Abstract
Recent advances in the statistical analysis of microsatellite data permit calculation of sex-specific dispersal rates through sex- and age-specific comparisons of genetic variation. This approach, developed for the analysis of data derived from co-dominant autosomal markers, should be applicable to a sex-specific marker such as mitochondrial DNA. To test this premise, we amplified a 449 bp control region DNA sequence from the mitochondrial genome of the collared peccary (Pecari tajacu), and estimated intra-class correlations among herds sampled from three Texas populations. Analyses on data partitioned by breeding group showed a clear signal of male-biased dispersal; sex-specific fixation indices associated with genetic variation among social groups within populations yielded values for females (FGP=0.91), which were significantly larger than values for males (FGP=0.24; P=0.0015). The same general pattern emerged when the analyses were conducted on age classes (albeit nonsignificantly), as well as categories of individuals that were predicted a posteriori to be dispersers (adult males) and philopatric (adult females and all immatures). By extending a previously published methodology based on biparentally inherited markers to matrilineally inherited haploid data, we calculated sex-specific rates of contemporary dispersal among social groups within populations (m♂=0.37). These results support the idea that mitochondrial DNA haplotype frequency data can be used to estimate sex-specific instantaneous dispersal rates in a social species.
Similar content being viewed by others
Introduction
Sex bias in natal dispersal is common; in most mammalian species, males are dispersers whereas females are philopatric, and the opposite trend is shown in birds (Greenwood, 1980). Exploring why the sexes differ in their dispersal patterns can shed light on the evolutionary causes of dispersal in general (Goudet et al., 2002), and accurate characterization of dispersal behavior is integral to our understanding of the social structure, mating system and population genetic structure of a species. Yet detection of sex-biased dispersal can be tricky because a dispersal event may occur once in an animal's lifetime, and such events can be difficult to observe directly.
Measuring dispersal
In the last few decades molecular genetics has provided a means of investigating sex-biased dispersal within and among populations (reviewed in Lawson-Handley and Perrin, 2007). Several powerful approaches have been developed to detect individual dispersers through assignment tests or to characterize general patterns of dispersal through summary statistics of population genetic structure (F-statistics, relatedness). Most of these approaches utilize autosomal microsatellites as molecular markers, either alone (Mossman and Waser, 1999; Petit et al., 2001; Waser et al., 2001; Goudet et al., 2002) or in tandem with a uniparentally inherited marker such as mitochondrial DNA (mtDNA) or a Y-chromosome locus (Girman et al., 1997; Escorza-Trevino and Dizon, 2000). The expectation inherent to all these approaches is that greater genetic structure will be evident in the philopatric sex compared with the dispersing sex, thus comparisons of sex-specific FST estimates should reveal the direction (and suggest the relative strength) of sex-bias in dispersal (Goudet et al., 2002).
Because mitochondrial DNA is matrilineally inherited, it is commonly used to infer female-biased dispersal rates (Prugnolle and de Meeus, 2002). When mtDNA haplotype distribution patterns are examined in isolation, inferences can be made about female dispersal behavior without respect to males, but this approach is qualitative and not widely applied (Hoelzer et al., 1994). However, it is possible to use mtDNA alone to infer the relative dispersal of both sexes by extending methods developed for autosomal, bi-parentally inherited markers. For instance, the comparisons of sex-specific population differentiation from haplotype frequency data can indicate which sex disperses more (Escorza-Trevino and Dizon, 2000; Yang et al., 2003).
Using sex-specific fixation indices to estimate instantaneous dispersal rates
Vitalis (2002) developed a method to quantitatively measure sex bias in instantaneous dispersal rates using data from biparentally inherited markers such as microsatellites. This approach allows the inference of sex-specific dispersal rates by comparing sex-specific estimates of genetic differentiation (FST) measured before and after dispersal. This intuitive method can be further extended to incorporate the hierarchical structure within social species (Fontanillas et al., 2004), as it has been recognized that social organization can strongly influence correlations of gene frequencies (Chesser, 1991; Slatkin and Voelm, 1991; Sugg and Chesser, 1994; Chesser and Baker, 1996; Vigouroux and Couvet, 2000). In this study, we develop and use an extension of the Vitalis' (2002) method to estimate instantaneous dispersal rates through analyses of mtDNA haplotype distribution patterns in a social mammal, the collared peccary (Pecari tajacu, family Tayassuidae).
We sampled extensively within three populations separated by long distances, with the goal of quantifying local dispersal among breeding groups within populations. We then compared sex- and age-specific estimates of population differentiation based solely on mtDNA haplotype frequencies, using probability-based estimates of intra-class correlations of gene frequencies among social groups within populations. We used a resampling approach to test for the significance of the observed age and sex-bias in dispersal. Last, the fixation indices generated by these analyses were used to calculate single-generation sex-specific dispersal rates. Earlier mtDNA was used primarily to infer female dispersal patterns, but we show that this matrilineally inherited genetic marker can be used to quantify male dispersal rates in the absence of nuclear population genetic data.
Materials and methods
Study species
The collared peccary is a socially complex, pig-like ungulate that forms stable, mixed sex herds of 3 to 30 individuals (Sowls, 1978). These groups associate throughout the year and vigorously defend territories against other social groups (Ellisor and Harwell, 1969; Bissonette, 1982; Hellgren et al., 1984). Herds are socially cohesive and attempts to immigrate may be met with aggression, although direct observational data on dispersal behavior are still scarce. Male exchange between groups and solitary wandering of both sexes has been observed but natal dispersal has not been adequately described (Ellisor and Harwell, 1969; Day, 1985; Gabor and Hellgren, 2000). Earlier little population genetic data existed for P. tajacu (but see Gongora et al., 2006). Theimer and Keim (1994) utilized mtDNA variation to measure sequence divergence and geographic partitioning in Arizona populations, but their samples were not associated with social groups. There was sufficient heterogeneity in mtDNA haplotype distribution to indicate limited female dispersal across regions (rather than among neighboring herds as is considered here), although it was not clear if the patterns observed were also a signature of founding events (Theimer and Keim, 1994).
Sampling
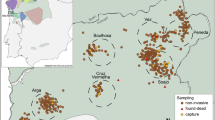
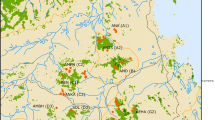
Data were collected from three wild populations of P. tajacu in Texas. In the mid-1990s, 102 whole blood samples were collected from the Chaparral Wildlife Management Area (CWMA) in south Texas (Gabor and Hellgren, 2000). These samples were taken from live-trapped animals from 13 social groups, but not all group members were sampled. In 2005, we collected 31 ear snip tissue samples from live-trapped animals from four groups in the Welder Wildlife Refuge (WWR) in south Texas. In 2006–2007 we similarly sampled 134 animals from 13 groups in Big Bend Ranch State Park (BB) in west Texas, along the Texas–Mexico border. The WWR and BB populations were sampled extensively; every social group at these locations was identified through direct and remote camera observation and trapped in large corrals over several sessions. Groups ranged in size from 2 to 18 animals and mean group size was 8.9. Individuals were uniquely marked with numbered ear tags and the strongest possible effort was made to trap and sample every unmarked individual. All samples include associated data on age class (adult, subadult, juvenile, infant), sex, territory location and social group affiliation. Age class was assigned according to behavior and morphological traits such as pelage, body size and testicular development. Individuals showing immature characteristics such as ginger or spotted pelage, undescended or partially descended testicles, adult-oriented following behavior, or estimated body size of less than 9 kg were classed as infants or juveniles, whereas individuals that weighed 10–13 kg were classed as ‘subadults’ that were on the cusp of sexual maturity. Whole blood samples were frozen at −20 °C, and tissue samples were stored in lysis buffer at room temperature until DNA was extracted for long-term storage at 4 °C.
Genetic analysis
Blood clot samples (∼0.5 g) were digested by rotating for 12 h at 55 °C in 750 μl of lysis buffer (100 mM Tris-Cl pH 8, 10 mM EDTA, 1% SDS, ddH2O), 40 μl of proteinase K (10 mg ml−1) and 2 μl of streptokinase (10 U μl−1). Tissue samples (∼5 × 5 mm) were digested by rotating for 24 h at 55 °C in 750 μl of lysis buffer and 20 μl of proteinase K (10 mg ml−1). Genomic DNA was extracted from blood using a standard phenol–chloroform method, and from tissue samples using either a phenol–chloroform–isopropanol method or ammonium acetate method (Sambrook and Russell, 2001). All DNA precipitations were washed twice in 70% ethanol, and DNA pellets were resuspended in 250 μl of TLE (10 mM Tris-Cl, 0.1 mM EDTA). A 449 bp region between sites 15 390 and 15 900 of the collared peccary mtDNA D-loop was amplified from genomic DNA using porcine primers (Alves et al., 2003). This sequence lies in the hypervariable 5′ end of the mitochondrial control region and does not code for any known protein product. PCR volumes were 25 μl and contained final concentrations of the following reagents: 1.5 mM MgCl2; 0.5 μM each primer; 0.21 mM dNTPs; 1.25 U Taq polymerase. PCRs were performed in an Eppendorf MasterCycler using the following temperature profile: denaturation for 3 min. at 94 °C, followed by 30 cycles of 94 °C for 4 s, 55 °C for 4 s, and 72 °C for 12 s; finishing with a 15 min extension step at 72° C. PCR products were cleaned using a low sodium protocol; 28 μl of a mixture containing 500 ml of absolute ethanol and 20 μl of 3M NaOAc (pH 5.2) was added to each sample, shaken for 15 min, and centrifuged at 2051 g for 35 min. This step was followed by 70% ethanol precipitation under centrifugation (twice) and resuspension in 20 μl ddH20.
PCR products were then directly sequenced in both directions using Big Dye 3.1 chemistry. Sequencing products were purified using the low sodium protocol described above, and then electrophoresed using an AB Prism 3730XL sequencer (Applied Biosystems, Foster City, CA, USA). Sequence data were aligned and edited with Sequencher 4.5 (gene codes). Nuclear copies of mtDNA genes (numts) can greatly confound evolutionary analyses, and we avoided numts using methods described in Triant and DeWoody (2007). For example, a few individuals (<5%), harbored apparently heterozygous sites so we reamplified their DNA and completely resequenced the amplicons in both directions. In every case, this procedure completely resolved the mismatch and suggested the initial discrepancy was probably a result of Taq error.
We converted sequences into NEXUS format and imported them into PAUP* 4.0 (Swofford, 2003) for haplotype assignment. Haplotypes were determined through reconstruction of unrooted phylogenetic trees using a neighbor-joining algorithm. Direct sequencing of a sub-set of the CWMA population revealed that some of the mtDNA haplotypes could be discriminated by restriction digest with the MboI enzyme, but all individuals from WWR and BB were typed by direct sequencing.
Statistical analyses
Among-population differentiation
MtDNA haplotype frequencies were calculated by hand for all three populations. Genetic differentiation among populations was inferred from FST estimates (Weir and Cockerham, 1984) and exact tests of population differentiation (Raymond and Rousset, 1995) using the software package Arlequin Version 3.1 (Excoffier et al., 2005). For the latter, P-values were estimated from a Markov chain set to 110 000 steps including 10 000 dememorization steps. All analyses were based on pure haplotype frequency data rather than nucleotide differences.
Within-population differentiation
Because P. tajacu populations are subdivided into breeding groups, we incorporated breeding group as a hierarchical level. We calculated identity probabilities by simple counting of identical pairs of genes at different hierarchical levels (Q1 for pairs of genes within groups, Q2 for pairs of genes sampled among groups within populations, and Q3 for pairs of genes sampled in different populations). We then estimated the intra-class correlations by taking appropriate ratios of identity probabilities, weighted according to the number of pairs in each sample (see Rousset, 2007), following the definitions of F-statistics as functions of identity probabilities between pairs of genes (see Appendix). As the distances among populations are large in this study (range of 225 km–945 km among the three sampling sites), we considered the three populations as independent replicates in the analysis, and we restricted our analyses to estimate within-population dispersal. We focused on the level of genetic differentiation among social groups within populations as measured by the parameter FGP. The notation is adapted from Wright (1965). This approach is different from that of Fontanillas et al. (2004) who considered dispersal both among populations and among breeding groups. Although the samples from each site were collected in different years, FGP estimates do not depend upon identity between pairs of genes from different populations and temporally discontinuous sampling is therefore unlikely to undermine the approach. We employed a bootstrapping procedure to calculate confidence limits around estimates of FGP for each class of individuals. Using the statistical software package R (R Development Core Team, 2008), we generated 25 000 bootstrap samples, with each sample being produced by random resampling (with replacement) of the 255 nucleotide sites from the mtDNA haplotypes (254 sites+1 indel). This allowed us to calculate FGP estimates for each sample and generate a distribution; confidence intervals end points were then calculated as the 2.5% and the 97.5% percentiles of this distribution. This procedure is strictly equivalent to that implemented in the software package Arlequin Version 3.1 (Excoffier et al., 2005) to generate 95% confidence limits by bootstrapping genetic differentiation values in a locus-by-locus AMOVA (see, for example, Langergraber et al., 2007).
Class-specific analyses
Dispersal is a trait that can be partitioned into pre- and post-dispersal conditions, therefore our first analysis partitioned the data by age. We performed independent analyses on data partitioned into two age sets, respectively for adults and immatures (the latter including both juveniles and infants). Subadults were classed as immatures and then as adults in sequential analyses. Each age-specific data set was composed of individuals assigned to their respective populations and social groups, and intra-class correlations (FGP) were calculated among social groups within populations from identity probabilities of pairs of genes (see the previous section: ‘Within-population differentiation’). Only those social groups containing a representative individual from each treatment were included in the analysis (for example, in the independent analyses on adult and immature data sets, a social group must have contained at least 1 adult and 1 immature to be included). We then duplicated the analysis with the data partitioned by sex rather than age. From these results, we were able to distinguish a putative class of dispersing individuals, from a putative class of non-dispersers. We therefore performed a posteriori, independent analyses on data sets of putative dispersers and non-dispersers.
We used a resampling scheme after Goudet et al. (2002) to test whether the estimated fixation indices among social groups within replicate populations (FGP) for specific classes (age, sex or putative dispersal class) departed significantly from the null hypothesis that dispersal is independent from the class of individuals. Resampling tests were all performed with the statistical software package R (R Development Core Team, 2008). For each class, we generated 25 000 randomized datasets, by re-assigning the age (or sex, or dispersal class) of each haplotype randomly within each breeding group. By doing so, we kept the number of individuals from each class constant within each breeding group. We calculated the probabilities of identities between pairs of genes for each resampled dataset, and obtained the distribution of class-specific FGP estimates under the null hypothesis that dispersal behavior or capability is independent from age, sex, or dispersal class. We then calculated P-values as the proportion of times where FGP from the randomized data sets was larger than or equal to the observed FGP on the original dataset.
Estimating dispersal
To calculate a sex-specific dispersal rate within a single generation, we adapted the Vitalis' (2002) approach and extended it to mtDNA data. In Vitalis (2002), the ratio of the sex-specific differentiation evaluated after juvenile dispersal (F̂GPXX) divided by the differentiation evaluated before dispersal (F̂GP*) gives the sex-specific dispersal rate. Appendix 1 shows that this relationship also applies to uniparentally inherited markers, and:

gives the sex-specific dispersal rate. In this study, we use this simple model to compare fixation indices before and after dispersal at the within-population level, focusing on dispersal of individuals among breeding groups. This equation assumes that the number of breeding groups, n, is large (infinite); by considering an infinitely large n, we slightly overestimate dispersal rate mx (for example, 10% relative bias with n=10). We estimated instantaneous sex-specific dispersal rates for P. tajacu by applying equation (1), using fixation indices estimates for adult males, adult females and all immatures of both sexes (2). Confidence intervals for dispersal rates were obtained by means of a bootstrap procedure, similar to that used for FGP (see the previous section: ‘Within-population differentiation’), modified as follows. For each bootstrap sample, FGP estimates were calculated for adult males (resp. adult females) and all immatures, and male- (resp. female-) specific migration rates were calculated using equation (1). Confidence intervals for sex-specific dispersal rates were then derived from the 0.025 and 0.975 percentiles of the bootstrap distribution.
Results
mtDNA haplotype distribution patterns
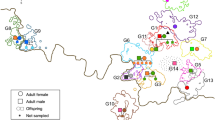
A total of 18 nucleotide sites were variable (17 substitutions and a single indel) over 449 bp. We recovered six mtDNA haplotypes from 267 individual collared peccaries among the three sites sampled (Table 1). Haplotype A was observed in all sampling sites, but haplotype B was unique to the CWMA, and haplotype C was found in both the WWR and the CWMA. The BB population was almost fixed for haplotype E (96%). Haplotypes F and G were only found in the CWMA, and were represented by single individuals (both males).
We overlaid mtDNA haplotype distribution onto the social group territory distribution for all populations. At the local level, haplotype distribution did not show geographic structuring in the CWMA or the WWR; all haplotypes present at each sampling site were found distributed throughout that site. In the BB population, haplotype A was found only in the eastern portion of the sampling site. At the regional level across Texas, we observed significant population differentiation. Pairwise FST estimates ranged from 0.31 to 0.86 between populations and pairwise exact tests of population differentiation were highly significant (P=0.001), indicating that these populations are significantly divergent from one another.
Patterns of genetic variation revealed by F-statistics as functions of identity probabilities
Because dispersal status is often dependent upon age, we tested for an age bias in dispersal. To that end, we pooled infants and juveniles (categorized hereafter as ‘immatures’) in one class, and adults in another class. It was not clear if individuals categorized as subadults were sufficiently developed to be considered as adults, therefore we performed a preliminary analysis on adult-only and immature-only data sets partitioned into social groups, which revealed a decrease in FGP when subadults were included in the adult class (not shown). This result indicates that individual genetic variation in the subadult class is apportioned among rather than within social groups, and therefore subadults were classed as adults in all subsequent analyses. We estimated fixation indices among social groups for each sex, with individuals partitioned into known breeding groups (Table 2). It is clear that FGP for adults (0.30 [0.03, 0.38]) is much smaller than that for immatures (0.60 [0.60, 1.00]), as would be expected if the adult class included dispersed individuals. To test for significance of these quantitative differences, we used a randomization approach, and generated randomized data sets by assigning an age randomly to each mtDNA haplotype. Under the null hypothesis that dispersal is not age-biased, we expect the observed FGP of adults and immatures not to depart significantly from the null distribution. For adults, there was a large proportion of randomized data sets with a differentiation among groups within populations (FGP) larger than the observed, although this proportion did not achieve significance (P=0.79; Figure 1a). In contrast, for immatures of both sexes, there was only a small proportion of randomized data sets giving an FGP larger than the observed, although the test was not significant (P=0.20; Figure 1b). In general terms, these results clearly indicate a greater amount of dispersal among social groups for adults when compared to immatures.
Re-sampled data null distributions for each class of individuals: (a) adults, (b) immatures, (c) males, (d) females, (e) dispersers, (f) non-dispersers. Observed FGP for each analysis represented by hatched vertical line. Significance tested over 25 000 permutations. Histogram class heights are represented as black dots, and the smoothed density was obtained using the Average Shifted Histogram (ASH) algorithm (Scott, 1992) with smoothing parameter m=20.
To test for a signal of sex-biased dispersal, intra-class correlations (FGP) were estimated for each sex with individuals partitioned into known breeding groups (Table 2). It can be seen from these results that FGP among social groups is much smaller for males (0.23 (0.09, 0.36)) than it is for females (0.90 (0.87, 1.00)), which indicates that even when pre-dispersal age individuals are included in the male class the sex difference is still apparent. To test the significance of the sex difference, we used a randomization approach identical to the one described for age bias, and generated randomized data sets by assigning a sex randomly to each mtDNA haplotype. For males, there was a very large proportion of randomized data sets with a larger FGP than the observed (P=0.98; Figure 1c). In contrast, for females, there was only a very small proportion of randomized data sets giving a FGP larger than the observed, and the test was therefore highly significant (P<0.001; Figure 1d). These results suggest that dispersal is strongly male-biased in P. tajacu.
The inferred dispersal pattern of P. tajacu being of adult male dispersal, we conducted a further analysis, a posteriori, on data partitioned by putative dispersal condition: the data were partitioned into ‘dispersers’ (adult males) and ‘philopatrics’ (immature males and all females) and separate analyses performed on individuals assigned to breeding groups. As expected, FGP for the philopatric class was much larger (0.76 (0.73, 0.99)) than was seen for adult males (0.24 (0.07, 0.36)). For adult males, there was a very large proportion of randomized data sets with a FGP larger than the observed (P=0.99; Figure 1e). In contrast, for putative non-dispersers, the test was highly significant, with very few data sets giving an FGP larger than the observed (P<0.001; Figure 1f).
Dispersal rate estimates
The instantaneous sex-specific dispersal rate among social groups within populations was estimated using equation (1). We used the FGP estimates among social groups within populations (Table 2) for adult males (dispersers) (FGP=0.24), for adult females (FGP=0.91), and for pre-dispersal individuals of both sexes (also categorized as ‘immature’; FGP=0.60). This yielded a male-specific dispersal rate estimate (m♂) of 0.37 (0.32, 0.65). Equation (1) only makes sense if there is a significant difference between FGP measured after dispersal and before dispersal. As the confidence limits of FGP for adult females ((0.88; 1.00)) and immatures ((0.60; 1.00)) largely overlap, we were unable to calculate a female-specific dispersal rate from equation (1).
Discussion
We have demonstrated that maternally inherited genes can be used to describe the contemporary dispersal patterns of males (and the overall dispersal patterns of females) within an analytical framework based on intra-class genetic correlations. This was accomplished through comparisons of age- and sex-specific intra-class correlations partitioned hierarchically within populations. A second aim was to show that instantaneous sex-specific dispersal rates could be calculated from sex-specific estimates of differentiation using single-locus haplotypic data.
Dispersal in Pecari tajacu
In our study, we quantitatively showed that dispersal in collared peccaries is strongly biased toward males, and that approximately one-third of males dispersed from their natal groups in this single generation. This is a minimum estimate, as some individuals die before or during dispersal, and the lack of mtDNA variation undoubtedly prevented our detection of dispersal between some groups. Moreover, the pronounced local genetic structure indicates that males preferentially disperse over short distances, perhaps into neighboring herds; this is congruent with trapping data (Gabor and Hellgren, 2000). The results from the age-based analysis indicate that dispersal in this species is usually accomplished by subadults (18–24 months). At this age, they have not reached their full body mass and may be forced out by larger, resident males.
Measuring dispersal biases
Our approach allowed us to organize data into age classes, sex classes, social groups and populations and then test hypotheses about the dispersal rate of each class. For example, by performing separate analyses on sex-specific data sets, we were able to both detect a sex-bias in dispersal and also determine, which sex contributed to the pattern. Because the method relies on contrasts of sex-specific estimates of population differentiation, rather than absolutes, the power to detect differences among hierarchies is limited only by the intensity of the bias (Vitalis, 2002). In this study, there was sufficient contrast between pre- and post-dispersal age classes in males to provide a direct estimate of the instantaneous dispersal rate.
The method presented here should be applicable to any species in which there is a bias in dispersal, whether that bias is conditional on sex, age or some other phenotype, so long as trait variation can be readily distinguished and assigned to different hierarchical levels. This approach does not impose spatial distance (or a distance proxy) onto the analysis, as is seen in other approaches such as spatial autocorrelation (Smouse and Peakall, 1999). Such approaches force investigators to make assumptions about how distance interacts with the social organization when it may be inappropriate or irrelevant (for example, when sampling a highly mobile species, or at a scale where an individual is equally likely to disperse to any location under consideration). Our approach removes metric distance and location from the equation, and shifts the focus onto how the genetic variation is distributed across space irrespective of distance, which is especially useful for addressing questions of how sociality influences dispersal.
Measuring sex-biased dispersal with uni-parentally inherited markers
The approach discussed herein relies upon contrasts: we compared the genetic structure of the pre-dispersal class to the sex-specific genetic structure of the post-dispersal class to estimate instantaneous dispersal within a single generation (Lawson-Handley and Perrin, 2007). When autosomal markers are used, the expectation is that genetic structure will be more apparent in the pre-dispersal class compared with the post-dispersal class as a whole, and even more apparent in the non-dispersing sex (whichever sex it may be). When a uniparentally inherited marker is used the expectation is similar, but not identical, to what is seen for biparentally inherited markers. For instance, under a system of male-biased dispersal mtDNA haplotypes are carried within males into breeding groups, but males do not contribute mtDNA to the subsequent generation and thus the contrast between pre-dispersal individuals and adult males is substantial. However, under a system of female-biased dispersal, haplotypes would be re-distributed within and among populations in each generation. Thus a contrast between genetic differentiation for pre- and post-dispersal individuals would be difficult to detect. As a result, this approach is most useful for deriving instantaneous sex-specific dispersal rates with mtDNA data under a system of male-biased dispersal, or a double uniparental system of mitochondrial DNA inheritance (for example, Mytilus mussels). Here, we use mtDNA haplotypes as a tag, but any physical or genetic tag that could be identified in males and females before and after dispersal may have the same role as mtDNA markers in this context.
We have shown that mtDNA can be used in isolation to estimate sex-specific dispersal in the current generation. The main caveat is that mtDNA is, in effect, a single genetic marker that might be biased by selection (Bazin et al., 2006). Yet, because we based our analyses upon differences of variation in male and female within a single generation, it is difficult to imagine a pattern of selection that would undermine the approach.
References
Alves E, Ovilo C, Rodriguez MC, Silio L (2003). Mitochondrial DNA sequence variation and phylogenetic relationships among Iberian pigs and other domestic and wild pig populations. Anim Genet 34: 319–324.
Bazin E, Glemin S, Galtier N (2006). Population size does not influence mitochondrial genetic diversity in animals. Science 312: 570–572.
Bissonnette JA (1982). Social behavior and ecology of the collared peccary in Big Bend National Park. National Park Scientific Monograph Series No. 16. US Dept of the Interior: Washington, DC.
Chesser RK (1991). Gene diversity and female philopatry. Genetics 127: 437–447.
Chesser RK, Baker RJ (1996). Effective sizes and dynamics of uniparentally and diparentally inherited genes. Genetics 144: 1225–1235.
Cockerham CC, Weir BDS (1987). Correlations, descent measures: drift with migration and mutation. Proc Natl Acad Sci USA 84: 8512–8514.
Day GI (1985). Javelina Research and Management in Arizona. Arizona Game and Fish Department: Phoenix.
Ellisor JE, Harwell WF (1969). Mobility and home range of collared peccary in southern Texas. J Wildl Manage 33: 425–427.
Escorza-Trevino S, Dizon AE (2000). Phylogeography, intraspecific structure and sex-biased dispersal of Dall's porpoise, Phocoenoides dalli, revealed by mitochondrial and microsatellite DNA analyses. Mol Ecol 9: 1049–1060.
Excoffier L, Laval G, Schneider S (2005). Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol Bioinform Online 1: 47–50.
Fontanillas P, Petit E, Perrin N (2004). Estimating sex-specific dispersal rates with autosomal markers in hierarchically structured populations. Evolution 58: 886–894.
Gabor TM, Hellgren EC (2000). Variation in peccary populations: landscape composition or competition by an invader? Ecology 81: 2509–2525.
Girman DJ, Mills MGL, Geffen E, Wayne R (1997). A molecular genetic analysis of social structure, dispersal and interpack relationships of the African wild dog (Lycaon pictus). Behav Ecol Sociobiol 40: 187–198.
Gongora J, Morales S, Bernal JE, Moran C (2006). Phylogenetic divisions among Collared peccaries (Pecari tajacu) detected using mitochondrial and nuclear sequences. Mol Phylogenet Evol 41: 1–11.
Goudet J, Perrin N, Waser PM (2002). Tests for sex-biased dispersal using bi-parentally inherited genetic markers. Mol Ecol 11: 1103–1114.
Greenwood PJ (1980). Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28: 1140–1162.
Hellgren EC, Lochmiller RL, Grant WE (1984). Demographic, morphologic, and reproductive status of a herd of collared peccaries (Tayassu tajacu) in south Texas. Am Midl Nat 112: 402–407.
Hoelzer GA, Dittus WPJ, Ashley MV, Melnicks DJ (1994). The local distribution of highly divergent mitochondrial DNA haplotypes in toque macaques Macaca sinica at Polonnaruwa, Sri Lanka. Mol Ecol 3: 451–458.
Langergraber KE, Siedel H, Mitani JC, Wrangham RW, Reynolds V et al. (2007). The genetic signature of sex-biased migration in patrilocal chimpanzees and humans. PLoS ONE 2: e973. doi:10.1371/journal.pone.0000973.
Lawson-Handley LJ, Perrin N (2007). Advances in our understanding of mammalian sex-biased dispersal. Mol Ecol 16: 1559–1578.
Mossman CA, Waser PM (1999). Genetic detection of sex-biased dispersal. Mol Ecol 8: 1063–1067.
Petit E, Balloux F, Goudet J (2001). Sex-biased dispersal in a migratory bat: a characterization using sex-specific demographic parameters. Evolution 55: 635–640.
Prugnolle F, de Meeus T (2002). Inferring sex-biased dispersal from population genetic tools: a review. Heredity 88: 161–165.
R Development Core Team (2008). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. ISBN 3-900051-07-0, URL http://R-project.org.
Raymond M, Rousset F (1995). An exact test for population differentiation. Evolution 49: 1283–1286.
Rousset F (1996). Equilibrium values of measures of population subdivision for stepwise mutation processes. Genetics 142: 1357–1362.
Rousset F (2007). Inferences from spatial population genetics. In: Balding D, Bishop M, Cannings C (eds) Handbook of Statistical Genetics 3rd edn. John Wiley & Sons: Chichester. pp 945–979.
Sambrook J, Russell DW (2001). Molecular Cloning: A Laboratory Manual, 3rd edn. Cold Spring Harbor Laboratory Press: New York.
Scott DW (1992). Multivariate Density Estimation: Theory, Practice, and Visualization. John Wiley: New York.
Slatkin M (1991). Inbreeding coefficients and coalescence times. Genet Res 58: 167–175.
Slatkin M, Voelm L (1991). FST in a hierarchical island model. Genetics 127: 627–629.
Smouse PE, Peakall R (1999). Spatial autocorrelation analysis of multi-allele and multi-locus genetic microstructure. Heredity 82: 561–573.
Sowls LK (1978). Collared peccary. In: Schmidt JL, Gilbert DL (eds) Big game of North America. Stackpole Books: Harrisburg, Pennsylvania. pp 191–205.
Sugg DW, Chesser RK (1994). Effective population sizes with multiple paternity. Genetics 137: 1147–1155.
Swofford DL (2003). PAUP*: Phylogenetic Analysis Using Parsimony (* and Other Methods), Version 4.0b 10. Sinauer Associates: Sunderland, Massachusetts.
Theimer TC, Keim P (1994). Geographic patterns of mitochondrial-DNA variation in collared peccaries. J Mammal 75: 121–128.
Triant DA, DeWoody JA (2007). The occurrence, detection, and avoidance of mitochondrial DNA translocations in mammalian systematic and phylogeography. J Mammal 88: 908–920.
Vigouroux V, Couvet D (2000). The hierarchical island model revisited. Genet Sel Evol 32: 395–402.
Vitalis R (2002). Sex-specific genetic differentiation and coalescence times: estimating sex-biased dispersal rates. Mol Ecol 11: 125–138.
Waser PM, Strobeck C, Paetkau D (2001). Estimating interpopulation dispersal rates. In: Gittleman JL, Funk SM, MacDonald DW, Wayne RK (eds) Arnivore Conservation. Cambridge University Press, Cambridge. pp 484–497.
Weir B, Cockerham C (1984). Estimating F-Statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wright S (1951). The genetical structure of populations. Ann Eugen 15: 323–354.
Wright S (1965). The interpretation of population structure by F-statistics with special regard to systems of mating. Evolution 19: 395–420.
Yang G, Lui S, Ren WH, Zhou KY, Wei FW (2003). Mitochondrial control region variability of baiji and the Yangtze finless porpoises, two sympatric small cetaceans in the Yangtze river. Acta Theriologica 48: 469–483.
Acknowledgements
This research project was supported by a Grant in Aid of Research from the American Society of Mammalogists to JD Cooper and by Purdue University funding to JA DeWoody. JD Cooper is a Welder Wildlife Refuge Fellow, and many thanks are offered to the faculty and staff at the Welder for their encouragement and assistance. We also thank Jim Cathey and Texas Parks and Wildlife Department personnel for their help and cooperation during sample collection. Special thanks goes to volunteers Edmund Harriss, Rachel Bonner and Raul Martinez for their dedication and hard work in trapping collared peccaries. Finally, we thank the members of the DeWoody lab and Peter Smouse for their insightful comments during the preparation of this article. Part of this work was carried out by using the resources of the Computational Biology Service Unit from the Museum National d'Histoire Naturelle (MNHN—CNRS UMS 2700), which was partially funded by Saint Gobain.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Sex-specific differentiation before and after (instantaneous) dispersal
F-statistics are defined as intra-class correlations for the probability of identity in state (IIS correlations) (Cockerham and Weir, 1987; Rousset, 1996). Yet, the infinite allele model (IAM) provides the value of the probability of identity by descent (IBD probabilities) and in the low mutation rate limit, for two given classes, IIS and IBD correlations converge to the same value (Slatkin, 1991; Rousset, 1996). Thus, the properties of F-statistics can be deduced from the properties of intra-class correlations for IBD probabilities (Rousset, 1996).
In the following, we consider mitochondrial DNA (mtDNA) markers, that is, haploid markers, transmitted by females only. Let Q1XY, Q2XY and Q3XY be the IBD probability of two mtDNA gene copies sampled from two individuals of sex X and Y after dispersal among individuals within the same breeding group, among breeding groups within the same populations and among distinct populations, respectively. These individuals may be two males, two females, or one male and one female. Let Q1*, Q2* and Q3* be the corresponding IBD probabilities for gene copies sampled before dispersal. For pairs of genes sampled before dispersal there is no need to consider distinct coefficients for different pairs of individuals of the same or opposite sex (Vitalis, 2002).
Let us consider an infinite island model of population structure (Wright, 1951) in which each population is isolated and further subdivided into n breeding groups, and dispersal among breeding groups is achieved by juveniles, before reproduction. Here, because we consider that populations are independent, we restrict our analyses to estimate within-population dispersal only. We therefore focus on the IBD probabilities among individuals within social groups (Q1XY and Q1*), and among social groups within populations (Q2XY and Q2*), and further consider the IBD probabilities between pairs of genes from distinct populations (Q3XY and Q3*) to be nil in the model. A migrant individual in a breeding group is equally likely to come from any of the (n−1) other breeding groups. Generations do not overlap. Let mX denote the probability that an individual of sex X has immigrated. Each generation, after migration, the frequency of pairs of individuals taken at random in one breeding group that come from a single group before migration is aXY=(1−mX)(1−mY)+mX mY/(n−1), for pairs of individuals of sex X and Y with X∈{♂,♀} and Y∈{♂,♀}. Conversely, the frequency of pairs of individuals taken at random from two breeding groups after migration that originate from the same group before migration is bXY=(1−aXY)/(n−1) (see, for example, Rousset, 1996). We assume that the mutation rate is low (that is, μ<<mX), so that virtually no mutation arises over a single generation.
The genes sampled among individuals after dispersal in a breeding group at any generation come from the same breeding group before dispersal with probability aXY and from different breeding groups with probability (1−aXY). These mtDNA gene copies are then IBD with probabilities Q1* and Q2*, respectively. Thus,

Similarly, for genes sampled before dispersal,

We can rewrite equations (A.1) and (A.2) as:

and:

Subtracting equations (A.3) and (A.4), we obtain:

where dXY=(aXY−bXY). Rearranging equation (A.4) gives:

Taking the ratio of equation (A.6) over (A.5) gives:

As we consider independent populations, the relevant parameter to infer dispersal among breeding groups is FGP. From the definition of sex-specific F-statistics as appropriate ratios of sex-specific IBD probabilities (Vitalis, 2002), we get:

before dispersal, and:

after dispersal. We get from equation (A.7):

that is, multiplying both sides by (dXY × FGPXY) and rearranging:

which is equation 15 in Vitalis (2002) except that here it holds for mtDNA markers, and it does not assume equilibrium. This relation is valid for any generation, for samples taken before and after dispersal. The last term in the right-hand side of equation (A.11) is negligible compared with dXY. Thus, FGPXY differs from FGP* by a factor dXX=(1−mX[n/(n−1)])2≈(1−mX)2, for large n. Therefore, taking the ratio of sex-specific FGPXY evaluated after juvenile dispersal, over FGP* evaluated before dispersal gives the sex-specific migration rate:

Following Rousset (2007), we estimated the intra-class correlations by taking appropriate ratios of identity probabilities, weighted according to the number of pairs in each sample. From the definitions of F-statistics as functions of identity probabilities between pairs of genes (equations (A.8) and (A.9)), we get:

and

where Q̂i* and Q̂iXY denote the estimates of identity probabilities between pairs of genes, calculated by simple counting of identical pairs of genes at the ith hierarchical level. An estimate of the sex-specific migration rate therefore reads:

Rights and permissions
About this article
Cite this article
Cooper, J., Vitalis, R., Waser, P. et al. Quantifying male-biased dispersal among social groups in the collared peccary (Pecari tajacu) using analyses based on mtDNA variation. Heredity 104, 79–87 (2010). https://doi.org/10.1038/hdy.2009.102
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.102
Keywords
This article is cited by
-
Spatial patterns of the first groups of collared peccaries (Pecari tajacu) reintroduced in South America
Tropical Ecology (2020)
-
Mitochondrial DNA paradox: sex-specific genetic structure in a marine mussel – despite maternal inheritance and passive dispersal
BMC Genetics (2012)
-
The spatial distribution does not affect host–parasite coevolution in Rossomyrmex ants
Insectes Sociaux (2012)
-
Is sexual monomorphism a predictor of polygynandry? Evidence from a social mammal, the collared peccary
Behavioral Ecology and Sociobiology (2011)