Abstract
Purpose: We evaluated the experiences of 19 Italian laboratories concerning 241 small supernumerary marker chromosomes (sSMCs) with the aim of answering questions arising from their origin from any chromosome, their variable size and genetic content, and their impact on the carrier's phenotype.
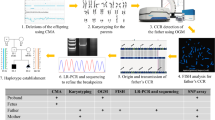
Methods: Conventional protocols were used to set up the cultures and chromosome preparations. Both commercial and homemade probes were used for the fluorescent in situ hybridization analyses.
Results: A total of 113 of the 241 sSMCs were detected antenatally, and 128 were detected postnatally. There were 52 inherited and 172 de novo cases. Abnormal phenotype was present in 137 cases (57%), 38 of which were antenatally diagnosed. A mosaic condition was observed in 87 cases (36%). In terms of morphology, monocentric and dicentric bisatellited marker chromosomes were the most common, followed by monocentric rings and short-arm isochromosomes. The chromosomes generating the sSMCs were acrocentric in 132 cases (69%) and non-acrocentric chromosomes in 60 cases (31%); a neocentromere was hypothesized in three cases involving chromosomes 6, 8, and 15.
Conclusion: The presented and published data still do not allow any definite conclusions to be drawn concerning karyotype–phenotype correlations. Only concerted efforts to characterize molecularly the sSMCs associated or not with a clinical phenotype can yield results suitable for addressing karyotype–phenotype correlations in support of genetic counseling.
Similar content being viewed by others
Main
The widespread use of molecular cytogenetic techniques in diagnostic laboratories has improved diagnostic quality, especially in prenatal cases. However, one of the few major problems remaining is the identification of the nature and origin of small supernumerary marker chromosomes (sSMCs).
sSMCs display a wide range of morphology and occur at highly variable incidence,1,2 thus giving rise to considerable problems in genetic counseling, particularly during prenatal testing. Only the combined use of conventional and molecular cytogenetics can define the origin and genetic content of sSMCs. However, the rarity of some sSMCs and the lack of comparative fluorescent in situ hybridization (FISH) characterization of those sharing the same chromosomal origin still hinder the genotype–phenotype correlations and appropriate genetic counseling. We evaluate the experiences of 19 Italian laboratories concerning 241 sSMCs with the aim of answering questions such as the following: Is FISH feasible and informative in the short times required for prenatal diagnosis? Should panels of genomic clones covering the pericentromeric regions of all chromosomes be set up for this purpose? How far can FISH take us? When is a uniparental disomy (UPD) test needed? What recommendations should be given for genetic counseling? And should they be generic or specifically designed on the basis of the type of sSMC?
MATERIALS AND METHODS
The analyses were performed in accordance with the Italian guidelines,3 which are consistent with the European guidelines4 (see Appendix 2). The analyzed tissues were peripheral and cord blood, amniotic fluid, and chorionic villi. Conventional protocols were used to set up the cultures and chromosome preparations. The applied banding techniques were GTG, QFQ, CBG, DA-DAPI, and AgNOR. The karyotypes were formulated following the International System for Human Cytogenetic Nomenclature 1995 indications.5 Both commercial and homemade probes were used for the FISH analyses. The commercial probes were specific α-satellite DNA of centromeres and chromosome libraries. The FISH experiments were performed according to the suppliers' protocols. Twelve previously published cases were characterized by means of YACs, BACs, and PACs, using previously described FISH protocols.6–14
UPD tests for chromosomes 2, 7, 8, 13, 14, 15, and 22 were used in only nine cases. The tests were performed by means of the dinucleotide microsatellite segregation study of fetal and parental DNAs.
RESULTS
Ascertainment
The 241 sSMCs described here are the fruit of 10 years of activity in 19 Italian laboratories; 113 (43%) were antenatally ascertained, and 128 were postnatal diagnoses (Fig. 1, first column). The indications in the prenatal diagnoses are listed in Table 1.
Parental inheritance and mosaicism
Information about familial origin was achieved for 224 of the 241 sSMCs, of which 172 (77%) were de novo, 35 (16%) were maternally inherited, and 17 (7%) were paternally inherited. In five cases (9.6%), the contributing parent showed a mosaic karyotype for the marker chromosome, and the probands also carried the sSMC in a mosaic status. Seventy-nine of the de novo cases (46%) were prenatal ascertainments, and 93 (54%) were identified in the postnatal period (Fig. 1, second column). Thirty-four of the inherited sSMCs (65%) were detected during prenatal diagnoses, and 18 (35%) were postnatal ascertainments (Fig. 1, third and fourth columns). Mosaicism was observed in 87 (36%) of the 241 sSMCs, with the abnormal cell line ranging from 3% to 5% to 60% to 70%. Information about familial occurrence was obtained for 82 (94%) of these cases, 9 (11%) of which were found to be inherited. Information about familial occurrence was obtained in 146 of the 154 nonmosaic sSMCs, 43 (29%) of which were inherited.
Chromosomal origin
On the basis of banding techniques, 231 of 241 sSMCs could be morphologically subgrouped as shown in Figure 2. FISH analyses successfully identified 192 (80%) of the 241 sSMCs. The sSMCs derived from chromosome 15 accounted for 59% of the acrocentric sSMCs derived and 40% of all of the FISH characterized sSMCs.
The combined morphologic and FISH results allowed us to establish that 54% of the 241 sSMCs derived from acrocentric chromosomes, and the remaining 46% derived from nonacrocentric chromosomes. The chromosomal origin of 29 i(p), representing 12% of all sSMCs, was non-acrocentric in 25 cases (8 deriving from chromosome 12, 15 deriving from chromosome 18, 2 deriving from chromosome 20, 3 deriving from chromosome 15, and 1 deriving from chromosome 2212), accounting for a 10% frequency. The chromosomal origins of 33 rings from non-acrocentric chromosomes were also identified, that is, chromosomes X (n = 4), 2 (n = 4), 3 (n = 4), 8 (n = 4), 5 (n = 3), 16 (n = 3), 1 (n = 2), 7 (n = 2), 6 (n = 1), 12 (n = 1), 19 (n = 1), and 1/5/19 (n = 4). Three cases, classified as acentric fragments, were negative after hybridization of α-satellite probes for all centromeres, that is, the common or specific α-satellite DNAs and positive with chromosome libraries of chromosomes 6, 8, and 15. The chromosome 6-derived sSMC was antenatally detected, whereas the other two were postnatally detected. Multiple sSMCs were found in two prenatal diagnoses. In the first, two bisatellited and apparently monocentric but morphologically different sSMCs were present in all of the analyzed amniotic fluid cells (one maternally derived and the second de novo). The woman decided to interrupt the pregnancy and refused any other investigation. In the second case, amniotic fluid culture showed three cell lines: one with a normal karyotype, the second with a psu dic(15)(q13), and a third with a very small ring originating from chromosome 15: ish r(15)(p11.2q11.2)(D15Z1+;D15S11−). The woman decided to interrupt the pregnancy because of the adverse prognosis.
Phenotype
Information concerning the patient's phenotype was obtained in 224 cases, and the relative results are summarized in Table 2.
An abnormal phenotype was observed in 137 of the 224 cases (61%), 38 (17%) of which were antenatally investigated and 99 (44%) of which were postnatally ascertained. Among the postnatal cases the abnormal phenotypes [with the exclusion of the classical syndromes associated with psu dic(15), psu dic(22), i(18)(p), i(12p)] ranged from a generic psychomotor/mild mental retardation with or without dysmorphisms observed in the youngest probands to infertility ascertained by reproductive failure in the eldest patients. The abnormal phenotypes of the prenatal cases were mainly malformations or intrauterine growth retardation.
The abnormal phenotypes were associated with 37 (27%) of the prenatal and 84 (61%) of the postnatal de novo sSMCs, and 15 (11%) of the postnatal and one (<1%) of the prenatal inherited sSMCs. Of the prenatal cases, the sSMCs associated with an abnormal phenotype were, in order of decreasing incidence: psu dic(15) (q13), psu dic(22) (q11), and rings from nonacrocentric autosomes. Table 2 also shows that 83% of the postnatal cases with inherited sSMCs (15/18) were associated with clinical manifestations, compared with only one (3%) of the 34 cases detected antenatally. Conversely, 87 patients (39%) had a normal phenotype, of which 75 (86%) were antenatally investigated and 12 (14%) were postnatally ascertained. Moreover, the normal phenotypes were associated with 42 prenatal (48%) and 9 postnatal (10%) de novo sSMCs and with 33 prenatal (38%) and 3 postnatal (3%) inherited sSMCs. The prevalent “de novo” sSMCs morphology among the 42 normal fetuses was monocentric bisatellited. The karyotype–phenotype correlation in the 192 cases with FISH-characterized sSMCs showed that the abnormal phenotypes ranged from a generic psychomotor/mild mental retardation to classic syndromes such as Prader-Willi (because of a concomitant mat 15UPD),8 cat-eye, i(18) (p), and so forth. As can be deduced from Figure 3A, the overall ratio of normal to abnormal phenotypes was 1:2 (66 vs. 126), with a 1:3.2 ratio (14 vs. 46) in the case of sSMCs originating from non-acrocentric chromosomes. The ratio of acrocentric-derived sSMCs seems to be more balanced (52 normal vs. 80 abnormal: a ratio of 1:1.5). The phenotypical abnormalities in this last group were prevalently caused by psudic(15) (58/80) and psudic(22) (10/80). Figures 3B and C, respectively, summarize the chromosomal origin and associated phenotypes of the 60 nonacrocentric (31%) and the 132 acrocentric-derived sSMCs (69%).
The UPD test was performed in nine cases with sSMCs (excluding the published cases)8,11–14 derived from chromosomes 2, 7, 8, 13 to 21, 14 to 22, 15, and 22, but the results were always negative.
DISCUSSION
sSMCs are a particular group of aberrant chromosomes that originate as a result of numeric errors and/or structural rearrangements. On the basis of previous surveys, their frequency in the general population can be estimated as ranging from 0.18 in 1000 to 1.5 in 1000.1,2
They are morphologically highly heterogeneous, and it is known that they may derive from all chromosomes and that a few show neocentromeres.16 It is generally held that, although limited, conventional cytogenetic studies are necessary to plan subsequent FISH analyses (e.g., the choice of probes and the number of cells to analyze) and evaluate the results correctly. FISH is and will remain an obligatory technique because it makes it possible to determine the origin and genetic content of sSMCs, which is a premise for establishing the genotype–phenotype correlations. This latter aspect plays an essential role in prenatal diagnoses.
Some sSMCs, such as psu dic(15), psu dic(22), i(18) (p), and so forth, have been described as being responsible for well-defined clinical pictures, whereas 60% of all known sSMCs are associated with clinical signs but do not cause any precise syndromic pictures.17–19 The use of PACs/BACs and the evolution of FISH techniques (reverse painting and CGH), CGH microarray, and so forth have made it possible to “size” sSMCs and define their content very finely. However, the literature contains few descriptions of sSMCs grouped by marker type from which karyotype–phenotype correlations can be deduced to allow correct genetic counseling. Starke et al.20 provided a first step toward the identification of pericentromeric disease-related genes by showing that only a few proximal trisomies underlie clinical manifestations. The phenotypic variability associated with sSMCs originating from the same chromosome may reflect the degree of mosaicism,17,19 the UPD status of the originating chromosome pair, the DNA sequence content (repeated vs. coding DNA), and the size in terms of the multiplicities of genes and/or imprinted regions involved. As in the case of contiguous gene syndromes, in which one or a few genes may be responsible for disease,21–23 the presence or absence of a specific genomic fragment can determine a normal/mild/severe phenotype, even if the sSMCs comes from the same chromosome.
The data described in this article are from 10 years of diagnoses made in 19 Italian laboratories. As can be seen in Figure 1, the sample is equally divided between antenatal (43%) and postnatal (57%) diagnoses. Liehr et al.24 reported 61% de novo sSMCs when reviewing 174 cases from several studies, and Crolla et al.25 reported 70% in 109 cases. Our results support the high prevalence of de novo sSMCs, 77%, of which 46% were antenatally diagnosed and 54% postnatally. Moreover, 65% of the familial cases in our series were prenatally ascertained (Fig. 1). The 2:1 ratio of maternally/paternally derived sSMCs is very similar to the 1,75:1 observed by Crolla et al.25
A mosaic karyotype was observed in 36% of our cases (87/241), which is lower than the 54% (78/144) and 60% (81/137) reported by Crolla et al.25,26 and the 60% (21/35) reported by Starke et al.20 This difference can be attributed to the differences in sample sizes and the relative proportions of sSMCs with a given chromosomal origin. As underlined by Crolla et al.25 there are striking differences in the distribution of mosaicism when non-acrocentric and acrocentric-derived sSMCs are compared. In our sample, sSMC(15)s accounted for 40% of all of the FISH characterized sSMCs.
In line with their known morphologic heterogeneity (Fig. 2), the major classes of our sSMCs were pseudodicentric marker chromosomes (36%), monocentric rings (22%), and monocentric bisatellited (21%) and nonsatellited iso(p) chromosomes (12%). On the basis of their morphologic appearance (satellited vs. nonsatellited), 54% of the sSMCs originated from acrocentric chromosomes. By FISH analyses we identified the chromosomal origin of 80% of the sSMCs and established that 68% originated from acrocentric chromosomes (Fig. 3B and C), a percentage that is similar to that observed by Crolla et al.25 Among the sSMCs originated from acrocentric chromosomes, the most predominant (59%) was the so-called inv dup(15), thus confirming data available on the http://mti-n.mti.uni-jena.de/huwww/MOL_ZYTO/sSMC.htm website including published and unpublished sSMCs cases. The sSMCs derived from nonacrocentric chromosomes were rings (55%) and iso(p) (42%). Three of our cases were classified as analphoid sSMCs, because they were negative after hybridization with α-satellite probes for all centromeres: They were found to originate from chromosomes 6, 8, and 15. In their review, Liehr et al.24 reported that 45% of their analphoid sSMCs derived from chromosomes 13, 3, 8, and 1, and 25% from chromosome 15.
We identified only two cases with multiple sSMCs, both of which were antenatally diagnosed. In the first case, the sSMCs were present in different cell lines and, although both derived from chromosome 15, they were morphologically different. In the second case, the sSMCs were morphologically different but simultaneously present in a homogeneous karyotype. The fact that multiple sSMCs can originate from different chromosomes induced Daniel and Malafiej19 to propose a stimulating causative hypothesis based on the presence of a superfluous pronucleus at fertilization, the incomplete digestion of which leads to minichromosomes persisting in the cytoplasm and then being absorbed in the nuclei of blastomeres. This hypothesis may explain some findings in two to eight cell human embryos,27 showing that chromatin desegregation and altered mitotic spindle are not so rare in vitro.
Phenotype–karyotype correlations in the presence of a sSMCs remain a major problem, especially in the case of prenatal diagnoses. Our results indicate 61% of pathologic phenotypes: 44% detected postnatally and 17% detected antenatally (Table 2). When an abnormal phenotype was observed, the associated sSMCs were de novo in 97% of the prenatal and 85% of the postnatal cases, but only 47% of the de novo sSMCs diagnosed prenatally had an abnormal phenotype compared with 90% of those postnatally diagnosed (Table 2). The inherited sSMCs in our sample were diagnosed twice as frequently prenatally than postnatally. This reflects the differences in the indications for prenatal and postnatal analyses, and highlight the fact that a marker chromosome can also be found in the absence of a clinical indication. Our 34 prenatally diagnosed familial sSMCs did not seem to be related to phenotypical alterations, with the exception of a single case with a 48,XXY,+mar karyotype. The same conclusion was reached in the previously reported studies, recently reviewed by Liehr et al.24 On the contrary, 15 of 18 postnatally diagnosed inherited sSMCs (83%) were associated with clinical manifestations, mainly inherent to reproductive defects.
Only two previously published studies have provided indications concerning the pathologic phenotype and the risk accordingly associated with sSMCs: Warburton2 identified 10 of 68 cases with an abnormal phenotype (14.8%), and Crolla et al.26 described 6 of 21 cases (28.6%). These two surveys are not statistically comparable, but they provide the only available data for genetic counseling. We found that 137 of 224 (61%) of our cases had anomalous phenotypes (Table 2): If only the FISH-verified cases are considered (Fig. 3A), this percentage increases to 66% (126/192), whereas, if the known syndromes [associated with psudic(15), i(12p), i(18p), and i(22)] are excluded, it decreases to 33% (33/99), a figure that is close to that of Crolla et al.26 However, these percentages do not apply when the data are analyzed on the basis of the individual type of sSMC. For example, of the 28 cases with sSMCs containing centromere 22, 15 were normal and 13 (46%) were pathologic, whereas only 3 of the 16 cases with sSMCs containing centromere 13 (19%) had an abnormal phenotype.
This calculation can be made in the case of the sSMCs deriving from acrocentric chromosomes because they are frequent in case samples, but the situation concerning rings is so fragmentary that it is impossible to obtain percentages specific for each marker. On the basis of their previous experience (five cases, all normal) and the published data (six cases) concerning rings originating from the pericentromeric region of chromosome 3, Anderlid et al.17 concluded that r(3) has little or no effect on the carriers' phenotype. Two of our three r(3) cases were phenotypically normal, and only one showed dysmorphisms and mild mental retardation (but the size of the sSMCs could not be investigated). All of these specific observations allow more tailored genetic counseling than that based on a generic risk of 14% (Warburton2), 28% (Crolla et al.26), or 31% (the present series). Given the size of the heterochromatic centromeric region of chromosome 3, it is particularly important to establish whether r(3) cases are only heterochromatic. The combined efforts of independent groups will make it possible to define a morbidity map of sSMCs.
Only nine of the cases in our series were checked for the presence of UPD, which was excluded on the basis of biparental heredity in all probands. Kotzot,28 in a review of the literature concerning UPD and sSMCs, concluded that the data available until 2002 were too scarce to give random or nonrandom significance to the UPD findings and suggested that systematic studies be performed to provide more information on this topic. Such a systematic approach was not applied on our sSMC carriers, but in the future perspective we believe that UPD search should be recommended at least in cases of sSMC originating from chromosomes carrying imprinted genes.
It is still not possible to correlate specific sSMCs with defined clinical pictures, mainly because of their highly heterogeneous nature, their origin from any chromosome, and their variable size and genetic content, which is not always finely identified. Only concerted efforts in defining phenotypes associated with subgroups of sSMCs or chromosome-specific sSMCs characterized by FISH analysis may lead to results suitable for establishing karyotype–phenotype correlations that can be used in genetic counseling.
References
Nielsen J, Wohlert M . Chromosome abnormalities found among 34910 newborn children: results from a 13-year incidence study in Arhus, Denmark. Hum Genet 1991; 87: 81–83.
Warburton D . De novo balanced chromosome rearrangements and extra marker chromosomes identified at prenatal diagnosis: clinical significance and distribution of breakpoints. Am J Hum Genet 1991; 49: 995–1013.
Diagnostica Citogenetica, Consensus 1995. Associazione Italiana di Citogenetica Medica. Analysis 1995; 8.
EUCROMIC Quality Assessment Group. Quality guide-lines and standards for genetic laboratories/clinics in prenatal diagnosis on fetal samples obtained by invasive procedures. Eur J Hum Genet 1997; 5: 342–350.
Mitelman F . An International System for Human Cytogenetic Nomenclature (ISCN). Basel: S. Karger; 1995; 8.
Doneda L, Dalprà L, Larizza L . Prenatal diagnosis of an extranimerary i(22p) with normal phenotype. Ann Genet 1993; 36: 154–158.
Tibiletti MG, Sala E, Colombo D, Arlati S, Varisco T, La Placa G . Chromosome 22 marker in a child with Duane syndrome and urogenital abnormalities. Ann Genet 1996; 39: 168–172.
Bettio D, Rizzi N, Giardino D, Guerrieri F, et al. FISH characterization of small supernumerary marker chromosomes in two Prader-Willi patients. Am J Med Genet 1997; 68: 99–104.
Pierluigi M, Battaglia P, Perfumo C, Baroncini A, Bricarelli FD . Combined use of cytogenetic analysis and FISH for the identification of two antenatal de novo markers as Robertsonian translocations involving the p arms. Ann Genet 1997; 40: 99–103.
Giardino D, Bettio D, Gottardi G, Rizzi N, et al. FISH characterization of two supernumerary r(1) associated with distinct clinical phenotypes. Am J Med Genet 1999; 84: 377–380.
Villa N, Riva P, Colombo D, Sala E, et al. Identification of a small supernumerary marker chromosome, r(2)(p10;q11.2), and the problem of determining prognosis. Pren Diagn 2001; 21: 801–805.
Borgatti R, Piccinelli P, Passoni D, Dalprà L, et al. Relationship between clinical and genetic features in “inverted duplicated chromosome15” patients. Pediatr Neurol 2001; 24: 111–116.
Giardino D, Finelli P, Russo S, Gottardi G, et al. Small familial supernumerary ring 2: FISH characterization and genotype-phenotype correlation. Am J Med Genet 2002; 111: 319–323.
Demori E, Devescovi R, Benussi DG, Dolce S, et al. Supernumerary ring chromosome 8: clinical and molecular cytogenetic characterization in a case report. Am J Med Genet 2004; 130A( 3): 288–294.
Wald NJ, Kennard A . Prenatal biochemical screening for Down's syndrome and neural tube defects. Curr Opin Obstet Gynecol 1992; 4( 2): 302–307.
Amor DJ, Choo KH . Neocentromeres: role in human disease, evolution, and centromere study. Am J Hum Genet 2002; 71: 695–714.
Anderlid BM, Sahlen S, Schoumans J, Holmberg E, et al. Detailed characterization of 12 supernumerary ring chromosomes using micro-FISH and search for uniparental disomy. Am J Med Genet 2001; 99: 223–233.
Stankiewicz P, Bocian E, Jakubów-Durska K, Obersztyn E, et al. Identification of supernumerary marker chromosomes derived from chromosomes 5, 6, 19, and 20 using FISH. J Med Genet 2000; 37: 114–120.
Daniel A, Malafiej P . A series of supernumerary small ring marker autosomes identified by FISH with chromosome probe arrays and literature review excluding chromosome 15. Am J Med Genet 2003; 117A: 212–222.
Starke H, Nietzel A, Weise A, Heller A, et al. Small supernumerary marker chromosomes (SMCs): genotype-phenotype correlation and classification. Hum Genet 2003; 114: 51–67.
Yagi H, Furutani Y, Hamada H, Sasaki T, et al. Role of TBX1 in human del22q11.2 syndrome. Lancet 2003; 362: 1366–1373.
Bartsch O, Nemeckova M, Kocarek E, Wagner A, et al. DiGeorge/velocardiofacial syndrome: FISH studies of chromosomes 22q11 and 10p14, and clinical reports on the proximal 22q11 deletion. Am J Med Genet 2003; 117A: 1–5.
Whittington J, Holland A, Webb T, Butler J, Clarke D, Boer H . Relationship between clinical and genetic diagnosis of Prader-Willi syndrome. (Letter). J Med Genet 2002; 39: 926–932.
Liehr T, Claussen U, Starke H . Small supernumerary marker chromosomes (sSMC) in humans. Cytogenet Genome Res 2004; 107: 55–67.
Crolla JA, Youings SA, Ennis S, Jacobs PA . Supernumerary marker chromosomes in man: parental origin, mosaicism and maternal age revisited. Eur J Hum Genet 2005; 13: 154–160.
Crolla JA . FISH and molecular studies of autosomal supernumerary marker chromosomes excluding those from chromosome 15: review of the literature. Am J Med Genet 1998; 75: 367–381.
Dalprà L, Tibiletti MG, Cristiani C, Ragni G, Crosignani PM . Morphology and nuclear contents in IVF embryos. Ann Genet 1993; 36: 100–106.
Kotzot D . Supernumerary marker chromosomes (SMC) and uniparental disomy (UPD): coincidence or consequence?. J Med Genet 2002; 39: 775–778.
Acknowledgements
This work was partially supported by Italian Ministry of Health to Istituto Auxologico Italiano (grant number: 030.11/RF01.154).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dalprà, L., Giardino, D., Finelli, P. et al. Cytogenetic and molecular evaluation of 241 small supernumerary marker chromosomes: Cooperative study of 19 Italian laboratories. Genet Med 7, 620–625 (2005). https://doi.org/10.1097/01.gim.0000182876.57766.2d
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/01.gim.0000182876.57766.2d
Keywords
This article is cited by
-
Molecular delineation of small supernumerary marker chromosomes using a single nucleotide polymorphism array
Molecular Cytogenetics (2020)
-
Familiar unbalanced complex rearrangements involving 13 p-arm: description of two cases
Molecular Cytogenetics (2018)
-
Genetic dosage and position effect of small supernumerary marker chromosome (sSMC) in human sperm nuclei in infertile male patient
Scientific Reports (2015)