- OUTLOOK
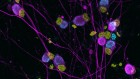
Olfactory receptors are not unique to the nose
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 606, S14-S17 (2022)
doi: https://doi.org/10.1038/d41586-022-01631-0
This article is part of Nature Outlook: Smell, an editorially independent supplement produced with the financial support of third parties. About this content.
References
Buck, L. & Axel, R. Cell 65, 175–187 (1991).
Orecchioni, M. et al. Science 375, 214–221 (2022).
Parmentier, M. et al. Nature 355, 453–455 (1992).
Zhang, X. et al. Proc. Natl Acad. Sci. USA 101, 14168–14173 (2004).
Spehr, M. et al. Science 299, 2054–2058 (2003).
Neuhaus, E. M. et al. J. Biol. Chem. 284, 16218–16225 (2009).
Busse, D. et al. J. Invest. Dermatol. 134, 2823–2832 (2014).
Jimenez, F. et al. J. Cosmet. Dermatol. 20, 784–791 (2021).
Pluznick, J. L. et al. Proc. Natl Acad. Sci. USA 106, 2059–2064 (2009).
Pluznick, J. L. et al. Proc. Natl Acad. Sci. USA 110, 4410–4415 (2013).
Pluznick, J. Gut Microbes 5, 202–207 (2014).
Shepard, B. D. et al. Sci. Rep. 6, 35215 (2016).
Li, E. et al. Cell Metab. 30, 319–328 (2019).
Cheng, J. et al. Cell Metab. 34, 240–255 (2022).

 The science of smell steps into the spotlight
The science of smell steps into the spotlight
 Unpicking the link between smell and memories
Unpicking the link between smell and memories
 The science behind COVID’s assault on smell
The science behind COVID’s assault on smell
 How to bring back the sense of smell
How to bring back the sense of smell
 The dogs learning to sniff out disease
The dogs learning to sniff out disease
 Restoring smell with an electronic nose
Restoring smell with an electronic nose
 Sniffing out smell’s effects on human behaviour
Sniffing out smell’s effects on human behaviour
 Building neural networks that smell like a brain
Building neural networks that smell like a brain