Abstract
Mammary-digital-nail syndrome is a novel phenotypic association consisting of anonychia onychodystrophy with hypoplasia or absence of distal phalanges in males and females, accompanied by juvenile hypertrophy of the breast in affected females. This newly described genetic trait presents an autosomal dominant inheritance pattern, with either reduced penetrance or germ-line mosaicism. Analysis of the pedigree, linkage studies followed by a genome-wide screen and by haplotype analysis defined the locus for the phenotype within a 12 cM (4.3 Mb) interval on chromosome 22q12.3-13.1. This chromosomal region has not been implicated before in genetic disorders of the mammary tissue or limbs. These data suggest a possibly novel signaling pathway affecting the organogenesis of limbs and mammary glands in humans.
Similar content being viewed by others
Introduction
Familial juvenile hypertrophy of the breast (JHB) is a rare condition with very little known documentation1, 2, 3 characterized by gigantomastia in peripubertal females. The pathology in JHB is limited to the breast, with otherwise normal growth and development.1, 2, 4 We previously described a familial case of JHB comprising four affected females.2 Another affected family consists of a mother–daughter pair.1 A third familial case of breast hypertrophy was reported in two sisters3 who experienced abnormal breast enlargement during pregnancy, though this phenotype seems different from JHB. Over 100 cases of sporadic JHB have been documented.4, 5 Clinical features include bilateral or unilateral rapid breast enlargement, skin hyperemia, and widened subcutaneous veins.2 The growth of the glands begins between ages 8 and 16, just before or shortly after menarche. Gigantomastia is treated surgically by breast reduction or mastectomy.1, 2, 4 JHB is a major source of psychological and social distress.2, 4
The genetic basis for familial JHB has not been deciphered. A single report links a mutation in PTEN to virginal breast hypertrophy and neoplasia in the mammary gland.6
Familial onychodystrophy/anonychia with abnormalities of the distal phalanges (ODP; Cooks syndrome, OMIM 106995) is another rare disorder, characterized by onychodystrophy, anonychia, brachydactyly of the fifth finger, and digitalization of the thumbs, with absence or hypoplasia of the distal phalanges of the hands and feet. Generally, the nails of the first to third digits are progressively deformed with total anonychia in the two last digits and in all toes.7, 8 Although all four documentations of this syndrome report that the affected individuals in the first generation were born to unaffected parents, an autosomal dominant (AD) mode of inheritance has been proposed based on the pedigree structure seen in the following generations, with affected offspring born to an affected parent (male and female) along two to three generations.7, 8, 9
We present here for the first time a clinical, molecular, and genetic study of three successive generations of a large kindred, in which ODP afflicted eight males, and co-segregation of JHB and ODP presented in six females. We have denoted this newly described genetic syndrome Mammary-Digital-Nail syndrome (MDN).
Patients and methods
The institutional ethics review committee and the national committee for genetic studies of the Israeli Ministry of Health approved the study protocol. All participants or their legal guardians signed informed consent.
Patients
Patient IV25, the first affected individual to be identified, was referred for breast reduction surgery due to premenarche massive breast enlargement, as we previously reported.2 Her immediate and extended family lives in an isolated village of 13 000 Druze residents in northern Israel. Ancestors of this kindred immigrated to Israel from Lebanon in 1711.10 Two physicians conducted complete physical examinations and detailed interviews of the medical and family history of the 41 members of the kindred who agreed to participate in the study. Patients III19, IV20, IV25, and IV28 (Figure 2) underwent X-rays of hands, feet, and long bones.
Skin and breast tissues were frozen and fibroblast cultures were established from patients who underwent surgeries.2
Genotyping
Blood was drawn and DNA was extracted from 41 members of the investigated kindred, and from 100 unaffected Druze individuals residing in the same village. Whole-genome screening was performed using 400 polymorphic microsatellite markers (Linkage Mapping Set version 2.5 MD10; Applied Biosystems, Foster City, CA, USA) spaced at an average of 10 cM, and with an average heterozygosity of 79%.
DNA fragments were PCR amplified using fluorescently labeled PCR primers according to the manufacturer's instructions. Genotypes were determined using the ABI Prism 3100 DNA analyzer (Applied Biosystems) and GeneScan 3.1, Genotyper 3.1, and GeneMapper 1.1 software. The data were evaluated for Mendelian inheritance of marker alleles by two-point analysis and haplotype analysis programs, using the Superlink-online software.11, 12 Additional polymorphic microsatellite markers, based on the Genethon human linkage map, were used for fine-mapping and to refine the boundaries of suspected candidate regions.
Linkage analyses
Genome-wide linkage analyses were initially performed for 17 family members (9 affected, 8 unaffected) using the Superlink-online software.11, 12 Two point and multipoint LOD score analyses were performed under both AD and AR modes of inheritance, based on the compatibility of the pedigree with both modes of inheritance. We used a penetrance of 50–99% for AD and 99% for AR modes of inheritance, with a hypothetical prevalence of 0.1% for the disease, based on the rarity of this syndrome. Fine-mapping data were analyzed using the same software with an increasing number of markers in the suspect areas, and with an increasing number of genotyped persons. Using genotyping data from 11 affected and 31 unaffected individuals, we constructed haplotypes and a recombination map for all genomic intervals suggestive for linkage.
Cytogenetic and SNP array analyses
Formal cytogenetic analysis was carried out for two affected females using routine Giemsa staining. Whole-genome SNP analyses used the Affymetrix GeneChip Mapping 250K Sty array, which is enriched with chromosome 22 markers. Analyses were performed for eight individuals, including three affected (two males and one female), and five unaffected, including two carriers of the haplotype (one family member and one from the control group) associated with the disease (see text in Results). Following genotyping and copy number variation (CNV) analysis using Partek software (Weizmann Institute Facilities), haplotypes and a recombination map were constructed.
Sequence analysis
The sequences of all exons and the flanking splice junctions of 38 candidate genes in the linked interval were determined using genomic DNA from unaffected and affected family members. Sequences were determined using the BigDye Terminator cycle sequencing kit version 1.1 (Applied Biosystems), and analyzed using the ABI PRISM 3100 Genetic Analyzer (Applied Biosystems), according to the manufacturer's instructions.
Expression studies
Real-time PCR (RT-PCR) analyses were carried out for five sequenced genes (C1QTNF6, SSTR3, RAC2, LGALS1, and ANKRD54) that exhibited sequence alterations segregating with the phenotype in the investigated kindred. We compared cDNA isolated from skin and/or mammary tissues of four affected family members (two males and two females), IV19, IV25, IV21, and IV28 (Figure 2) to that of five unrelated unaffected individuals (three males and two females). Breast tissue was removed solely from females. The samples were electronically homogenized using Kinematica polytron homogenizer PT 2100. Total RNA was extracted using the RNeasy lipid tissue mini kit (Qiagen GmbH, Hilden, Germany). cDNA was synthesized using the high-capacity cDNA reverse transcription kit (Applied Biosystems). RT-PCR analysis was performed using the miScript reverse transcription kit (Qiagen) and commercial primers suitable to SYBR Green (QuantiTect Primer Assays; Qiagen). The primary cDNA levels were calibrated according to the expression of the housekeeping genes GAPDH and GUSB. Samples were run on the ABI PRISM 7300 analyzer and the relative quantification rates were calculated based on the Ct values. Statistical analysis was performed using the Wilcoxon rank sum test.
Results
Clinical phenotype
Eleven of the individuals examined in this extended family (six males and five females), presented with ODP since birth (Figure 1a–c). In addition, ODP was described in three deceased members of the family (two males and one female). The clinical presentation comprises digitalization of thumbs, hypoplasia of the distal phalanges of digits 2–4, total absence of distal phalanges of digit 5, bilateral absence of all distal ceases of digits 2–5, and hypoplasia and complete absence of distal phalanges and nails of the great toes (Figure 1a–c).
The onychodystrophy/anonychia with abnormalities of the distal phalanges (ODP) phenotype. (a) The hands of patients III19 and IV28 (male) showing onychodystrophy, anonychia, brachydactyly of the fifth digits, and digitalization of thumbs. The hands of IV25 (female) showing anonychia and brachydactyly of the fifth digits. (b) X-rays showing digitalizing of both thumbs (III19, IV28), hypoplasia of the distal phalanges of digits 2–4 and total absence of all distal phalanges of digit 5 (III19, IV25, IV28). (c) X-rays showing total bilateral absence of all distal phalanges of digits 2–5; hypoplasia of distal phalanges of great toes in III19 and IV28; complete absence of distal phalanges of the great toes in IV25. Patient numbers are according to a. (d) Gigantomastia in patient IV25 that appeared 3 months before and shortly after menarche. Congenital anonychia of hands can also be seen2 (published with permission).
The JHB phenotype presented only in females. In patients IV25, III10, IV19, and IV20 (Figure 2) breasts enlarged dramatically, reaching enormous proportion during a 3-month period before and shortly after menarche.2 Of note, female patient III9, who died at the age of 3 years, well before her premenarche time, presented with ODP only. Female patient V1 is presently 2 years old, and presents with ODP since birth.
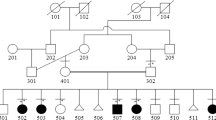
Extended pedigree with haplotypes. A five-generation pedigree of the investigated family with haplotypes for five markers that span ∼30 cM on chromosome 22q12.3–13.1. The order of markers is indicated to the left of the pedigree. Generation numbers appear above the haplotype data. Solid black shading represents the ODP phenotype: (III-9, III-10, III-15, III-16, III-17, III-19, IV-19, IV-20, IV-21, IV-23, IV-25, IV-28, IV-29, V-1) and grey shading represents the JHB phenotype: (III-10, IV-19, IV-20, IV-25). The common affected haplotype that segregates with the disease is denoted with grey shaded background. The affected individual V1 presents a crossover event between markers D22S424 and D22S277 leading to narrowing the linked disease interval to 12 cM (4.3 Mb). Individuals with irrelevant haplotypes are denoted with Eu. The haplotype of the deceased father II4 is deduced based on genotyping results of his family. The nine unaffected individuals and the deceased father II4, who share the common 'disease haplotype‘, are emphasized as bold numbers. The nuclear family, from which the disease started, is marked by a solid bold line.
The one or two breast reduction and reconstruction surgical operations performed in each of the four females affected with gigantomastia were previously described2 (Figure 1d). The four affected individuals have regular menses, and normal laboratory work-up including endocrine profile, lipid profile, liver and kidney function, and basic hematology. None was exposed to drug or hormonal therapy. All became victims of social ridicule and stopped attending school.
All unaffected females in the kindred have normal fingers, toes, and nails; and no symptoms of JHB. All examined males had normal breasts. Other than the symptoms of JHB and ODP described, the physical examinations for all 41 family members, including secondary sex characteristics, were unremarkable, as was independently verified with their respective family physicians. We thus conclude that the phenotype is ODP in males and ODP plus JHB in females.
Inheritance pattern and genetic mapping
None of the individuals in the first and second generations of the investigated pedigree was affected. In the third generation, the disease phenotype presented in 6 of 14 (43%) offspring of one unaffected couple. We examined the unaffected mother and received information regarding the deceased unaffected father from several sources. All affected individuals in the fourth and fifth generations, both males and females, were offspring of affected fathers. Affected females are socially isolated and have remained unmarried and without offspring.
In the fourth generation, four of five, and three of six, offspring from the nuclear families of two affected brothers III17 and III19 (Figure 2) were affected. The only affected individual in the fifth generation is a 2-year-old daughter of an affected father.
Because the kindred is part of an isolate with both overt and ancestral consanguinity, pseudodominant inheritance of a recessive disorder should be considered. However, the pedigree structure is also compatible with AD inheritance with either reduced penetrance or germ-line mosaicism (GM) in a parent of the sibship with the initial affected individuals. The log likelihood ratio between AD and AR inheritance was insignificant (LOD <1.5) for penetrance values in the range of 50–99%, assuming a disease prevalence of 0.1%. Hence, genome-wide LOD scores were computed under both modes of inheritance. Using genotypes from 25 family members, we could not obtain significant LOD score values under an AR model with pseudodominant inheritance throughout the genome. Because the pedigree contains a number of nuclear families with MDN in generations III, IV and V, and one family with parental consanguinity, we reanalyzed each nuclear family independently under the pseudodominant model. No common linkage was obtained among the nuclear families and no significant combined LOD score was achieved. When performing the analysis two considerations were applied: first, evidence of homozygosity for an ancestral mutation was the most likely explanation for this population under this model. Second and less likely was the presence of two mutations and compound heterozygosity in the affected individuals. The linkage studies ruled out all regions of the genome under a recessive model. A maximum LOD score of 4.27 was obtained under AD inheritance, with full penetrance and allele prevalence of 0.1%, at three continuous loci (D22S1154, D22S277, and D22S283) (Table 1), located within a 30 cM (19.3 Mb) interval on chromosome 22. Linkage analysis using 70% penetrance led to similar results with a maximum LOD score of 3.6 at the same three markers.
No significant linkage was obtained to any other chromosome. Nevertheless, to rule out other options for linkage throughout the genome, suspected regions with LOD scores higher than 1.0 were reanalyzed under both AD and AR inheritance models, using additional markers and additional members of the kindred. Also here, no supporting linkage was obtained. The possibility of X-linked inheritance was excluded.
Fine-mapping and haplotype analyses
Haplotype analysis using 22 markers along the primary linked interval revealed a common haplotype, derived from the unaffected founder father II4, in all 11 affected individuals. The genotypes of patient V1 identified a crossover event between the markers at loci D22S424 and D22S277, leading to significant narrowing of the suspected interval, from 30 cM (19.3 Mb) to 12.5 cM (4.3 Mb). The narrowed interval includes the markers at loci D22S277 and D22S283, which exhibited the highest LOD score.
Surprisingly, 9 of the 30 unaffected family members examined carry the same haplotype as the affected individuals. Possible explanations for these findings are reduced penetrance of MDN or GM in the patriarch (II4) of this family. The former implies a possible modifier gene responsible for the reduced penetrance. To evaluate this possibility, we analyzed genotypes to determine whether the nine unaffected individuals with the disease-associated haplotype shared an additional common genomic interval not carried by the affected individuals. The analysis was performed as described above, under both AD and AR inheritance models with full penetrance and with 1% theoretical allele prevalence, based on the assumption that the protective allele should be more common than the disease-causing allele. No significant linkage was obtained, based on low LOD scores across the genome, and haplotype analyses that excluded each locus in which LOD scores were positive.
Because the disease-associated haplotype was carried by three unaffected offspring, III4, III6, III13 (Figure 2) of the founding father (II4), and all of his affected offspring, it is possible that the founder is a germ-line mosaic for a dominant mutation. Under this model, the haplotype shared by the nine unaffected individuals is not relevant to the disease mutation and should not be included in the linkage analysis. Similar conclusions have been reached in the genetic analysis of other disorders.13, 14 Reanalysis of the genotypes under the germ-line mosaic model revealed a maximum LOD score of 4.87 at a recombination fraction of zero at the loci for four continuous fully informative linked markers within the narrowed linked interval (D22S277, D22S1142, D22S683, and D22S283) (Supplementary Table 1).
Candidate gene analysis
A total of 92 known genes, and about 40 hypothetical proteins, and ESTs are reported within the minimal common interval described above. None of the genes was known to be involved in the developmental processes of limbs and mammary glands, or in other limb- or mammary-gland-associated diseases. On the basis of published data for each gene (NCBI, GeneCard, UCSC), we set a priority list of candidate genes for direct sequencing. We considered any gene known to be expressed in tissues related to the MDN syndrome (mammary gland, bone, cartilage, and nails), or that could be involved in the development of those tissues through known molecular pathways.15, 16 No disease-associated sequence changes were found in any of the 38 candidate genes selected for sequence analysis. However, sequence changes were identified in six of the genes (C1QTNF6, SSTR3, RAC2, LGALS1, ANKRD54, and miRN658) (Supplementary Table 2), within the interval that co-segregate with the ‘disease haplotype’. These sequence alterations were also identified in four unaffected individuals from our 100 individual control group who carry three ‘disease alleles’ from the ‘disease haplotype’, and are thus unlikely to be causative.
The sequence alterations are not identified as known polymorphisms in the NCBI and ensemble databases, and are either in-frame or localized in intronic regions. RT-PCR and cDNA analyses in five of the genes did not detect significant quantitative or qualitative differences in the expression of these genes compared with normal controls. Regarding miRN658, the sequence alteration we found is located in a nonfunctional domain at the 3′ end of the gene, a region that is not conserved among lower species.
Cytogenetic and SNP array analyses
Cytogenetic analysis of patients III10 and IV25 (Figure 2) revealed normal karyotype 46, XX in both. No CNV was identified in any of the samples along the linked interval. We did not find short homozygous regions of adjacent polymorphic DNA markers and SNPs shared by the three affected individuals tested, heterozygous in their parents and different or heterozygous in the two healthy individuals from the control group. These data further exclude the possibility of AR inheritance of MDN in this family.
Discussion
This study presents a newly described AD genetic disorder involving the breast, digits, and nails, which we have denoted MDN syndrome. We localized the disease gene to a 12 cM (4.3 Mb) interval on chromosome 22q12.3-13.1. This region has not been associated with any mammary gland or limb/digits/nails disease phenotype.
Clinical manifestations
The clinical manifestation of ODP in this cohort is similar to that previously described.7, 8, 9 The clinical manifestations and natural history of JHB described here are similar to those of the previously described mother and daughter with familial JHB.1 The Ulnar mammary syndrome (OMIM 181450), the limb mammary syndrome (OMIM 603543), and the Holt Oram syndrome (OMIM 142900) are other genetic syndromes affecting both mammary glands and limbs. However, ODP concomitant with JHB has not been documented. Evidence for molecular connections between breast and bone17, 18, 19, 20 indicate complex interactions between Wnts, FGFs, and Tbx gene families. However, the precise genetic basis and signaling pathways in embryogenesis and in the peripubertal period that affect the normal biological development of breast, digits, and nails are not yet understood.
Inheritance pattern
The dichotomy of recessive versus dominant inheritance for a novel phenotype in an isolated population is indeed a challenge in the absence of disease-associated mutations.
The AR pattern of inheritance is neither supported by the pedigree analyses nor by the molecular analyses performed.
The AD inheritance pattern of the investigated pedigree structure is consistent with other documentations of ODP.7, 8 This is the first report of AD inheritance for JHB. Linkage and haplotype analyses performed in this study exclude pseudodominant inheritance of a recessive disorder, and support the possibility of a single mutation or of mutations in two tightly linked genes, causing both ODP and JHB transmitted in an AD mode of inheritance. Though we cannot exclude the possibility of reduced penetrance, we did not observe heterogeneity in phenotype or in disease severity, which is typical when a modifier gene is involved.21, 22
Haplotype analysis revealed a common haplotype encompassing a 12 cM interval (4.3 Mb) in all 11 affected individuals who originate from the unaffected founder father, and in nine unaffected individuals who originate from the same patriarch in the pedigree. Such a pattern supports GM for the MDN mutation in the patriarch II4 rather than AD inheritance with reduced penetrance in this pedigree.13, 23
MDN syndrome locus
The MDN syndrome was mapped to chromosome 22q12.3-13.1 with a maximum LOD score of 4.87 at a recombination fraction of zero, for four informative DNA markers. Direct sequencing of 38 candidate genes in the linked interval did not reveal any disease-associated mutations. Although complete exclusion is not possible at present, the sequence alterations that were identified in five genes seem to be polymorphisms common to the Druze population in this village, rather then causative mutations for MDN.
Identification of the causative gene and mutation for MDN is of particular interest, given the unique combination of symptoms. To determine the disease-causing gene and mutation, we plan to use next-generation sequencing platforms24, 25 within the linked interval, followed by further studies to elucidate the pathogenesis of the syndrome, and identify molecular and biochemical interactive pathways that may involve breast, bones, and nails.
References
Kupfer D, Dingman D, Broadbent R : Juvenile breast hypertrophy: report of familial pattern and review of the literature. Plast Reconstr Surg 1992; 2: 303–309.
Govrin-Yehudain J, Kogan L, Cohen I, Falik-Zaccai TC : Familial juvenile hypertrophy of the breast. J Adoles Health 2004; 35: 151–155.
Misirlioglu A, Akoz T : Familial severe gigantomastia and reduction with free nipple graft vertical mammoplasty technique: report of two cases. Aesth Plast Surg 2005; 29: 205–209.
Koves HI, Zacharin M : Virginal breast hypertrophy of an 11-year old girl. J Pediatr Child Health 2007; 43: 315–317.
Dancey A, Khan M, Dawson J, Peart F : Gigantomastia – a classification and review of the literature. J Plast Reconstr Aesthet Surg 2008; 61: 493–502.
Li G, Robinson GW, Lesche R et al: Condition loss of PTEN leads to precocious development and neoplasia in the mammary gland. Development 2002; 129: 4159–4170.
Nevin NC, Thomas PS, Eedy DJ, Shepherd C : Anonychia and absence/hypoplasia of distal phalanges (Cooks syndrome): report of a second family. J Med Genet 1995; 32: 638–641.
Cooks RG, Hertz M, Katznelson MB, Goodman RM : A new nail dysplasia syndrome with onychonychia and absence and/or hypoplasia if distal phalanges. Clin Genet 1985; 27: 85–91.
Castori M, Brancati F, Mingarelli R, Mundlos S, Dallapiccola B : A novel patient with Cooks syndrome supports splitting from ‘classic’ brachydactyly type B. Am J Med Genet A 2007; 143: 195–199.
Zgaier A : Yerca Village and its Surroundings: Druze, Archaeological and Natural Sites in the Village and its Surroundings. Nesher, Israel: 1993.
Fishelson M, Geiger D : Optimizing exact genetic linkage computations. J Comput Biol 2004; 11: 263–275.
Silberstein M, Tzemach A, Dovgolevsky N, Fishelson M, Schuster A, Geiger D : Online system for faster multipoint linkage analysis via parallel execution on thousands of personal computers. Am J Hum Genet 2006; 78: 922–935.
Zlotogora J : Germ-line mosaicism. Hum Gent 1998; 102: 381–386.
Yaron Y, Orr-Urtreger A : New genetic principles. Clin Obstet Gynecol 2002; 45: 593–604.
van Bokhoven H, Jung M, Smits APT et al: Limb mammary syndrome: a new genetic disorder with mammary hypoplasia, ectrodactyly, and or other hand/foot anomalies maps to human chromosome 3q27. Am J Hum Genet 1999; 64: 538–546.
Pelton L, Palotie A, Lange K : Use of population isolates for mapping complex traits. Nat Rev Genet 2000; 1: 182–190.
Shore P : A role for Runx2 in normal mammary gland and breast cancer bone metastasis. J Cell Biochem 2005; 96: 484–489.
Eblaghie MC, Song SJ, Kim JY, Akita K, Trckle C, Jung HS : Interactions between FGF and wnt signals and TBX3 gene expression in mammary gland initiation in mouse embryos. J Anat 2004; 205: 1–13.
Turashvili G, Bouchal J, Burkadze G, Kolar Z : Wnt signaling pathway in mammary gland development and carcinogenesis. Pathobiology 2006; 73: 213–223.
Katoh M : Cross talking of WNT and FGF signaling pathways at GSK3beta to regulate beta-catenin and SNAIL signaling cascades. Cancer Biol Ther 2006; 5: 1059–1064.
Summers KM : Relationship between genotype and phenotype in monogenic diseases: relevance to polygenic diseases. Hum Mutat 1996; 7: 283–293.
Gropman AL, Adams DR : Atypical patterns of inheritance. Semin Pediatr Neurol 2007; 14: 34–45.
Martin DI, Ward R, Suter CM : Germ-line epimutation: a basis for epigenetic disease in humans. Ann NY Acad Sci 2005; 1054: 68–77.
Chan EY : Next-generation sequencing methods: impact of sequencing accuracy on SNP discovery. Methods Mol Biol 2009; 578: 95–111.
Horner DS, Pavesi G, Castrignanò T et al: Bioinformatics approaches for genomics and post genomics applications of next-generation sequencing. Brief Bioinform 2009. [e-pub ahead of print 27 October 2009] PMID:19864250.
Acknowledgements
This work was supported by Grant No. 5727 from the Israeli Ministry of Health, Grant No. 842004 from the Isvonot Foundation of the Israeli Ministry of Justice, and Grant No. 3–934 from the Israeli Ministry of Science. This work was part of the PhD dissertation of MGN. We thank Dr Esti Feldmesser and Dr Shirley Horn-Saban who operate the SNP array facility at the Weizmann Institute of Science, Israel. We also thank Dan Cohn for enlightening discussions and review of the article, Cindy Cohen and Laura Golan for editing. We acknowledge the patients and their families who participated in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Electronic database information:
Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ UCSC Genome browser, http://www.genome.ucsc.edu/ (for reference sequences) Ensemble Genome browser, http://www.ensembl.org/index.html (for reference sequences and SNPs information).
GeneCards Home page, http://www.genecards.org/ (for microsatellite data and candidate genes data).
NCBI Map Viewer, http://www.ncbi.nlm.nih.gov/mapview/ (for reference sequences).
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Genzer-Nir, M., Khayat, M., Kogan, L. et al. Mammary-digital-nail (MDN) syndrome: a novel phenotype maps to human chromosome 22q12.3–13.1. Eur J Hum Genet 18, 662–667 (2010). https://doi.org/10.1038/ejhg.2009.236
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2009.236