100 ng of denatured probe, 10 mg ml−1 dextran sulfate, 2 × SSC and 50% formamide in a final volume of 30 μl) was applied to the slides, and the hybridization was performed overnight at 37 °C in a moist chamber containing 2 × SSC. Posthybridization washes were carried out at 37°C in 2 × SSC with 50% formamide for 15 min, followed by a second wash in 2 × SSC for 15 min and a final wash at room temperature in 4 × SSC for 15 min. The signal detection was by avidin-FITC (Sigma, St Louis, MO, USA) for the 5SHindIII-DNA probe and anti-digoxigenin-rhodamine (Roche) for the 18S rDNA and (TTAGGG)n probes. The posthybridization washes were performed on a shaker (150 r.p.m.). The chromosomes were counterstained with propidium iodide (50 μg ml−1) or DAPI (1.2 μg ml−1) and mounted in antifade solution (Vector, Burlingame, CA, USA). FISH analysis was carried out using an epifluorescence microscope Olympus BX50 (Olympus Corporation).
Results
Karyotypes, C-banding and fluorochrome staining
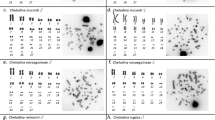
All specimens of the karyomorph C had 2n=40 chromosomes (14 m+26 sm) in both sexes (Figure 1). In karyomorph D, all females had 2n=40 chromosomes (14 m+26 sm) and all males had 2n=39 chromosomes (14 m+25 sm). The karyotype composition of females is the same as that found in karyomorph C. The specific male karyotype is determined by the characteristic multiple sex chromosome system of this karyomorph, with X1X1X2X2 females and X1X2Y males, and where the neo-Y corresponds to a large submetacentric chromosome (Figure 1).
Karyotypes of Hoplias malabaricus karyomorphs C and D (male/female) arranged from sequentially Giemsa-stained (left) and C-banded chromosomes (right). In karyomorph C, males and females present the same Giemsa-stained and C-banded karyotype. The X and Y chromosomes of karyomorph C (pair no. 11) and the X1, X2 and neo-Y chromosomes of karyomorph D are boxed. Note the conspicuous proximal heterochromatic band on the long arms of the X, Y and X1 chromosomes and on the short arms of the neo-Y chromosome. Bar=5 μm.
In both karyomorphs, C-positive heterochromatic bands were found in the centromeric/pericentromeric region of all chromosomes and in the telomeric region of several pairs (Figure 1). In general, C-banding did not identify sex chromosomes heteromorphy in karyomorph C. However, in some male metaphases, the proximal C-positive bands on the long arms of the chromosome pair no. 11 were slightly distinct in size. Moreover, the heterochromatic pattern of chromosome pair nos. 11 and 20 in both males and females were the same as those of chromosomes X1 and X2 of karyomorph D, suggesting that they might be homologous. The conspicuous proximal heterochromatic block found on the long arms of the chromosome pair no. 11 of karyomorph C was observed on the same region of the X1 chromosome, and on the short arms of the neo-Y chromosome.
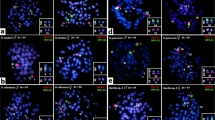
GC-rich heterochromatin characterized these marked C-bands of both karyomorphs (Figure 2). Remarkably, in all male specimens of karyomorph C, the chromosome pair no. 11 showed heteromorphic CMA3+ sites, with one of the chromosomes clearly bearing a larger site than the other. Coincidentally, in all males from karyomorph D, the neo-Y and the X1 chromosomes showed a smaller and a larger CMA3+ site, respectively (Figures 2 and 3). Because all females from both karyomorphs showed homomorphic CMA3+ sites, a statistical test to confirm the males' exclusive heteromorphism was not necessary. These results were supported by DAPI staining, which shows that the CMA3+ sites were DAPI-negative and with the same distinct sizes (Figures 2 and 3). Thus, the chromosome pair no. 11 of males from karyomorph C is heteromorphic on fluorochrome banding basis, allowing the identification of a cryptic XX/XY sex chromosome system in this karyomorph. However, this heteromorphism did not change the macrostructure of this chromosome pair because it kept the same general morphology and overall size.
Metaphase plates of Hoplias malabaricus karyomorphs C and D showing the location of the DAPI−, chromomycin A3+, 5SHindIII-DNA and 18S rDNA sites on the chromosomes. The DAPI, CMA3 and 18S rDNA sequences, respectively, occur as homomorphic and heteromorphic sites in all females and males examined. The correspondence between the X and Y chromosomes (karyomorph C) and X1 and neo-Y chromosomes (karyomorph D) is evident. Centromeric 5SHindIII-DNA sites are shown in chromosome 20 (karyomorph C), in the X2 (karyomorph D) as well as interstitially on the long arms of the neo-Y chromosome. Arrows indicate conspicuous proximal heterochromatic blocks on the propidium iodide-counterstained X, Y, X1 and neo-Y chromosomes. Bar=5 μm.
The X and Y chromosomes of karyomorph C and the X1 and neo-Y chromosomes of karyomorph D showing C-positive, DAPI− and CMA3+ bands as well as the hybridization sites of 5SHindIII-DNA and 18S rDNA. The patterns on the X and Y are similar to those on the X1 and neo-Y chromosomes. The correspondence between the interstitial 5SHindIII-DNA site on the long arms of the neo-Y chromosome and the centromeric site of the X2 and chromosome 20 of karyomorph C is also highlighted. The X, Y, X1 and neo-Y chromosomes lack a centromeric 5SHindIII-DNA site and show conspicuous proximal heterochromatic blocks after propidium iodide counterstaining. (a) Male metaphase plate of karyomorph D with telomeric hybridization signals on both telomeres of all chromosomes. (b) Inset showing the interstitial telomeric site (ITS) on the long arms of the neo-Y chromosome (arrowed). Bar=5 μm.
5SHindIII-DNA, 18S rDNA and (TTAGGG)n repeats
The satellite 5SHindIII-DNA was mapped in the centromeric region of several chromosome pairs of karyomorphs C and D, with a total of 22 corresponding sites. Among them, those located on chromosome pair no. 20 of karyomorph C and the corresponding X2 chromosome of karyomorph D were noteworthy. The neo-Y chromosome of karyomorph D displayed a characteristic interstitial site on the long arms, the only one located in a noncentromeric position in the whole karyotype (Figures 2 and 3).
Five chromosome pairs had 18S rDNA sites in both karyomorphs, mainly located in their telomeric regions. However, the X and Y chromosomes (pair no. 11) of karyomorph C and the X1 and neo-Y chromosomes of karyomorph D shared a conspicuous proximal site located on the long arms of the X, Y and X1 chromosomes, and on the short arms of the neo-Y chromosome. Again, as found with DAPI and CMA3 staining, the XY chromosomes in karyomorph C and the X1 and neo-Y chromosomes in karyomorph D showed heteromorphism in all males, with a homomorphic condition being found only in females of both karyomorphs. Therefore, these results also suggest the occurrence of an XY sex chromosome system in karyomorph C (Figures 2 and 3).
FISH analysis using the (TTAGGG)n repeats probe showed typical telomeric hybridization signals on both telomeres of all chromosomes. In addition, an exclusive interstitial telomeric site (ITS) was located on the long arms of the neo-Y chromosome (Figures 3a and b).
Meiotic analysis
Meiotic chromosomes were analyzed in male specimens of karyomorph D using FISH analysis with the 18S rDNA probe. Spermatogonial cells had 39 chromosomes and 10 sites of 18S rDNA, as expected on the basis of results for mitotic metaphases (Figure 4a). Cells in early prophase I showed five paired 18S rDNA sites because of the homologous pairing during the zygotene stage (Figure 4b). Eighteen bivalents and one characteristic trivalent were found during diplotene-diakinesis. These bivalents corresponded to the 18 synapsed autosomal pairs, whereas the trivalent corresponded to the X1, X2 and neo-Y sex chromosomes (Figures 4c and d). Consequently, five 18S rDNA sites were detected, including the one located on the sex trivalent. Spermatocytes in metaphase II had 19 or 20 chromosomes, as expected from spermatogonial cells with 2n=39 chromosomes. Cells with n=19 (18 autosomes+neo-Y chromosome), as well as those with n=20 (18 autosomes+X1X2 chromosomes), presented five 18S rDNA signals. Despite the large degree of condensation in meiotic chromosomes, the heteromorphic sites could also be detected between the neo-Y and the X1 chromosomes (Figures 4e and f).
Meiotic chromosomes of male Hoplias malabaricus karyomorph D analyzed with an 18S rDNA probe. (a) Spermatogonial cells showing 2n=39 chromosomes and ten 18S rDNA sites; (b) zygotene/pachytene stage and (c, d) diakinesis/metaphase I stage, displaying five 18S rDNA sites in the synapsed chromosomes; (e, f) metaphase II cells with 19 and 20 chromosomes, respectively, each showing five 18S rDNA sites. Arrows indicate the conspicuous ribosomal site on the sex trivalent; arrowheads indicate the heteromorphic 18S rDNA sites associated with the neo-Y and X1 chromosomes. Bar=5 μm.
Discussion
Chromosomal mapping of repetitive DNA sequences shows that karyomorphs C and D are similar in their karyotypic structure. However, the presence of cryptically differentiated XY chromosomes in karyomorph C provides insights into the likely evolutionary pathway leading to the multiple X1X2Y sex chromosome system in karyomorph D.
The X and Y chromosomes of karyomorph C differ only slightly because of the amplification of repetitive sequences on the X chromosome. This heteromorphic condition was not always clearly detected on the pattern of C-positive bands, but it was evident regarding the GC-rich heterochromatin and 18S rDNA sequences. Therefore, both repetitive DNA classes seem to have a function in this stage of the sex chromosome differentiation. However, the differences between the X and Y chromosomes were not enough to modify the overall size of this chromosome pair, thus reinforcing the presence of a nascent sex chromosome system in karyomorph C. The similar heteromorphic conditions of the DAPI−/CMA3+ and 18S rDNA sites could be explained if the ribosomal genes are interspersed throughout the GC-rich heterochromatic sequences or, alternatively, if the rDNA constitutes these sequences itself. However, the first hypothesis looks to be more plausible given that in Atlantic salmon, and perhaps in other fishes, the rRNA genes are interspersed with the heterochromatic regions (Pendás et al., 1993), and that CMA3 primarily stains the flanking heterochromatin of the NORs (Philips and Hartley, 1988). It is thought that nascent sex chromosomes retain extensive homology with some differentiated areas that are associated with sex determination (Charlesworth, 2004; Charlesworth et al., 2005). The redundancy of the rDNA sequences and the GC-rich-associated heterochromatin could make these chromosomal regions more susceptible to unequal crossing-over. Consequently, the amount of the repetitive sequences might be modified by duplications and deletions during the evolutionary process.
The chromosomal pattern of karyomorph C was compared with that of karyomorph D to analyze their karyological relationships and the origin of the multiple X1X2Y sex chromosome system. The same heteromorphism in size, concerning the patterns of DAPI−/CMA3+ and 18S rDNA sites found between the X and Y chromosomes of karyomorph C, was also found between the X1 and neo-Y chromosomes of karyomorph D. These results indicate that these chromosomes are probably homologous. Only one conspicuous site of 18S rDNA was observed in the meiotic trivalent of karyomorph D as a result of the X1 and neo-Y chromosome pairing. Similarly, a less pronounced 18S rDNA site on the neo-Y chromosome compared with the conspicuous one found in the X1 chromosome was observed. Thus, meiotic data in karyomorph D supported the heteromorphism observed in the mitotic chromosomes.
The chromosome mapping of 5SHindIII-DNA sequences also provided evidence of the evolutionary relationships between the sex chromosomes of both karyomorphs. The 5SHindIII is a satellite DNA family specific to the H. malabaricus genome, and it is located only in the centromeric region of some chromosome pairs (Martins et al., 2006). Such sequences were found neither on the centromeres of the X and Y chromosomes in karyomorph C and nor on the centromeres of the X1 and the neo-Y chromosomes in karyomorph D. However, the chromosomal pair no. 20 in karyomorph C and its homolog (X2) in the karyomorph D displayed centromeric signals, in addition to an exclusive interstitial site on the long arms of the neo-Y chromosome. These findings suggest that this interstitial site corresponds to the centromere of chromosome 20 fused onto the ancestral Y, giving rise to a dicentric neo-Y chromosome. These results also support the proposition that this chromosome behaves as a dicentric but stable component of the karyotype (Bertollo et al., 1997; Bertollo and Mestriner, 1998; Martins et al., 2006).
ITSs, mapped by FISH analysis on the long arms of the neo-Y chromosome, also emphasized the evolutionary hypothesis of the origin of the X1X2Y sex chromosome system. As expected, fluorescent sites were found on both telomeric regions of all chromosomes of the karyotype, whereas the neo-Y chromosome highlighted an unusual interstitial site on the long arms. ITSs have been found in the pericentromeric regions of many species of vertebrate, suggesting that chromosomal rearrangements can occur without the loss of telomeric sequences (Meyne et al., 1989). In fact, the general hypothesis that ITSs may be remnants of chromosome rearrangements that occurred during genome evolution has been supported by several observations (Ijdo et al., 1991). In the case of H. malabaricus, the ITS was mapped only to the neo-Y chromosome region where the ancestral Y was fused onto chromosome no. 20. Thus, this indicates the maintenance of some telomeric sequences from the rearranged chromosomes and supports the origin of the neo-Y chromosome. The chromosomal markers used in this study collectively suggest that the neo-Y chromosome resulted from a tandem fusion involving Ypter and 20pter (Figure 5).
Schematic diagram showing the derivation of the X1X2Y sex chromosome system of karyomorph D from the cryptic XX/XY sex chromosome system of karyomorph C, evidenced by the mapping of repetitive DNA sequences on the chromosomes. The central box illustrates the tandem fusion Ypter/20pter resulting in the neo-Y chromosome. The curled arrow is used to indicate the inverted orientation of Y.
The absence of differentiated sex chromosomes in most fish species (and because this type of analysis is generally not possible in higher vertebrates because of their relatively stable sex-determining systems, see Dettai et al., 2007) makes H. malabaricus an excellent model for the study of the differentiation and evolution of sex chromosomes. In addition to the previously identified XY, X1X2Y and XY1Y2 sex systems, a new variant is documented herein where XY chromosomes can be seen in an early stage of differentiation. Although the results show a close association between the XY and the X1X2Y systems of karyomorphs C and D, similar findings have not been reported for karyomorph B, where the morphology of the XY sex chromosomes is different from the one found in karyomorph C, thus highlighting the plasticity of sex chromosome differentiation in H. malabaricus.
References
Almeida-Toledo LF, Foresti F, Daniel MFZ, Toledo-Filho SA (2000). Sex chromosome evolution in fish: the formation of the neo-Y chromosome in Eigenmannia (Gymnotiformes). Chromosoma 109: 197–200.
Almeida-Toledo LF, Foresti F, Pequignot EV, Daniel-Silva MF (2001). XX:XY sex chromosome system with X heterochromatinization: an early stage of sex chromosome differentiation in the Neotropic electric eel Eigenmannia virescens. Cytogenet Cell Genet 95: 73–78.
Andreata AA, Almeida-Toledo LF, Oliveira C, Toledo-Filho SA (1992). Chromosome studies in Hypoptopomatinae (Pisces, Siluriformes, Loricariidae): I. XX/XY sex chromosome heteromorphism in Pseudotocinclus tietensis. Cytologia 57: 369–372.
Andreata AA, Almeida-Toledo LF, Oliveira C, Toledo-Filho SA (1993). Chromosome studies in Hypoptopomatinae (Pisces, Siluriformes, Loricariidae). II. ZZ/ZW sex-chromosome system, B chromosomes, and constitutive heterochromatin differentiation in Microlepidogaster leucofrenatus. Cytogenet Cell Genet 63: 215–220.
Artoni RF, Venere PC, Bertollo LAC (1998). A heteromorphic ZZ/ZW sex chromosome system in fish, genus Hypostomus (Loricariidae). Cytologia 63: 421–425.
Bertollo LAC, Born GG, Dergam JA, Fenocchio AS, Moreira-Filho O (2000). A biodiversity approach in the neotropical fish Hoplias malabaricus. Karyotypic survey, geographic distribution of cytotypes and cytotaxonomic considerations. Chromosome Res 8: 603–613.
Bertollo LAC, Cavallaro ZI (1992). A highly differentiated ZZ/ZW sex chromosome system in a Characidae fish, Triportheus guentheri. Cytogenet Cell Genet 60: 60–63.
Bertollo LAC, Fontes MS, Fenocchio AS, Cano J (1997). The X1X2Y sex chromosome system in the fish Hoplias malabaricus. I. G-, C- and chromosome replication banding. Chromosome Res 5: 493–499.
Bertollo LAC, Mestriner CA (1998). The X1X2Y sex chromosome system in the fish Hoplias malabaricus (Pisces, Erythrinidae). II. Meiotic analyses. Chromosome Res 6: 141–147.
Bertollo LAC, Moreira-Filho O, Galetti Jr. PM (1986). Cytogenetics and taxonomy considerations based on chromosome studies of freshwater fish. J Fish Biol 28: 153–159.
Bertollo LAC, Takahashi CS, Moreira-Filho O (1978). Cytotaxonomic considerations on Hoplias lacerdae (Pisces, Erythrinidae). Brazil J Genet 1: 103–120.
Born GG, Bertollo LAC (2000). An XX/XY sex chromosome system in a fish species, Hoplias malabaricus, with a polymorphic NOR-bearing X chromosome. Chromosome Res 8: 111–118.
Charlesworth B (2004). Sex determination: primitive Y chromosomes in fish. Curr Biol 14: 745–747.
Charlesworth D, Charlesworth B, Marais G (2005). Steps in the evolution of heteromorphic sex chromosomes. Heredity 95: 118–128.
Cioffi MB, Martins C, Bertollo LAC (2009). Comparative chromosome mapping of repetitive sequences. Implications for genomic evolution in the fish Hoplias malabaricus. BMC Genetics 10: 34.
Dettai A, Bouneau L, Fischer C, Schultheis C, Schmidt C, Zhou Q et al. (2007). FISH analysis of fish transposable elements: tracking down mobile DNA in teleost genomes. In: Pisano E, Ozouf-Costaz C, Foresti F, Kapoor BG (eds). Fish Cytogenetics. Science Publisher: New Hampshire, pp 361–383.
Feldberg E, Bertollo LAC, Almeida-Toledo LF, Foresti F, Moreira-Filho O, Santos AF (1987). Biological aspects of Amazonian fishes. IX. Cytogenetic studies in two species of the genus Semaprochilodus (Pisces, Prochilodontidae). Genome 29: 1–4.
Galetti Jr. PM, Foresti F (1986). Evolution of the ZZ/ZW system in Leporinus (Pisces, Anostomidae). Cytogenet Cell Genet 43: 43–46.
Ijdo JW, Wells RA, Baldini A, Reeders ST (1991). Improved telomere detection using a telomere repeat probe (TTAGGG)n generated by PCR. Nucleic Acids Res 19: 4780.
Kejnovsky E, Hobza R, Cermák T, Kubát Z, Vyskot B (2009). The role of repetitive DNA in structure and evolution of sex chromosomes in plants. Heredity 102: 533–541.
Kligerman AD, Bloom SE (1977). Rapid chromosome preparations from solid tissues of fishes. J Fish Res Board Can 34: 266–269.
Levan A, Fredga K, Sandberg AA (1964). Nomenclature for centromeric position on chromosomes. Hereditas 52: 201–220.
Liu ZY, Moore PH, Ma H, Ackerman CM, Ragiba M, Yu Q et al. (2004). A primitive Y chromosome in papaya marks incipient sex chromosome evolution. Nature 427: 348–352.
Martins C (2007). Chromosomes and repetitive DNAs: a contribution to the knowledge of fish genome. In: Pisano E, Ozouf-Costaz C Foresti F, Kapoor BG (eds). Fish Cytogenetics. Science Publisher: New Hampshire, pp 421–453.
Martins C, Ferreira IA, Oliveira C, Foresti F, Galetti Jr. PM (2006). A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica 127: 133–141.
Meyne J, Ratliff RL, Moyzis RK (1989). Conservation of the human telomere sequence (TTAGGG)n among vertebrates. Proc Natl Acad Sci USA 86: 7049–7053.
Moreira-Filho O, Bertollo LAC, Galetti PM Jr. (1993). Distribution of sex chromosome mechanisms in neotropical fish and description of a ZZ/ZW system in Parodon hilarii (Parodontidae). Caryologia 46: 115–125.
Nanda I, Feichtinger W, Schmid M, Schroder JH, Zischler H, Epplen JC (1990). Simple repetitive sequences are associated with differentiation of the sex chromosomes in the guppy fish. J Mol Evol 30: 456–462.
Nanda I, Volff JN, Weis S, Körting C, Froschauer A, Schmid M et al. (2000). Amplification of a long terminal repeat-like element on the Y chromosome of the platyfish, Xiphophorus maculates. Chromosoma 109: 173–180.
Nanda I, Kondo M, Hornung U, Asakawa S, Winkler C, Shimizu A et al. (2002). A duplicated copy of DMRT1 in the sex determining region of the Y chromosome of the medaka, Oryzias latipes. Proc Natl Acad Sci USA 99: 11778–11783.
Parise-Maltempi PP, Martins C, Oliveira C, Foresti F (2007). Identification of a new repetitive element in the sex chromosomes of Leporinus elongatus (Teleostei: Characiformes: Anostomidae): new insights into the sex chromosomes of Leporinus. Cytogenet Genome Res 116: 218–223.
Pendás AM, Morán P, Garcia-Vázquez E (1993). Ribosomal RNA genes are interspersed throughout a heterochromatic chromosome arm in Atlantic salmon. Cytogenet Cell Genet 63: 128–130.
Philips RB, Hartley SE (1988). Fluorescent banding patterns of the chromosomes of the genus Salmo. Genome 30: 193–197.
Pinkel D, Straume T, Gray J (1986). Cytogenetic analysis using quantitative, high sensitivity, fluorescence hybridization. Proc Natl Acad Sci USA 83: 2934–2938.
Sola L, Rossi AR, Laselli V, Rasch EM, Monaco PJ (1992). Cytogenetics of bisexual/unisexual species of Poecilia II. Analysis of heterochromatin and nucleolar organizer regions in Poecilia mexicana mexicana by C-banding and DAPI, quinacrine, chromomycin A3, and silver staining. Cytogenet Cell Genet 60: 229–235.
Stein J, Phillips RB, Devlin RH (2001). Identification of the Y chromosome in Chinook salmon (Oncorhynchus tshawytscha). Cytogenet Cell Genet 92: 108–110.
Sumner AT (1972). A simple technique for demonstrating centromeric heterochromatin. Exp Cell Res 75: 304–306.
Volff JN, Bouneau L, Ozouf-Costaz C, Fischer C (2003). Diversity of retrotransposable elements in compact pufferfish genomes. Trends Genet 19: 674–678.
Acknowledgements
We thank Dr Liano Centofante for providing fish specimens. This work was supported by the Brazilian agencies FAPESP (Fundação de Amparo à Pesquisa do Estado de São Paulo—Proc. No. 2007/05565-5), CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico—Proc. No. 305483/2006-4) and CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Proc. No. 083/2007).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Cioffi, M., Bertollo, L. Initial steps in XY chromosome differentiation in Hoplias malabaricus and the origin of an X1X2Y sex chromosome system in this fish group. Heredity 105, 554–561 (2010). https://doi.org/10.1038/hdy.2010.18
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2010.18
Keywords
This article is cited by
-
Homeology of sex chromosomes in Amazonian Harttia armored catfishes supports the X-fission hypothesis for the X1X2Y sex chromosome system origin
Scientific Reports (2023)
-
Sex determination mechanisms and sex control approaches in aquaculture animals
Science China Life Sciences (2022)
-
Against the mainstream: exceptional evolutionary stability of ZW sex chromosomes across the fish families Triportheidae and Gasteropelecidae (Teleostei: Characiformes)
Chromosome Research (2021)
-
Chromosomal mapping of repetitive sequences in Hyphessobrycon eques (Characiformes, Characidae): a special case of the spreading of 5S rDNA clusters in a genome
Genetica (2020)
-
Isolation of a Male-Specific Molecular Marker and Development of a Genetic Sex Identification Technique in Spotted Knifejaw (Oplegnathus punctatus)
Marine Biotechnology (2020)