Abstract
First described as a variant of Larsen syndrome in Reunion Island (LRS) in the southern Indian Ocean, ‘Larsen of Reunion Island syndrome’ is characterized by dwarfism, hyperlaxity, multiple dislocations and distinctive facial features. It overlaps with Desbuquois dysplasia, Larsen syndrome and spondyloepiphyseal dysplasia with dislocations ascribed to CANT1, FLNB and CHST3 mutations, respectively. We collected the samples of 22 LRS cases. After exclusion of CANT1, FLNB and CHST3 genes, an exome sequencing was performed in two affected second cousins and one unaffected sister. We identified a homozygous missense mutation in B4GALT7, NM_007255.2: c.808C>T p.(Arg270Cys) named p.R270C, in the two affected cases, not present in the unaffected sister. The same homozygous mutation was subsequently identified in the remaining 20 LRS cases. Our findings demonstrate that B4GALT7 is the causative gene for LRS. The identification of a unique homozygous mutation argues in favor of a founder effect. B4GALT7 encodes a galactosyltransferase, required for the initiation of glycoaminoglycan side chain synthesis of proteoglycans. This study expands the phenotypic spectrum of B4GALT7 mutations, initially described as responsible for the progeroid variant of Ehlers–Danlos syndrome. It further supports a common physiopathological basis involving proteoglycan synthesis in skeletal disorders with dislocations.
Similar content being viewed by others
Introduction
Larsen of Reunion Island syndrome (LRS) [MIM 245600] has been described as a specific entity from Reunion Island in the south of the Indian Ocean in 1975.1 The clinical manifestations of LRS include dislocations of large joints with ligamentous hyperlaxity, short stature and characteristic facial features, namely, round flat face, prominent forehead, prominent bulging eyes, under-eye shadows and microstomia. Radiological features include dislocations of knees, hips, elbows and fingers, advanced carpal ossification, widened metaphyses, particularly at the knees, and often aradioulnarsynostosis.1, 2
LRS shares many clinical features with other conditions from the multiple dislocation group,3 namely, Larsen syndrome (MIM 150250, 245600, LS),4, 5, 6 spondyloepiphyseal dysplasia with dislocations (MIM 608637) and Desbuquois dysplasia (MIM 251450), ascribed to filamin B (FLNB),7 carbohydrate sulfotransferase 3 (CHST3)8 and calcium-activated nucleotidase 1 (CANT1) mutations,9 respectively.
In 1992, Bonaventure et al10 studied seven LRS children from three families from the Reunion Island, and excluded by linkage analysis four collagen candidate genes, namely, COL1A1, COL1A2, COL3A1 and COL5A2.
The aim of our study was to identify LRS gene. Among the 53 LRS patients followed in our department in the last 20 years, only 22, originating from 20 distinct families, had detailed clinical data and were therefore included in the study. We first excluded FLNB, CANT1 and CHST3 by direct sequencing (data not shown). Considering the lack of multiplex families and the absence of consanguinity in our families, an exome sequencing strategy was undertaken, leading to the identification of B4GALT7 as the causative gene of LRS.
Subjects and methods
Clinical and radiological features
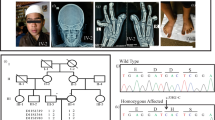
The 22 cases fulfilled the diagnostic criteria for LRS, namely, (i) characteristic facial features, (ii) multiple joint dislocations (knees, elbows, hips) with hyperlaxity and (iii) severe short stature. The series included 11 males and 11 females, ranging in age from 4 to 46 years (Figure 1a–d). The clinical and radiological features of all patients are presented in Tables 1 and 2.
(a) Left: the female proband at 2 years of age, with round, flattened midface, prominent forehead, circles under eyes and microstomia, right: paternal uncle at 29 years of age with very small stature (122 cm) and obesity (BMI: 45.9). (b) Left: hand at 28 months of age with short fingers, right: X-Rays showing advanced carpal ossification (3½ years contrasting with the chronological age of 28 months), delta phalanx and proximal radioulnar synostosis. (c) Clinically unstable knee joint (left) confirmed by the femorotibial misalignment (right). (d) X-rays showing Swedish key appearance of the proximal femur (left) and radioulnar synostosis (right). (e) Pedigree of the family studied by exome sequencing. (f) Chromatograms showing the mutation in B4GALT7 gene in control (wild type), affected (homozygous patients) and carrier (heterozygous parents) cases.
Molecular analysis
We selected two second cousins (one male and one female) and the unaffected sister of the affected male to perform exome sequencing (Figure 1e). For each subject, DNA samples from index cases and their parents and unaffected sibs (if any) were obtained following written informed consents. Genomic DNA was extracted from peripheral blood using standard procedure.
Exome sequencing
Exome sequencing was performed by IntegraGen (Evry, France). Enrichment was performed using 3 mg of genomic DNA and an Agilent SureSelect Human All Exon Kit V2, 46 Mb (Agilent Technologies, Santa Clara, CA, USA) in accordance with the manufacturer’s protocols.11 After sonication, DNA was fragmented and purified to yield fragments of mean size 150–200 bp. Each elute-enriched DNA sample was then sequenced on an IlluminaHiSEQ 2000 (Illumina Inc, San Diego, CA, USA) as paired-end 75b reads. Image analysis and base calling were performed using Illumina Real Time Analysis Pipeline version 1.9 with default parameters. On average, 7.4 Gb of sequence was generated for each subject to achieve 74 × and 90% (X10X) coverage. The bioinfomatics analysis of sequencing data was based on the Illumina pipeline (CASAVA 1.7). The alignment algorithm used was ELANDv2 (Illumina Inc) (performs multiseed and gapped alignments). Genetic variation annotation was performed using ANNOVAR software12 including annotations (RefSeq hg19).
Sanger sequencing
A 666-bp fragment harboring the candidate mutation on chr5:g.177035995(hg Built 37.6) was amplified using primer couples with melting temperature of 58 °C (primers on request). Sequencing was performed using ABI PRISM 3130 BigDye terminator chemistry according to the manufacturer’s instruction (Applied Biosystems, Foster City, CA, USA). Sequences were analyzed using SeqScape v2.7 (Applied Biosystems).
Bioinformatics analysis
We predicted the effects of amino-acid substitution on protein stability and function on the basis of the following methods including in Annovar: SIFT, PolyPhen, PhyloP, LRT, MutationTaster, GERP++.
Results
We focused our analyses on non-synonymous variants located in coding sequence by assuming that synonymous variants were less likely to be pathogenic. DNA variants were filtered on the basis of the presence of variants in the same gene in the two affected individuals and on their absence in the unaffected sister. First, a total of 117 310 variants (single-nucleotide polymorphisms and small insertions/deletions) was identified; after using various computational filtering criteria including the hypothesis of a recessive inheritance of rare mutations, 4146 homozygous variants present in the two patients and not in the healthy sister were found. After filtering with Exome Sequencing Project database, only seven synonymous variants were selected and dbSNP 137 NonFlagged allowed to retain an unique variant in B4GALT7 gene: it was predicted to have a functional effect by the five functional prediction algorithms in Annovar. This mutation was confirmed by direct sequencing in the two affected individuals, although their parents were heterozygous (Figure 1f). The B4GALT7 mutation was submitted to the corresponding LOVD database at http://databases.lovd.nl/shared/variants/0000031236. We sequenced B4GALT7 gene in the other 20 patients: all were homozygous for this mutation.
A total of 500 control cases (chosen among the same ethnic group ‘white creoles’ than that of our patients) was studied. No control case was homozygous for the mutation. In this ethnic group, the allelic frequency was 2%, corresponding to a prevalence of 1/2500 births.
Discussion
We report here the identification of a homozygous B4GALT7 mutation c.808C>T p.(Arg270Cys) in 22/22 LRS and confirm that LRS is a distinct condition from previously reported skeletal disorders with multiple dislocations. The B4GALT7 mutation was submitted to the corresponding LOVD database at http://databases.lovd.nl/shared/variants/0000031236
The identification of a unique homozygous B4GALT7 mutation c.808C>T p.(Arg270Cys) in our cohort supports a founder effect. Our patients belong to the same ethnic group named ‘white creoles’, representing about 15.5% of the population from Reunion Island.
With a total of 22 patients ranging from 4 to 46 years, our cohort of patients with Larsen of Reunion Island constitutes the largest one ever reported in the literature, leading to define key features for clinical diagnosis (Tables 1 and 2). Indeed, LRS must be considered in the presence of multiple dislocations (21/21, 100%), severe pre- and postnatal dwarfism (19/19, 100%) with height often inferior to −6 SD (growth hormone therapy was administered in none of the cases), advanced carpal ossification (11/15, 73%) and facial features. All the patients presented with high forehead, hypertelorism, protruding eyes with under-eye shadows, hypoplastic midface and microstomia with protruding lips; a thin prominent nasal bridge tented to appear with age. Progeroid aspect was absent in all cases. Learning difficulties were reported in more than a half of the patients; however, repeated surgeries were also leading to significant difficulties in attending school daily. Other frequently observed features were a Swedish key appearance of the proximal femora (9/19, 47%) and radioulnar synostosis (10/21, 48%) without progressive severe arthritic changes. By contrast, progressive scoliosis, accessory ossification center at the base of the proximal phalange of the second digit and/or a delta phalanx of the thumb, which are hallmarks of Desbuquois dysplasia, were not observed in LRS. In addition, platyspondyly and intervertebral space narrowing, which are characteristic of spondyloepiphyseal dysplasia with dislocations, were never observed in LRS.
Our series of mutations in B4GALT7 constitutes the largest one ever recorded in the literature; it gives us the opportunity to also revise the clinical phenotype related to B4GALT7 mutations. Only four patients have been reported13, 14, 15, 16 with B4GALT7 mutations including either compound heterozygous mutations c.[557C>A];[617T>C] and c.[808C>T];[122T>C] or the homozygous c.[808C>T] also identified in LRS patients. The clinical manifestations of these four patients have been recently reviewed16 and included wrinkled loose facial skin, curly fine hair, scanty eyebrows and eyelashes, short stature and development anomalies of forearm and elbows with proximal radioulnar synostosis. Importantly, progeroid facial appearance was not consistently observed15 questioning the name of ‘progeroid type of Ehlers–Danlos syndrome’.16 The detailed description of the 22 LRS patients confirms the absence of progeroid symptoms and highlights the frequency and the severity of large joint dislocations requiring repeated surgical procedures. The joint hypermobility is present in all patients reported with B4GALT7 mutations and particularly severe in the young girl reported by Faiyaz-Ul-Haque et al.15 She was firstly suspected of Larsen syndrome but joint dislocations have never been reported in the previous reports. These clinical differences might be explained by different functional consequences of the reported mutations with variable quantitative effect on glycosaminoglycan (GAG) biosynthesis. Whereas the c.557C>A p.(Arg186Asp) mutation does not affect GAG biosynthesis severely, the c.617T>C p.(Leu206Pro) mutation leads to a complete inhibition and the c.808C>T p.(Arg270Cys) mutation to a significant reduction of GAG biosynthesis.17 Nevertheless, the phenotypic differences between the patients reported by Faiyaz-Ul-Haque et al15 and LRS patients who are sharing the same mutation may have other explanations. Given the insular peopling pattern and the high level of homozygosity in the white creoles population, the interaction with other variants in close linkage disequilibrium to B4GALT7 could be hypothesized as well as the involvement of modifier genes.
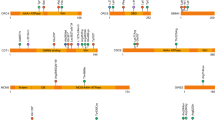
B4GALT7 is directly involved in the biosynthesis of proteoglycans. These macromolecules consist in GAG polymer chains attached to core proteins. The synthesis of GAG chains is initiated by the formation of a tetrasaccharide linker region attached to a serine residue of the core protein. The synthesis of this linker region starts by the transfer of xylose onto a serine residue of the core protein catalyzed by xylosyltransferases and, subsequently, two galactose residues are added by galactosyltransferase I (B4GALT7, encoded by B4GALT7 gene [MIM 604327])18, 19 and galactosyltransferase II (B3GALT6, encoded by B3GALT6)20 followed by the transfer of glucuronic acid by glucuronosyltransferase I (GlcAT-I, encoded by B3GAT3) [MIM 606374].21 After this tetrasaccharide linkage region synthesis, the GAG chain is modified by epimerization and sulfation.
Previous functional studies have shown that the c.808C>T p.(Arg270Cys) mutation identified in B4GALT7 causes a reduction in galactosyl transferase activity, directly involved in the tetrasaccharide linker synthesis. A lack of decorin and biglycan synthesis and reduced epimerization of the chain of decorin glycosaminoglycans have been demonstrated in p.(Arg270Cys)-mutant fibroblasts from progeroid Ehlers–Danlos patient.22 This deficiency results in a loss of cohesion of the tissue, which has been further demonstrated by the ultrastructural study showing abnormal connective tissue.23
In the last few years, the impairment of proteoglycan synthesis has been involved in a few human connective tissue disorders, namely, (1) SED with dislocations due to CHST3 mutations responsible for a defect in carbohydrate chondroitin 6-sulfotransferase with defective sulfation of chondroitin proteoglycan,8, 24 (2) Desbuquois dysplasia due to CANT1 mutations responsible for a defect in GAG synthesis,9, 25 (3) a chondrodysplasia with joint dislocations due to mutations in IMPAD1 (inositol monophosphatase domain-containing protein 1) encoding the Golgi resident nucleotide phosphatase gPAPP, and responsible for a defect in sulfated substrates,26 (4) ‘a Larsen-like’ phenotype with cardiac defects due to B3GAT3 (galactosyltransferase) mutations27 and affecting, through the linker synthesis defect, dermatan heparan and chondrotin sulfate proteoglycans and (5) very recently, a spondyloepimetaphyseal dysplasia with joint laxity type 1 and a spectrum of disorders affecting a broad range of skeletal and connective tissues characterized by lax skin, muscle hypotonia, joint dislocation and spinal deformity due to B3GALT6 mutations28 (Supplementary Table).
In conclusion, our study demonstrates that LRS shares a common physiopathological basis with the other multiple dislocation disorders involving proteoglycan synthesis from the linker synthesis (B4GALT7, B3GAT3, B3GALT6) to GAG chain addition and modification (CANT1, CHST3, IMPAD1). It also highlights that LRS constitutes a specific entity, and expandes the clinical spectrum of B4GALT7 mutations, supporting the screening of B4GALT7 in the presence of severe pre- and postnatal growth retardation, joint dislocations, radioulnar synostosis and advanced carpal bone age, without progeroid appearance, especially in a patient from Reunion Island.
References
Payet G : Nanisme et hyperlaxité dysmorphie faciale et luxations multiples: syndrome de Larsen? Arch Fr Pediatr 1975; 32: 601–608.
Laville JM, Lakermance P, Limouzy F : Larsen’s syndrome: review of the literature and analysis of thirty-eight cases. J Pediatr Orthop 1994; 14: 63–67.
Warman ML, Cormier-Daire V, Hall C et al: Nosology and classification of genetic skeletal disorders: 2010 revision. Am J Med Genet A 2011; 155A: 943–968.
Larsen LJ, Schottstaedt ER, Bost FC : Multiple congenital dislocations associated with characteristic facial abnormality. J Pediatr 1950; 37: 574–581.
Harris R, Cullen CH : Autosomal dominant inheritance in Larsen’s syndrome. Clin Genet 1971; 2: 87–90.
Latta RJ, Graham CB, Aase J, Scham SM, Smith DW : Larsen’s syndrome: a skeletal dysplasia with multiple joint dislocations and unusual facies. J Pediatr 1971; 78: 291–298.
Krakow D, Robertson SP, King LM et al: Mutations in the gene encoding filamin B disrupt vertebral segmentation, joint formation and skeletogenesis. Nat Genet 2004; 36: 405–410.
Thiele H, Sakano M, Kitagawa H et al: Loss of chondroitin 6-O-sulfotransferase-1 function results in severe human chondrodysplasia with progressive spinal involvement. Proc Natl Acad Sci USA 2004; 101: 10155–10160.
Huber C, Oules B, Bertoli M et al: Identification of CANT1 mutations in Desbuquois dysplasia. Am J Hum Genet 2009; 85: 706–710.
Bonaventure J, Lasselin C, Mellier J, Cohen-Solal L, Maroteaux P : Linkage studies of four fibrillar collagen genes in three pedigrees with Larsen-like syndrome. J Med Genet 1992; 29: 63–73.
Gnirke A, Melnikov A, Maguire J et al: Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat Biotechnol 2009; 27: 182–189.
Wang K, Li M, Hakonarson H : ANNOVAR: functional annotation of genetic variants from next-generation sequencing data. Nucleic Acids Res 2010; 38: e164.
Kresse H, Rosthoj S, Quentin E et al: Glycosaminoglycan-free small proteoglycan core protein is secreted by fibroblasts from a patient with a syndrome resembling progeroid. Am J Hum Genet 1987; 41: 436–453.
Okajima T, Fukumoto S, Furukawa K : Molecular basis for the progeroid variant of Ehlers Danlos-syndrome: identification and characterization of two mutations in galactosyltransferase I gene. J Biol Chem 1999; 274: 28841–28844.
Faiyaz-Ul-Haque M, Zaidi SH, Al-Ali M et al: A novel missense mutation in the galactosyltransferase-I (B4GALT7) gene in a family exhibiting facioskeletal anomalies and Ehlers-Danlos syndrome resembling the progeroid type. Am J Med Genet A 2004; 128A: 39–45.
Guo MH, Stoler J, Lui J et al: Redefining the progeroid form of Ehlers-Danlos syndrome: report of the fourth patient with B4GALT7 deficiency and review of the literature. Am J Med Genet A 2013; Part A 9999: 1–9.
Rahuel-Clermont S, Daligault F, Piet MH et al: Biochemical and thermodynamic characterization of mutated β1,4-galactosyltransferase 7 involved in the progeroid form of the Ehlers-Danlos syndrome. Biochem J 2010; 432: 303–311.
Okajima T, Yoshida K, Kondo T, Furukawa K : Human homolog of Caenorhabiditiselegans sqv-3 is galactosyltransferase I. A seventh member of the human beta4-galactosyltransferase gene family. J Biol Chem 1999; 274: 26165–26171.
Almeida R, Levery SB, Mandel U, Kresse H, Schwientek T, Benett EP, Clausen H : Cloning and expression of a proteoglycan UPD-galactose: beta-xylose beta 1,4-galactosyltransferase gene family. J Biol Chem 1999; 274: 26165–26171.
Bai X, Zhou D, Brown JR, Crawford BE, Hennet T, Esko JD : Biosynthesis of the linkage region of glycosaminoglycans: cloning and activity of galactosyl-transferase II, the sixth member of the beta 1,3-galactosyltransferase family (bet3 GalT6). J Biol Chem 2001; 276: 48189–48195.
Kitagawa H, Tone Y, Tamura J et al: Molecular cloning and expression of glucuronyltransferase I involved in the biosynthesis of the glycosaminoglycan-protein linkage region of protein linkage region of proteoglycans. J Biol Chem 1998; 273: 6615–6618.
Seidler DG, Faiyaz-Ul-Haque M, Hansen U et al: Defective glycosylation of decorin and biglycan, altered collagen structure, and abnormal phenotype of the skin fibroblasts of an Ehlers-Danlos syndrome patient carrying the novel Arg270Cys substitution in galactosyltransferase I (beta4GalT-7). J Mol Med (Berl) 2006; 84: 583–594.
Cetta G, Lenzi L, Ruggeri A, Tenni R, Boni M : Biochemical and structural abnormalities of the connective tissue in Larsen’s syndrome. Int Orthop 1979; 3: 47–53.
Hermanns P, Unger S, Rossi A et al: Congenital joint dislocations caused by carbohydrate sulfotransferase 3 deficiency in recessive Larsen syndrome and humero-spinal dysostosis. Am J Hum Genet 2008; 82: 1368–1374.
Nizon M, Huber C, De Leonardis F et al: Further delineation of CANT1 phenotypic spectrum and demonstration of its role in proteoglycan synthesis. Hum Mutat 2012; 33: 1261–1266.
Vissers LE, Lausch E, Unger S et al: Chondrodysplasia and abnormal joint development associated with mutations in IMPAD1, encoding the Golgi-resident nucleotide phosphatase, gPAPP. Am J Hum Genet 2011; 88: 608–615.
Baasanjav S, Al-Gazali L, Hashiguchi T et al: Faulty initiation of proteoglycan synthesis causes cardiac and joint defects. Am J Hum Genet 2011; 89: 15–27.
Malfait F, Kariminejedy A, Van Damme T et al: Defective initiation of glycosaminoglycan synthesis due to β3GALT6 mutations causes a pleiotropic Ehlers-Danlos-Syndrome-like connective tissue disorder. Am J Hum Genet 2013; 92: 935–945.
Acknowledgements
This work was supported by the Institut National de la Santé et de la Recherche Médicale (INSERM). We would like to thank all individuals with Larsen of Reunion Island syndrome and their family members for participation in this study. We thank the DNA bank (Centre de Ressources Biologiques, La Réunion) and Ms G.FRIDERICI for her help and generous material gift. Ethics approval was provided by Ethics Committee of Centre Hospitalo-Universitaire of Reunion Island.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Author Contributions
FC, BD and VC-D wrote the paper; FC, M-LJ and HR provided the clinical data; PM, CP and JV performed PCR and Sanger sequencing; FC and PM performed the exome analysis; VC-D, AM and FC analyzed the data.
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Cartault, F., Munier, P., Jacquemont, ML. et al. Expanding the clinical spectrum of B4GALT7 deficiency: homozygous p.R270C mutation with founder effect causes Larsen of Reunion Island syndrome. Eur J Hum Genet 23, 49–53 (2015). https://doi.org/10.1038/ejhg.2014.60
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2014.60
This article is cited by
-
Report of two siblings with spondylodysplastic Ehlers-Danlos syndrome and B4GALT7 deficiency
BMC Pediatrics (2021)
-
Expansion of B4GALT7 linkeropathy phenotype to include perinatal lethal skeletal dysplasia
European Journal of Human Genetics (2019)
-
SLC10A7 mutations cause a skeletal dysplasia with amelogenesis imperfecta mediated by GAG biosynthesis defects
Nature Communications (2018)
-
Expanding the clinical and mutational spectrum of B4GALT7-spondylodysplastic Ehlers-Danlos syndrome
Orphanet Journal of Rare Diseases (2017)
-
Clinical utility gene card for: B4GALT7-defective congenital disorder of glycosylation
European Journal of Human Genetics (2017)