- NEWS FEATURE
- Correction 22 October 2021
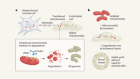
How the world’s biggest brain maps could transform neuroscience
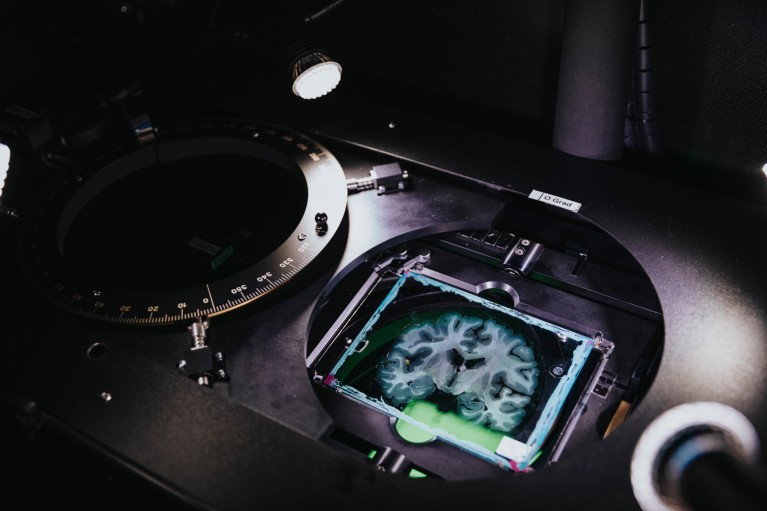
A human brain slice is placed in a microscope to visualize nerve fibres. Credit: Mareen Fischinger
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 598, 22-25 (2021)
doi: https://doi.org/10.1038/d41586-021-02661-w
Updates & Corrections
-
Correction 22 October 2021: This feature gave the wrong affiliation for Christof Koch. Koch is no longer president of the Allen Institute, but rather chief scientist of its MindScope Program. The story has also been modified to reflect the fact that John Ngai no longer has an active role at the University of California, Berkeley.
References
Witvliet, D. et al. Nature 596, 257–261 (2021).
Xu, C. S. et al. Preprint at bioRxiv https://doi.org/10.1101/2020.01.21.911859 (2020).
Motta, A. et al. Science 366, eaay3134 (2019).
Shapson-Coe, A. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.05.29.446289 (2021).
BRAIN Initiative Cell Census Network (BICCN). Nature 598, 86–102 (2021).
Bakken, T. E. et al. Nature 598, 111–119 (2021).
Musall, S. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.09.27.461599 (2021).
Berg, J. et al. Nature 598, 151–158 (2021).
Yao, Z. et al. Nature 598, 103–110 (2021).
Matho, K. S. et al. Nature 598, 182–187 (2021).
Sahel, J.-A. et al. Nature Med. 27, 1223–1229 (2021).

 The BRAIN Initiative Cell Census Network—Motor Cortex
The BRAIN Initiative Cell Census Network—Motor Cortex
 A census of cell types in the brain’s motor cortex
A census of cell types in the brain’s motor cortex
 Read the paper: A multimodal cell census and atlas of the mammalian primary motor cortex
Read the paper: A multimodal cell census and atlas of the mammalian primary motor cortex
 The search for secrets of the human brain
The search for secrets of the human brain
 Documentary follows implosion of billion-euro brain project
Documentary follows implosion of billion-euro brain project
 Worldwide brain-mapping project sparks excitement — and concern
Worldwide brain-mapping project sparks excitement — and concern