- TECHNOLOGY FEATURE
How to make spatial maps of gene activity — down to the cellular level
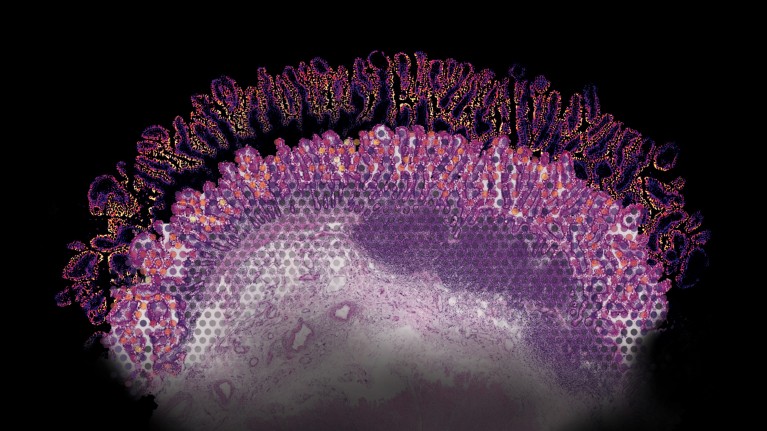
Gene-expression data captured using an array-based method for spatial transcriptomics, overlaid on a stained section of human gut tissue. Credit: Ludvig Bergenstråhle
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 606, 1036-1038 (2022)
doi: https://doi.org/10.1038/d41586-022-01743-7
References
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. Science 348, aaa6090 (2015).
Fang, R. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.11.01.466826 (2021).
Wang, X. et al. Science 361, aat5691 (2018).
Eng, C.-H. L. et al. Nature 568, 235–239 (2019).
Ståhl, P. L. et al. Science 353, 78–82 (2016).
Stickels, R. R. et al. Nature Biotechnol. 39, 313–319 (2021).
Chen, A. et al. Cell 185, 1777–1792 (2022).
Chen, A. et al. Preprint at bioRxiv https://doi.org/10.1101/2022.03.23.485448 (2022).
Wang, M. et al. Dev. Cell 57, 1271–1283 (2022).
Liu, C. et al. Dev. Cell 57, 1284-1298 (2022).
Cable, D. M. et al. Nature Biotechnol. 40, 517–526 (2022).
Bergenstråhle, L. et al. Nature Biotechnol. 40, 476–479 (2022).
Lopez, R. et al. Nature Biotechnol. https://doi.org/10.1038/s41587-022-01272-8 (2022).

 Revealing chromosome contours, one dot at a time
Revealing chromosome contours, one dot at a time
 Single-cell analysis enters the multiomics age
Single-cell analysis enters the multiomics age
 Starfish enterprise: finding RNA patterns in single cells
Starfish enterprise: finding RNA patterns in single cells
 Method of the Year: spatially resolved transcriptomics
Method of the Year: spatially resolved transcriptomics
 NatureTech hub
NatureTech hub